Abstract
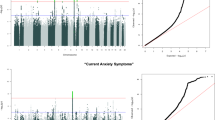
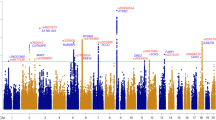
Anxiety disorders (ADs), namely generalized AD, panic disorder and phobias, are common, etiologically complex conditions with a partially genetic basis. Despite differing on diagnostic definitions based on clinical presentation, ADs likely represent various expressions of an underlying common diathesis of abnormal regulation of basic threat–response systems. We conducted genome-wide association analyses in nine samples of European ancestry from seven large, independent studies. To identify genetic variants contributing to genetic susceptibility shared across interview-generated DSM-based ADs, we applied two phenotypic approaches: (1) comparisons between categorical AD cases and supernormal controls, and (2) quantitative phenotypic factor scores (FS) derived from a multivariate analysis combining information across the clinical phenotypes. We used logistic and linear regression, respectively, to analyze the association between these phenotypes and genome-wide single nucleotide polymorphisms. Meta-analysis for each phenotype combined results across the nine samples for over 18 000 unrelated individuals. Each meta-analysis identified a different genome-wide significant region, with the following markers showing the strongest association: for case–control contrasts, rs1709393 located in an uncharacterized non-coding RNA locus on chromosomal band 3q12.3 (P=1.65 × 10−8); for FS, rs1067327 within CAMKMT encoding the calmodulin-lysine N-methyltransferase on chromosomal band 2p21 (P=2.86 × 10−9). Independent replication and further exploration of these findings are needed to more fully understand the role of these variants in risk and expression of ADs.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Kessler RC, Berglund P, Demler O, Jin R, Merikangas KR, Walters EE . Lifetime prevalence and age-of-onset distributions of DSM-IV disorders in the National Comorbidity Survey Replication. Arch Gen Psychiatry 2005; 62: 593–602.
Hettema JM, Neale MC, Kendler KS . A review and meta-analysis of the genetic epidemiology of anxiety disorders. Am J Psychiatry 2001; 158: 1568–1578.
Smoller JW, Block SR, Young MM . Genetics of anxiety disorders: the complex road from DSM to DNA. Depress Anxiety 2009; 26: 965–975.
Maron E, Hettema JM, Shlik J . Advances in molecular genetics of panic disorder. Mol Psychiatry 2010; 15: 681–701.
Erhardt A, Czibere L, Roeske D, Lucae S, Unschuld PG, Ripke S et al. TMEM132D, a new candidate for anxiety phenotypes: evidence from human and mouse studies. Mol Psychiatry 2011; 16: 647–663.
Otowa T, Kawamura Y, Nishida N, Sugaya N, Koike A, Yoshida E et al. Meta-analysis of genome-wide association studies for panic disorder in the Japanese population. Transl Psychiatry 2012; 2: e186.
Logue MW, Baldwin C, Guffanti G, Melista E, Wolf EJ, Reardon AF et al. A genome-wide association study of post-traumatic stress disorder identifies the retinoid-related orphan receptor alpha (RORA) gene as a significant risk locus. Mol Psychiatry 2013; 18: 937–942.
Xie P, Kranzler HR, Yang C, Zhao H, Farrer LA, Gelernter J . Genome-wide association study identifies new susceptibility loci for posttraumatic stress disorder. Biol Psychiatry 2013; 74: 656–663.
Guffanti G, Galea S, Yan L, Roberts AL, Solovieff N, Aiello AE et al. Genome-wide association study implicates a novel RNA gene, the lincRNA AC068718.1, as a risk factor for post-traumatic stress disorder in women. Psychoneuroendocrinology 2013; 38: 3029–3038.
Stewart SE, Yu D, Scharf JM, Neale BM, Fagerness JA, Mathews CA et al. Genome-wide association study of obsessive-compulsive disorder. Mol Psychiatry 2013; 18: 788–798.
Mattheisen M, Samuels JF, Wang Y, Greenberg BD, Fyer AJ, McCracken JT et al. Genome-wide association study in obsessive-compulsive disorder: results from the OCGAS. Mol Psychiatry 2014; 20: 337–344.
Walter S, Glymour MM, Koenen K, Liang L, Tchetgen Tchetgen EJ, Cornelis M et al. Performance of polygenic scores for predicting phobic anxiety. PLoS One 2013; 8: e80326.
Craske MG, Rauch SL, Ursano R, Prenoveau J, Pine DS, Zinbarg RE . What is an anxiety disorder? Depress Anxiety 2009; 26: 1066–1085.
Comorbidity of Mood and Anxiety Disorders American Psychiatric Press: Washington DC, USA, 1990.
Hettema JM, Prescott CA, Myers JM, Neale MC, Kendler KS . The structure of genetic and environmental risk factors for anxiety disorders in men and women. Arch Gen Psychiatry 2005; 62: 182–189.
Middeldorp CM, Cath DC, van Dyck R, Boomsma D . The co-morbidity of anxiety and depression in the perspective of genetic epidemiology. A review of twin and family studies. Psychol Med 2005; 35: 611–624.
Kendler KS, Prescott CA, Myers J, Neale MC . The structure of genetic and environmental risk factors for common psychiatric and substance use disorders in men and women. Arch Gen Psychiatry 2003; 60: 929–937.
Plomin R, Haworth CM, Davis OS . Common disorders are quantitative traits. Nat Rev Genet 2009; 10: 872–878.
Hettema JM, An SS, Neale MC, Bukszar J, van den Oord EJ, Kendler KS et al. Association between glutamic acid decarboxylase genes and anxiety disorders, major depression, and neuroticism. Mol Psychiatry 2006; 11: 752–762.
Howie B, Fuchsberger C, Stephens M, Marchini J, Abecasis GR . Fast and accurate genotype imputation in genome-wide association studies through pre-phasing. Nat Genet 2012; 44: 955–959.
Li Y, Willer CJ, Ding J, Scheet P, Abecasis GR . MaCH: using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet Epidemiol 2010; 34: 816–834.
Muthen LK, Muthen BO . Mplus User's Guide. Sixth Edition, Muthen & Muthen: Los Angeles, CA, 2010.
Willer CJ, Li Y, Abecasis GR . METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 2010; 26: 2190–2191.
Storey JD, Tibshirani R . Statistical significance for genomewide studies. Proc Natl Acad Sci USA 2003; 100: 9440–9445.
Li MX, Sham PC, Cherny SS, Song YQ . A knowledge-based weighting framework to boost the power of genome-wide association studies. PLoS One 2010; 5: e14480.
Li MX, Gui HS, Kwan JS, Sham PC . GATES: a rapid and powerful gene-based association test using extended Simes procedure. Am J Hum Genet 2011; 88: 283–293.
van den Oord EJ, Sullivan PF . False discoveries and models for gene discovery. Trends Genet 2003; 19: 537–542.
Yang J, Lee SH, Goddard ME, Visscher PM . GCTA: a tool for genome-wide complex trait analysis. Am J Hum Genet 2011; 88: 76–82.
Bulik-Sullivan BK, Loh PR, Finucane HK, Ripke S, Yang J, Patterson N et al. LD Score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 2015; 47: 291–295.
Wray NR, Lee SH, Mehta D, Vinkhuyzen AA, Dudbridge F, Middeldorp CM . Research review: Polygenic methods and their application to psychiatric traits. J Child Psychol Psychiatry 2014; 55: 1068–1087.
Martens K, Jaeken J, Matthijs G, Creemers JW . Multi-system disorder syndromes associated with cystinuria type I. Curr Mol Med 2008; 8: 544–550.
Saravakos P, Kokkinou V, Giannatos E . Cystinuria: current diagnosis and management. Urology 2014; 83: 693–699.
Martens K, Derua R, Meulemans S, Waelkens E, Jaeken J, Matthijs G et al. PREPL: a putative novel oligopeptidase propelled into the limelight. Biol Chem 2006; 387: 879–883.
Magnani R, Dirk LM, Trievel RC, Houtz RL . Calmodulin methyltransferase is an evolutionarily conserved enzyme that trimethylates Lys-115 in calmodulin. Nat Commun 2010; 1: 43.
Neale BM, Sklar P . Genetic analysis of schizophrenia and bipolar disorder reveals polygenicity but also suggests new directions for molecular interrogation. Curr Opin Neurobiol 2015; 30C: 131–138.
Yang J, Wray NR, Visscher PM . Comparing apples and oranges: equating the power of case-control and quantitative trait association studies. Genet Epidemiol 2010; 34: 254–257.
van der Sluis S, Posthuma D, Nivard MG, Verhage M, Dolan CV . Power in GWAS: lifting the curse of the clinical cut-off. Mol Psychiatry 2013; 18: 2–3.
Visscher PM, Brown MA, McCarthy MI, Yang J . Five years of GWAS discovery. Am J Hum Genet 2012; 90: 7–24.
Demirkan A, Penninx BW, Hek K, Wray NR, Amin N, Aulchenko YS et al. Genetic risk profiles for depression and anxiety in adult and elderly cohorts. Mol Psychiatry 2011; 16: 773–783.
Otowa T, Maher BS, Aggen SH, McClay JL, van den Oord EJ, Hettema JM . Genome-wide and gene-based association studies of anxiety disorders in European and African American samples. PLoS One 2014; 9: e112559.
Insel T, Cuthbert B, Garvey M, Heinssen R, Pine DS, Quinn K et al. Research domain criteria (RDoC): toward a new classification framework for research on mental disorders. Am J Psychiatry 2010; 167: 748–751.
Bienvenu OJ, Samuels JF, Wuyek LA, Liang KY, Wang Y, Grados MA et al. Is obsessive-compulsive disorder an anxiety disorder, and what, if any, are spectrum conditions? A family study perspective. Psychol Med 2012; 42: 1–13.
Bienvenu OJ, Hettema JM, Neale MC, Prescott CA, Kendler KS . Low extraversion and high neuroticism as indices of genetic and environmental risk for social phobia, agoraphobia, and animal phobia. Am J Psychiatry 2007; 164: 1714–1721.
Cross-Disorder Group of the Psychiatric GWAS Consortium. Identification of risk loci with shared effects on five major psychiatric disorders: a genome-wide analysis. Lancet 2013; 381: 1371–1379.
Sanders AR, Levinson DF, Duan J, Dennis JM, Li R, Kendler KS et al. The Internet-based MGS2 control sample: self report of mental illness. Am J Psychiatry 2010; 167: 854–865.
Preisig M, Waeber G, Vollenweider P, Bovet P, Rothen S, Vandeleur C et al. The PsyCoLaus study: methodology and characteristics of the sample of a population-based survey on psychiatric disorders and their association with genetic and cardiovascular risk factors. BMC Psychiatry 2009; 9: 9.
Hofman A, Breteler MM, van Duijn CM, Krestin GP, Pols HA, Stricker BH et al. The Rotterdam Study: objectives and design update. Eur J Epidemiol 2007; 22: 819–829.
Schmidt CO, Watzke AB, Schulz A, Baumeister SE, Freyberger HJ, Grabe HJ . [The lifetime prevalence of mental disorders in north-eastern Germany. What is the influence of earlier mental morbidity on survey participation and prevalence estimates? Results from the SHIP-study]. Psychiatr Prax 2013; 40: 192–199.
Jardine R, Martin NG, Henderson AS . Genetic covariation between neuroticism and the symptoms of anxiety and depression. Genet Epidemiol 1984; 1: 89–107.
Oldehinkel AJ, Rosmalen JG, Buitelaar JK, Hoek HW, Ormel J, Raven D et al. Cohort profile update: the tracking adolescents' individual lives survey (TRAILS). Int J Epidemiol 2015; 44: 76–76n.
Penninx BW, Beekman AT, Smit JH, Zitman FG, Nolen WA, Spinhoven P et al. The Netherlands Study of Depression and Anxiety (NESDA): rationale, objectives and methods. Int J Methods Psychiatr Res 2008; 17: 121–140.
Boomsma DI, Vink JM, Van Beijsterveldt TC, de Geus EJ, Beem AL, Mulder EJ et al. Netherlands Twin Register: a focus on longitudinal research. Twin Res 2002; 5: 401–406.
Boomsma DI, de Geus EJ, Vink JM, Stubbe JH, Distel MA, Hottenga JJ et al. Netherlands Twin Register: from twins to twin families. Twin Res Hum Genet 2006; 9: 849–857.
Acknowledgements
This overall project was supported by NIH grant R01MH87646 to JMH. TO was supported by a research fellowship from the Japan Society for the Promotion of Science (no. 21-8373). Samples and associated phenotype data for the MGS study were collected under the following grants: NIMH Schizophrenia Genetics Initiative U01s: MH046276 (CR Cloninger), MH46289 (C Kaufmann), and MH46318 (MT Tsuang); and MGS Part 1 (MGS1) and Part 2 (MGS2) R01s: MH67257 (NG Buccola), MH59588 (BJ Mowry), MH59571 (PV Gejman), MH59565(R Freedman), MH59587 (F Amin), MH60870 (WF Byerley), MH59566 (DW Black), MH59586 (JM Silverman), MH61675 (DF Levinson) and MH60879 (CR Cloninger). Analyses in the Rotterdam Study were supported by the Research Institute for Diseases in the Elderly (014-93-015; RIDE2), The Netherlands Genomics Initiative (NGI)/Netherlands Consortium for Healthy Ageing (NCHA) project No. 050-060-810. The work of HT is supported by Vidi (grant 017.106.370). The Rotterdam Study is funded by Erasmus Medical Center, Rotterdam, The Netherlands Organization for the Health Research and Development (ZonMw), the Ministry of Education, Culture and Science, and the Ministry for Health, Welfare and Sports. SHIP is part of the Community Medicine Research net of the University of Greifswald, Germany, which is funded by the Federal Ministry of Education and Research (grants no. 01ZZ9603, 01ZZ0103 and 01ZZ0403), the Ministry of Cultural Affairs and the Social Ministry of the Federal State of Mecklenburg-West Pomerania. Genome-wide data have been supported by the Federal Ministry of Education and Research (grant no. 03ZIK012) and a joint grant from Siemens Healthcare, Erlangen, Germany and the Federal State of Mecklenburg-West Pomerania. This work was also funded by the German Research Foundation (DFG: GR 1912/5-1). The University of Greifswald is a member of the Caché Campus program of the InterSystems GmbH. The QIMR samples are made available through the generous and willing participation of twins and their families registered at the Australian Twin Registry and through grant funding awarded from many grant funding bodies including the Australian National Health and Medical Research Council (241944, 339462, 389927, 389875, 389891, 389892, 389938, 442915, 442981, 496675, 496739, 552485, 552498, 613608), the FP-5 GenomEUtwin Project (QLG2-CT- 2002-01254), the US National Institutes of Health (NIH grants AA07535, AA10248, AA13320, AA13321, AA13326, AA14041, MH66206, DA12854, DA019951) and the Center for Inherited Disease Research, Baltimore. We thank Dixie Statham (sample collection); Leanne Wallace, Anthony Caracella and staff of the Molecular Epidemiology Laboratory (DNA processing); David Smyth, Harry Beeby and Daniel Park (IT support). Statistical analyses were partly conducted at the Genetic Cluster Computer (http://www.geneticcluster.org), which is financially supported by the Netherlands Scientific Organization (NWO 480-05-003). EMB (1053639), SEM, GM and NRW (613602, 1078901) are supported by the National Health and Medical Research Council Fellowship Scheme. The CoLaus|PsyCoLaus study was and is supported by research grants from GlaxoSmithKline, the Faculty of Biology and Medicine of Lausanne, and the Swiss National Science Foundation (grants 3200B0–105993, 3200B0-118308, 33CSCO-122661, 33CS30-139468 and 33CS30-148401). Participating centers of TRAILS include various departments of the University Medical Center and University of Groningen, the Erasmus University Medical Center Rotterdam, the University of Utrecht, the Radboud Medical Center Nijmegen and the Parnassia Bavo group, all in the Netherlands. TRAILS has been financially supported by various grants from the Netherlands Organization for Scientific Research NWO (Medical Research Council program grant GB-MW 940-38-011; ZonMW Brainpower grant 100-001-004; ZonMw Risk Behavior and Dependence grants 60-60600-97-118; ZonMw Culture and Health grant 261-98-710; Social Sciences Council medium-sized investment grants GB-MaGW 480-01-006 and GB-MaGW 480-07-001; Social Sciences Council project grants GB-MaGW 452-04-314 and GB-MaGW 452-06-004; NWO large-sized investment grant 175.010.2003.005; NWO Longitudinal Survey and Panel Funding 481-08-013), the Dutch Ministry of Justice (WODC), the European Science Foundation (EuroSTRESS project FP-006), Biobanking and Biomolecular Resources Research Infrastructure BBMRI-NL (CP 32), the participating universities, and Accare Center for Child and Adolescent Psychiatry. Statistical analyses were carried out on the Genetic Cluster Computer (http://www.geneticcluster.org), which is financially supported by the Netherlands Scientific Organization (NWO 480-05-003) along with a supplement from the Dutch Brain Foundation. Funding for the NESDA/NTR studies was obtained from the Netherlands Organization for Scientific Research (Geestkracht program grant 10-000-1002; 904-61-090, 985-10- 002, 904-61-193, 480-04-004, 400-05-717, 912-100-20; Spinozapremie 56-464-14192); the Center for Medical Systems Biology (CSMB, NWO Genomics), Biobanking and Biomolecular Resources Research Infrastructure (BBMRI-NL), VU University’s Institutes for Health and Care Research (EMGO+) and Neuroscience Campus Amsterdam, European Research Council (ERC, 230374),National Institutes of Health (NIH, R01D0042157-01 A, MH081802, Grand Opportunity grants 1RC2 MH089951 and 1RC2 MH089995). Part of the genotyping and analyses were funded by the Genetic Association Information Network (GAIN) of the Foundation for the National Institutes of Health. Computing was supported by BiG Grid, the Dutch e-Science Grid, which is financially supported by NWO.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Supplementary information
Rights and permissions
About this article
Cite this article
Otowa, T., Hek, K., Lee, M. et al. Meta-analysis of genome-wide association studies of anxiety disorders. Mol Psychiatry 21, 1391–1399 (2016). https://doi.org/10.1038/mp.2015.197
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2015.197
This article is cited by
-
Body weight in neurological and psychiatric disorders: a large prospective cohort study
Nature Mental Health (2024)
-
Causal relationships between blood lipids and major psychiatric disorders: Univariable and multivariable mendelian randomization analysis
BMC Medical Genomics (2023)
-
Depressive symptoms, anxiety and cognitive impairment: emerging evidence in multiple sclerosis
Translational Psychiatry (2023)
-
Investigating the shared genetic architecture and causal relationship between pain and neuropsychiatric disorders
Human Genetics (2023)
-
Epigenome-wide DNA methylation analysis of whole blood cells derived from patients with GAD and OCD in the Chinese Han population
Translational Psychiatry (2022)