Abstract
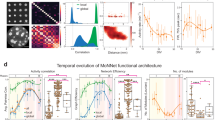
Numerous risk genes have recently been implicated in susceptibility to autism and schizophrenia. Translating such genetic findings into disease-relevant neurobiological mechanisms is challenging due to the lack of throughput assays that can be used to assess their functions on an appropriate scale. To address this issue, we explored the feasibility of using a micro-electrode array (MEA) as a potentially scalable assay to identify the electrical network phenotypes associated with risk genes. We first characterized local and global network firing in cortical neurons with MEAs, and then developed methods to analyze the alternation between the network active period (NAP) and the network inactive period (NIP), each of which lasts tens of seconds. We then evaluated the electric phenotypes of neurons derived from Shank3 knockout (KO) mice. Cortical neurons cultured on MEAs displayed a rich repertoire of spontaneous firing, and Shank3 deletion led to reduced firing activity. Enhancing excitation with CX546 rescued the deficit in the spike rate in the Shank3 KO network. In addition, the Shank3 KO network produced a shorter NIP, and this altered network firing pattern was normalized by clonazepam, a positive modulator of the GABAA receptor. MEA recordings revealed electric phenotypes that displayed altered excitation and inhibition in the network lacking Shank3. Thus, our study highlights MEAs as an experimental framework for measuring multiple robust neurobiological end points in dynamic networks and as an assay system that could be used to identify electric phenotypes in cultured neuronal networks and to analyze additional risk genes identified in psychiatric genetics.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Iossifov I, O'Roak BJ, Sanders SJ, Ronemus M, Krumm N, Levy D et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 2014; 515: 216–221.
Fromer M, Pocklington AJ, Kavanagh DH, Williams HJ, Dwyer S, Gormley P et al. De novo mutations in schizophrenia implicate synaptic networks. Nature 2014; 506: 179–184.
Schizophrenia Working Group of the Psychiatric Genomics Consortium. Biological insights from 108 schizophrenia-associated genetic loci. Nature 2014; 511: 421–427.
Sharma K, Choi SY, Zhang Y, Nieland TJ, Long S, Li M et al. High-throughput genetic screen for synaptogenic factors: identification of LRP6 as critical for excitatory synapse development. Cell Rep 2013; 5: 1330–1341.
Hempel CM, Sivula M, Levenson JM, Rose DM, Li B, Sirianni AC et al. A system for performing high throughput assays of synaptic function. PLoS One 2011; 6: e25999.
Maeda E, Robinson HP, Kawana A . The mechanisms of generation and propagation of synchronized bursting in developing networks of cortical neurons. J Neurosci 1995; 15: 6834–6845.
Chiappalone M, Bove M, Vato A, Tedesco M, Martinoia S . Dissociated cortical networks show spontaneously correlated activity patterns during in vitro development. Brain Res 2006; 1093: 41–53.
Wagenaar DA, Pine J, Potter SM . An extremely rich repertoire of bursting patterns during the development of cortical cultures. BMC Neurosci 2006; 7: 11.
Mok SY, Nadasdy Z, Lim YM, Goh SY . Ultra-slow oscillations in cortical networks in vitro. Neuroscience 2012; 206: 17–24.
Jimbo Y, Tateno T, Robinson HP . Simultaneous induction of pathway-specific potentiation and depression in networks of cortical neurons. Biophys J 1999; 76: 670–678.
Compte A, Sanchez-Vives MV, McCormick DA, Wang XJ . Cellular and network mechanisms of slow oscillatory activity (<1 Hz) and wave propagations in a cortical network model. J Neurophysiol 2003; 89: 2707–2725.
Crunelli V, Hughes SW . The slow (<1 Hz) rhythm of non-REM sleep: a dialogue between three cardinal oscillators. Nat Neurosci 2010; 13: 9–17.
Gray CM, Konig P, Engel AK, Singer W . Oscillatory responses in cat visual cortex exhibit inter-columnar synchronization which reflects global stimulus properties. Nature 1989; 338: 334–337.
Hales CM, Zeller-Townson R, Newman JP, Shoemaker JT, Killian NJ, Potter SM . Stimulus-evoked high frequency oscillations are present in neuronal networks on microelectrode arrays. Front Neural Circuits 2012; 6: 29.
Holcman D, Tsodyks M . The emergence of Up and Down states in cortical networks. PLoS Comput Biol 2006; 2: e23.
Persi E, Horn D, Volman V, Segev R, Ben-Jacob E . Modeling of synchronized bursting events: the importance on inhomogeneity. Neural Comput 2004; 16: 2577–2595.
Segev R, Baruchi I, Hulata E, Ben-Jacob E . Hidden neuronal correlations in cultured networks. Phys Rev Lett 2004; 92: 118102.
Cao Z, Hulsizer S, Tassone F, Tang HT, Hagerman RJ, Rogawski MA et al. Clustered burst firing in FMR1 premutation hippocampal neurons: amelioration with allopregnanolone. Hum Mol Genet 2012; 21: 2923–2935.
Bateup HS, Johnson CA, Denefrio CL, Saulnier JL, Kornacker K, Sabatini BL . Excitatory/inhibitory synaptic imbalance leads to hippocampal hyperexcitability in mouse models of tuberous sclerosis. Neuron 2013; 78: 510–522.
Peca J, Feliciano C, Ting JT, Wang W, Wells MF, Venkatraman TN et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 2011; 472: 437–442.
Baird G, Simonoff E, Pickles A, Chandler S, Loucas T, Meldrum D et al. Prevalence of disorders of the autism spectrum in a population cohort of children in South Thames: the Special Needs and Autism Project (SNAP). Lancet 2006; 368: 210–215.
Precht KS, Lese CM, Spiro RP, Huttenlocher PR, Johnston KM, Baker JC et al. Two 22q telomere deletions serendipitously detected by FISH. J Med Genet 1998; 35: 939–942.
Wilson HL, Wong AC, Shaw SR, Tse WY, Stapleton GA, Phelan MC et al. Molecular characterisation of the 22q13 deletion syndrome supports the role of haploinsufficiency of SHANK3/PROSAP2 in the major neurological symptoms. J Med Genet 2003; 40: 575–584.
Gauthier J, Spiegelman D, Piton A, Lafreniere RG, Laurent S, St-Onge J et al. Novel de novo SHANK3 mutation in autistic patients. Am J Med Genet B Neuropsychiatr Genet 2009; 150B: 421–424.
Durand CM, Betancur C, Boeckers TM, Bockmann J, Chaste P, Fauchereau F et al. Mutations in the gene encoding the synaptic scaffolding protein SHANK3 are associated with autism spectrum disorders. Nat Genet 2007; 39: 25–27.
Boeckers TM, Bockmann J, Kreutz MR, Gundelfinger ED . ProSAP/Shank proteins - a family of higher order organizing molecules of the postsynaptic density with an emerging role in human neurological disease. J Neurochem 2002; 81: 903–910.
Shcheglovitov A, Shcheglovitova O, Yazawa M, Portmann T, Shu R, Sebastiano V et al. SHANK3 and IGF1 restore synaptic deficits in neurons from 22q13 deletion syndrome patients. Nature 2013; 503: 267–271.
Baron MK, Boeckers TM, Vaida B, Faham S, Gingery M, Sawaya MR et al. An architectural framework that may lie at the core of the postsynaptic density. Science 2006; 311: 531–535.
Hayashi MK, Tang C, Verpelli C, Narayanan R, Stearns MH, Xu RM et al. The postsynaptic density proteins Homer and Shank form a polymeric network structure. Cell 2009; 137: 159–171.
Sheng M, Kim E . The Shank family of scaffold proteins. J Cell Sci 2000; 113: 1851–1856.
Ramamoorthi K, Fropf R, Belfort GM, Fitzmaurice HL, McKinney RM, Neve RL et al. Npas4 regulates a transcriptional program in CA3 required for contextual memory formation. Science 2011; 334: 1669–1675.
Mienville JM, Vicini S . Pregnenolone sulfate antagonizes GABAA receptor-mediated currents via a reduction of channel opening frequency. Brain Res 1989; 489: 190–194.
Nagarajan N, Quast C, Boxall AR, Shahid M, Rosenmund C . Mechanism and impact of allosteric AMPA receptor modulation by the ampakine CX546. Neuropharmacology 2001; 41: 650–663.
Lu C, Fu Z, Karavanov I, Yasuda RP, Wolfe BB, Buonanno A et al. NMDA receptor subtypes at autaptic synapses of cerebellar granule neurons. J Neurophysiol 2006; 96: 2282–2294.
Han S, Tai C, Jones CJ, Scheuer T, Catterall WA . Enhancement of inhibitory neurotransmission by GABAA receptors having alpha2,3-subunits ameliorates behavioral deficits in a mouse model of autism. Neuron 2014; 81: 1282–1289.
Han S, Tai C, Westenbroek RE, Yu FH, Cheah CS, Potter GB et al. Autistic-like behaviour in Scn1a+/- mice and rescue by enhanced GABA-mediated neurotransmission. Nature 2012; 489: 385–390.
Huntsman MM, Huguenard JR . Fast IPSCs in rat thalamic reticular nucleus require the GABAA receptor beta1 subunit. J Physiol 2006; 572: 459–475.
Chen X, Dzakpasu R . Observed network dynamics from altering the balance between excitatory and inhibitory neurons in cultured networks. Phys Rev E Stat Nonlin Soft Matter Phys 2010; 82: 031907.
Gogolla N, Takesian AE, Feng G, Fagiolini M, Hensch TK . Sensory integration in mouse insular cortex reflects GABA circuit maturation. Neuron 2014; 83: 894–905.
Aladjalova NA . Infra-slow rhythmic oscillations of the steady potential of the cerebral cortex. Nature 1957; 179: 957–959.
Wilson HR, Cowan JD . Excitatory and inhibitory interactions in localized populations of model neurons. Biophys J 1972; 12: 1–24.
Rubenstein JL, Merzenich MM . Model of autism: increased ratio of excitation/inhibition in key neural systems. Genes Brain Behav 2003; 2: 255–267.
Gogolla N, Leblanc JJ, Quast KB, Sudhof TC, Fagiolini M, Hensch TK . Common circuit defect of excitatory-inhibitory balance in mouse models of autism. J Neurodev Disord 2009; 1: 172–181.
Chao HT, Chen H, Samaco RC, Xue M, Chahrour M, Yoo J et al. Dysfunction in GABA signalling mediates autism-like stereotypies and Rett syndrome phenotypes. Nature 2010; 468: 263–269.
Valdivia P, Martin M, LeFew WR, Ross J, Houck KA, Shafer TJ . Multi-well microelectrode array recordings detect neuroactivity of ToxCast compounds. Neurotoxicology 2014; 44C: 204–217.
Acknowledgements
We thank Dr Stefano Vicini from Georgetown University for helpful discussions. We thank Caroline Maguire, Gregory Foos and Triana Dalia for technical support. We thank the members of the Feng Lab and the Pan Group for helpful discussions. This work was supported by National Institute of Health (R21MH099448 to JQP and R01MH097104 to GF), and by the Stanley Center for Psychiatric Research at the Broad Institute of MIT and Harvard, a grant from the Fidelity Bioscience Research Initiative, and the Poitras Center for Affective Disorders Research at the McGovern Institute for Brain Research at MIT.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Molecular Psychiatry website
Rights and permissions
About this article
Cite this article
Lu, C., Chen, Q., Zhou, T. et al. Micro-electrode array recordings reveal reductions in both excitation and inhibition in cultured cortical neuron networks lacking Shank3. Mol Psychiatry 21, 159–168 (2016). https://doi.org/10.1038/mp.2015.173
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/mp.2015.173
This article is cited by
-
Inhibitory regulation of calcium transients in prefrontal dendritic spines is compromised by a nonsense Shank3 mutation
Molecular Psychiatry (2021)
-
SHANK proteins: roles at the synapse and in autism spectrum disorder
Nature Reviews Neuroscience (2017)
-
Modeling psychiatric disorders: from genomic findings to cellular phenotypes
Molecular Psychiatry (2016)
-
A viral strategy for targeting and manipulating interneurons across vertebrate species
Nature Neuroscience (2016)