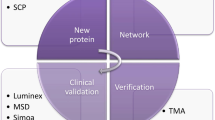
Abstract
Success in proteomics depends upon careful study design and high-quality biological samples. Advanced information technologies, and also an ability to use existing knowledge to the full, will be crucial in making sense of the data. Despite its genome-scale potential, proteome analysis is at a much earlier stage of development than genomics and gene expression (microarray) studies. Fundamental issues involving biological variability, pre-analytic factors and analytical reproducibility remain to be resolved. Consequently, the analysis of proteomics data is currently informal and relies heavily on expert opinion. Databases and software tools developed for the analysis of molecular sequences and microarrays are helpful, but are limited owing to the unique attributes of proteomics data and differing research goals.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Cambridge Healthtech Institute Conference on Human Proteome Project, 2–4 April 2001, McLean, Virgina 〈http://www.healthtech.com/2001/hpr/index.htm〉 (2001).
Krishna, R. G. & Wold, F. Post-translational modification of proteins. Adv. Enzymol. Relat. Areas Mol. Biol. 67, 265–298 (1993).
Keegan, L. P., Gallo, A. & O'Connell, M. A. The many roles of an RNA editor. Nature Rev. Genet. 2, 869–878 (2001).
Maniatis, T. & Tasic, B. Alternative pre-mRNA splicing and proteome expansion in metazoans. Nature 418, 236–243 (2002).
Dayhoff, M. O. & Eck, R. V. MASSPEC: a computer program for complete sequence analysis of large proteins from mass spectrometry data of a single sample. Comput. Biol. Med. 1, 5–28 (1970).
Anderson, N. G., Matheson, A. & Anderson, N. L. Back to the future: the human protein index (HPI) and the agenda for post-proteomic biology. Proteomics 1, 3–12 (2001).
Boguski, M. S. Bioinformatics. Curr. Opin. Genet. Dev. 4, 383–388 (1994).
Boguski, M. S. The turning point in genome research. Trends Biochem. Sci. 20, 295–296 (1995).
Zuckerkandl, E. & Pauling, L. Molecules as documents of evolutionary history. J. Theor. Biol. 8, 357–366 (1965).
Dayhoff, M. O. Computer aids to protein sequence determination. J. Theor. Biol. 8, 97–112 (1965).
Doolittle, R. F. Some reflections on the early days of sequence searching. J. Mol. Med. 75, 239–241 (1997).
Shortliffe, E. et al. (eds) Medical Informatics: Computer Applications in Health Care and Biomedicine (Springer, New York, 2000).
Hieter, P. & Boguski, M. Functional genomics: it's all how you read it. Science 278, 601–602 (1997).
Duyk, G. M. Sharper tools and simpler methods. Nature Genet. 32(Chipping Forecast II Suppl.), 465–468 (2002).
Kohane, I. S., Kho, A. T. & Butte, A. J. Microarrays For an Integrative Genomics (Massachusetts Institute of Technology Press, Cambridge, MA, 2003).
Potter, J. D. At the interfaces of epidemiology, genetics and genomics. Nature Rev. Genet. 2, 142–147 (2001).
McClatchey, K. D. (ed.) Clinical Laboratory Medicine (Lippincott, Philadelphia, 2002).
Huang, J. et al. Effects of ischemia on gene expression. J. Surg. Res. 99, 222–227 (2001).
Craven, R. A. & Banks, R. E. Laser capture microdissection and proteomics: possibilities and limitation. Proteomics 1, 1200–1204 (2001).
Craven, R. A. & Banks, R. E. Use of laser capture microdissection to selectively obtain distinct populations of cells for proteomic analysis. Methods Enzymol. 356, 33–49 (2002).
Margolin, J. From comparative and functional genomics to practical decisions in the clinic: a view from the trenches. Genome Res. 11, 923–925 (2001).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Statist. Soc. B 57, 289–300 (1995).
Dayhoff, M. O. & Eck, R. V. Atlas of Protein Sequence and Structure (National Biomedical Research Foundation, Silver Spring, MD, 1966).
Smith, T. F. The history of the genetic sequence databases. Genomics 6, 701–707 (1990).
Bairoch, A. & Boeckmann, B. The SWISS-PROT protein sequence data bank. Nucleic Acids Res. 19 (Suppl.), 2247–2249 (1991).
Maglott, D. R. et al. NCBI's LocusLink and RefSeq. Nucleic Acids Res. 28, 126–128 (2000).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nature Genet. 25, 25–29 (2000).
Bader, G. D. et al. BIND—The Biomolecular Interaction Network Database. Nucleic Acids Res. 29, 242–245 (2001).
Adkins, J. N. et al. Toward a human blood serum proteome: analysis by multidimensional separation coupled with mass spectrometry. Mol. Cell. Proteomics 1, 947–955 (2002).
Kratz, A. & Lewandrowski, K. B. Case records of the Massachusetts General Hospital. Weekly clinicopathological exercises. Normal reference laboratory values. N. Engl. J. Med. 339, 1063–1072 (1998).
Jung, E. et al. Annotation of glycoproteins in the SWISS-PROT database. Proteomics 1, 262–268 (2001).
Anderson, N. L. & Anderson, N. G. The human plasma proteome: history, character, and diagnostic prospects. Mol. Cell. Proteomics 1, 845–867 (2002).
Chakravarti, D. N., Chakravarti, B. & Moutsatsos, I. Informatic tools for proteome profiling. Biotechniques 32(Comput. Proteomics Suppl.), S4–S15 (2002).
Liebler, D. C. Introduction to Proteomics (Humana, Totowa, NJ, 2002).
The Association of Biomolecular Resource Facilities. Delta Mass: A Database of Protein Post Translational Modifications 〈http://www.abrf.org/index.cfm/dm.home〉 (2002).
Wilkins, M. R. et al. High-throughput mass spectrometric discovery of protein post-translational modifications. J. Mol. Biol. 289, 645–657 (1999).
Creasy, D. M. & Cottrell, J. S. Error tolerant searching of uninterpreted tandem mass spectrometry data. Proteomics 2, 1426–1434 (2002).
Choudhary, J. S. et al. Matching peptide mass spectra to EST and genomic DNA databases. Trends Biotechnol. 19 (Suppl.), S17–S22 (2001).
Choudhary, J. S. et al. Interrogating the human genome using uninterpreted mass spectrometry data. Proteomics 1, 651–667 (2001).
Bafna, V. & Edwards, N. SCOPE: a probabilistic model for scoring tandem mass spectra against a peptide database. Bioinformatics 17 (Suppl.) S13–S21 (2001).
Eng, J., McCormack, A. & Yates, J. An approach to correlate tandem mass spectral data of peptides with amino acid sequences in a protein database. J. Am. Soc. Mass Spectrom. 5, 976–989 (1994).
Fenyo, D. Identifying the proteome: software tools. Curr. Opin. Biotechnol. 11, 391–395 (2000).
Field, H. I., Fenyo, D. & Beavis, R. C. RADARS, a bioinformatics solution that automates proteome mass spectral analysis, optimises protein identification, and archives data in a relational database. Proteomics 2, 36–47 (2002).
Perkins, D. N. et al. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Efron, B. & Tibshirani, R. Empirical Bayes methods and false discovery rates for microarrays. Genet. Epidemiol. 23, 70–86 (2002).
Pepe, M. S. et al. Selecting differentially expressed genes from microarray experiments. Biometrics (in the press).
Keller, A. et al. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Adam, B. L. et al. Serum protein fingerprinting coupled with a pattern-matching algorithm distinguishes prostate cancer from benign prostate hyperplasia and healthy men. Cancer Res. 62, 3609–3614 (2002).
Petricoin, E. F. III et al. Serum proteomic patterns for detection of prostate cancer. J. Natl Cancer Inst. 94, 1576–1578 (2002).
Petricoin, E. F. et al. Use of proteomic patterns in serum to identify ovarian cancer. Lancet 359, 572–577 (2002).
Qu, Y. et al. Boosted decision tree analysis of surface-enhanced laser desorption/ionization mass spectral serum profiles discriminates prostate cancer from noncancer patients. Clin. Chem. 48, 1835–1843 (2002).
Pepe, M. S. et al. Phases of biomarker development for early detection of cancer. J. Natl Cancer Inst. 93, 1054–1061 (2001).
Judson, H. The Eighth Day of Creation: Makers of the Revolution in Biology expand. edn (Cold Spring Harbor Laboratory Press, New York, 1996)
Hayles, N. How We Became Posthuman: Virtual Bodies in Cybernetics, Literature, and Informatics (University of Chicago Press, Chicago, 1999).
Bonini, P. et al. Errors in laboratory medicine. Clin. Chem. 48, 691–698 (2002).
Narayanan, S. The preanalytic phase. An important component of laboratory medicine. Am. J. Clin. Pathol. 113, 429–452 (2000).
Spellman, P. T. et al. Design and implementation of microarray gene expression markup language (MAGE-ML). Genome Biol. 3, 46 (2002).
Brazma, A. et al. Minimum information about a microarray experiment (MIAME)—toward standards for microarray data. Nature Genet. 29, 365–371 (2001).
Editorial. Coming to terms with microarrays. Nature Genet. 32, 333–334 (2002).
Ball, C. et al. Standards for Microarray Data. Science 298, 539 (2002).
Orchard, S., Kersey, P., Hermjakob, H. & Apweiler, R. The HUPO proteomics standards initiative meeting: towards common standards for exchanging proteomics data. Comp. Funct. Genom. 4, 16–19 (2003).
Bader, G. D. & Hogue, C. W. BIND—a data specification for storing and describing biomolecular interactions, molecular complexes and pathways. Bioinformatics 16, 465–477 (2000).
Abiteboul, S., Buneman, P. & Suciu, D. Data on the Web: From Relations to Semistructured Data and XML (Morgan Kaufmann, San Francisco, 2000).
Coyle, F. XML, Web Services, and the Data Revolution (Addison-Wesley, Boston, 2002).
Acknowledgements
We thank L. Hartwell, J. Potter and G. Omenn for stimulating discussions and J. Gray, J. Pounds and L. Geer for valuable suggestions and critical readings of the manuscript.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Boguski, M., McIntosh, M. Biomedical informatics for proteomics. Nature 422, 233–237 (2003). https://doi.org/10.1038/nature01515
Issue Date:
DOI: https://doi.org/10.1038/nature01515
This article is cited by
-
Nucleic acid aptamers for clinical diagnosis: cell detection and molecular imaging
Analytical and Bioanalytical Chemistry (2011)
-
Open Biomedical Ontology-based Medline exploration
BMC Bioinformatics (2009)
-
A national clinical decision support infrastructure to enable the widespread and consistent practice of genomic and personalized medicine
BMC Medical Informatics and Decision Making (2009)
-
Evaluation of a joint Bioinformatics and Medical Informatics international course in Peru
BMC Medical Education (2008)
-
Pathogen profiling for disease management and surveillance
Nature Reviews Microbiology (2007)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.