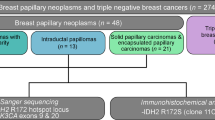
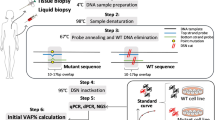
Abstract
The phosphatidylinositol-3-kinase pathway is one of the most commonly mutated pathways in invasive breast carcinoma, with PIK3CA mutations in ∼25% of invasive carcinomas, and AKT1 mutations in up to 5%. Ductal carcinoma in situ, and benign papillomas harbor similar mutations. However, activating point mutations in breast columnar cell lesions have been infrequently studied. Twenty-three breast resection specimens containing columnar cell lesions were identified; 14 with associated invasive carcinoma or carcinoma in situ. DNA extracts were prepared from formalin-fixed paraffin-embedded tissue and screened for a panel of point mutations (321 mutations in 30 genes) using a multiplex PCR panel with mass-spectroscopy readout. PIK3CA mutations were identified in 13/24 columnar cell lesions (54%) and 3/8 invasive carcinomas (37%). The mutation status of columnar cell lesions and associated carcinoma was frequently discordant. Of the 14 cases, only 5 demonstrated the same genotype in matched samples of columnar cell lesions and carcinoma (4 wild type, 1 PIK3CA H1047R). Interestingly, five patients had mutations in columnar cell lesions with wild-type carcinoma; two patients had different point mutations in columnar cell lesions and carcinoma. Only three cases had wild-type columnar cell lesion and mutated carcinoma. The 50% PIK3CA mutation prevalence in columnar cell lesions is greater than reported in most studies of invasive breast cancer. Further, columnar cell lesions and carcinoma were frequently discordant for PIK3CA/AKT1 mutation status. These findings raise interesting questions about the role of PIK3CA/AKT pathway in breast carcinogenesis, and the biologic/precursor potential of columnar cell lesions.
Similar content being viewed by others
Main
The phosphatidylinositol-3-kinase pathway is activated in numerous cancer types and is one of the most commonly mutated pathways in invasive breast carcinoma. Activating mutations in the phosphatidylinositol-3-kinase catalytic subunit (PIK3CA) are present in ∼25% of invasive carcinomas, with mutations clustering in ‘hotspots’ in exon 9 (helical domain) and exon 20 (kinase domain).1, 2, 3, 4, 5, 6, 7, 8, 9, 10 In addition, this pathway is activated by mutations in the plekstrin-homology domain of AKT1 in up to 5% of breast carcinomas, or by the loss of the phosphatase PTEN (phosphatase and tensin homolog) in nearly half of breast cancers.2, 11 Several groups have demonstrated a similar frequency of mutations in breast carcinoma in situ, with paired invasive and in situ carcinoma from the same patient concordant for PIK3CA mutation status in 66–100% of tested cases.12, 13, 14, 15 However, other breast proliferative or putative precursor lesions have been little studied. Li et al13 found PIK3CA hotspot mutations in only 6% of 52 tested cases of Ductal Intraepithelial Neoplasia 1A–B lesions (DIN1A–B, atypical ductal hyperplasia and flat epithelial atypia, also known as columnar cell change with atypia). Our group previously identified PIK3CA/AKT1 mutations in a substantial proportion of benign papillary lesions (65%), neoplasms that are generally not considered to represent precursors of breast cancer.16
Columnar cell lesions of the breast have been recognized historically under a wide variety of different names, and were recently renamed and studied in detail. Columnar cell lesions frequently coexist with atypical hyperplasias and low-grade ductal carcinoma in situ, lobular carcinoma in situ, or invasive breast carcinomas.17, 18, 19, 20 Columnar cell lesions are characterized by variably dilated acini lined by one to several layers of cells with ovoid to elongated nuclei, often with apical snouts (termed columnar cell change or columnar cell hyperplasia). According to the criteria proposed by Schnitt et al,17, 21 columnar cell lesions with cytologic atypia have rounded nuclei, may have nucleoli, and/or cytologic features overlapping low-grade carcinoma; the term flat epithelial atypia encompasses both columnar cell change with atypia, and columnar cell hyperplasia with atypia. The role, if any, of columnar cell lesions along the spectrum of morphologic and pathogenetic progression from normal breast, to in situ or invasive breast carcinoma remains speculative. Recent loss of heterozygosity, comparative genomic hybridization, and immunohistochemical studies have suggested that columnar cell lesions, might represent the ‘missing link’ in low-grade breast cancer progression, as the earliest identifiable non-obligate precursor.18, 22, 23, 24, 25, 26, 27, 28 In order to further address the pathogenetics of columnar cell lesions, we screened a cohort of columnar cell lesions, most with matched normal breast tissue and/or concurrent carcinoma, for a large panel of known activating point mutations, including PIK3CA and AKT1 mutations.
Materials and methods
This project was approved by the Institutional Review Boards of Stanford University and Oregon Health & Science University. Cases of columnar cell lesions were identified from the files of Stanford University Pathology, with two cases from Oregon Health & Science University. Slides were reviewed to identify columnar cell lesions, including columnar cell change, columnar cell hyperplasia, and flat epithelial atypia (columnar cell change with atypia and columnar cell hyperplasia with atypia) using the criteria of Schnitt et al.17, 21 In addition, representative blocks of normal breast tissue, and breast carcinoma from the same specimen/patient, where available, were also selected. Parameters of size, stage, grade, and hormone receptor status for carcinomas were obtained from the pathology reports.
Lesions of interest were marked on slides, and the corresponding area of the formalin-fixed, paraffin-embedded tissue block was isolated by punching the block with 1 or 2 mm core device. Multiple cores of each lesion were collected, some from different tissue blocks, and the cores were re-embedded. Paraffin shavings from the recipient block were collected, and slides were prepared from central depth in the recipient block, in order to allow re-evaluation of the lesional tissue for diagnostic confirmation (RBW and MLT). This method yielded samples comprised of lesional epithelium, surrounding myoepithelium, and stroma. As columnar cell lesions without hyperplasia often line dilated spaces and are associated with hypercellular stroma, the percentage of lesional cells was quite variable. Atypia in columnar cell lesions was assessed, and discrepant cases were resolved by consensus review; controversial or heterogeneous cases were further noted as borderline for atypia.
DNA was extracted from the paraffin shavings using standard protocols (RecoverAll Total Nucleic Acid Isolation Kit #AM1975, Ambion/Applied Biosystems, Austin, TX, USA). DNA extracts were screened for a panel of point mutations using a multiplex PCR panel as previously described.16, 29 In brief, sequences of interest were amplified by PCR; PCR products were then annealed to primers directly adjacent to targeted mutation sites. A primer extension reaction adds one base pair, and the resultant product is analyzed by mass spectroscopy (Sequenom MassARRAY), which allows identification of the added base pair by molecular weight. The mutation panel covers 321 mutations in 30 genes, including ABL, AKT1/2/3, BRAF, CDK4, CTNNB1, EGFR1, ERBB2, FBX4, FBXW7, FGFR1/2/3, FLT3, GNAQ, HRAS, JAK2, KIT, KRAS, MAPK2K1/2, MET, NRAS, PDGFRA, PIK3CA, PTPN11, RET, SOS1, and TP53. The panel includes 41 substitutions in 23 codons of the PIK3CA gene.29 Assays were previously validated in the laboratory using known controls, and 17 of the 20 mutations were independently confirmed by direct sequencing of the sample DNA on an ABI3130 sequencer using the BigDye Terminator method, with or without the use of a locked nucleic acid probe to suppress amplification of the wild-type allele (Ang et al, in preparation). For the remaining cases, sequence reactions were unsuccessful, or insufficient DNA remained. Patient identity across specimens in several of these cases was confirmed by the identification of a common single-nucleotide polymorphism in multiple samples, or by testing a panel of single-nucleotide polymorphisms for an unrelated gene.
Results
Mutational Status of Columnar Cell Lesions, Carcinoma, and Normal Breast Tissue
Twenty-four columnar cell lesions from 22 patients were identified for study, including columnar cell change, columnar cell hyperplasia, and flat epithelial atypia (columnar cell change with atypia). Normal breast tissue from the same specimen was available in 19 patients. Fourteen cases also had concurrent carcinoma, including ductal carcinoma in situ, lobular carcinoma in situ, invasive ductal carcinoma, and metastatic carcinoma (Table 1; Supplementary Information online).
DNA was prepared from formalin-fixed paraffin-embedded tissue, including epithelial lesional tissue and surrounding myoepithelial cells and stroma. DNA extracts were screened for a panel of 321 ‘hotspot’ point mutations in 30 genes; in total, extracts from 65 lesions were analyzed. Point mutations in the PIK3CA gene were identified in 13/24 (54%) columnar cell lesions (patients 1–12; 8 exon 20-H1047R; 3 exon 9-E542K; 1 exon 9-E545K; 1 exon 7-C420R; Table 1; Figures 1 and 2). In one case, columnar cell lesions from different areas of the specimen were analyzed separately, and the same mutation was demonstrated (patient 11L, both E542K; contralateral columnar cell lesion was wild type, 11R). Nine of the columnar cell lesions had atypia (flat epithelial atypia), including five cases with PIK3CA mutations, and four wild-type cases, not different than the mutation frequency of non-atypical lesions (P=0.9156 χ2, not significant). Other than a single-nucleotide polymorphism (MET T992I) in each of the specimens from one patient, no other panel mutations were found in columnar cell lesions.
Histopathology of representative columnar cell lesions. (a) Patient 4, columnar cell hyperplasia, borderline for atypia, with PIK3CA H1047R mutation (see Figure 2). (b) Patient 12, columnar cell hyperplasia without atypia, with PIK3CA C420R mutation (see Figure 2). (c) Patient 7, columnar cell lesion without atypia, with PIK3CA H1047R mutation. (d) Patient 20, columnar cell lesion with atypia, wild type for all mutations tested.
Mutation status across multiple samples from the same patient. (a) Patient 4. Top: Normal breast tissue shows a single wild-type (WT) peak for PIK3CA H1047 by PCR-mass-spectroscopy assay (PCR-MS). Middle: Columnar cell lesion (see Figure 1a) shows a WT and smaller H1047R mutant (*) peak by PCR-MS analysis, and is confirmed by direct sequencing below (CAT → CGT). The relatively small mutant peak is due to sample heterogeneity, including lesional columnar cell epithelium, myoepithelial cells, and stroma. Bottom: Invasive ductal carcinoma demonstrates the same H1047R mutation (*) peak by PCR-MS analysis. In this assay, the primer peak is non-zero, yet the mutation is confirmed by direct sequencing below. (The large peak in the middle of the PCR-MS tracing represent WT peaks from an unrelated, multiplexed assay). (b) Patient 12. Top: Normal breast tissue shows a single WT peak for PIK3CA C420 by PCR-MS assay. Second from top: Columnar cell lesion (see Figure 1b) shows a WT and smaller C420R mutant (*) peak by PCR-MS analysis, and is confirmed by direct sequencing below (CTG → CCG). Second from bottom: Invasive ductal carcinoma is WT for PIK3CA C420R. Bottom: Lymph node metastasis is also WT for PIK3CA C420R. (The large peak in the middle of the PCR-MS results represent WT peaks from an unrelated, multiplexed assay).
In accompanying normal tissue from 19 patients, 18 samples were wild type. One mostly normal sample had an exon 9-E542K mutation (patient 9), like the patient's columnar cell lesion; slide review demonstrated a small amount of contaminating columnar cell lesion (10%).
PIK3CA activating point mutations were identified in 4/8 (50%) of ductal carcinoma in situ tested (1 each exon 20-H1047R; exon 9-E542K; exon 9-E545K; exon 4-N345K; Table 1). Of the eight tested invasive ductal carcinomas, mutations were identified in three cases (3/8=37%), including an AKT1 mutation (AKT1 exon 2-E17K; 2 PIK3CA exon 20-H1047R). The size, grade, stage, and hormone receptor status of the invasive carcinomas are listed in Supplementary Information online. No activating hotspot mutations were identified in three nodal metastases, or two examples of lobular neoplasia (atypical lobular hyperplasia, lobular carcinoma in situ). Again, no other point mutations were identified with this large screening panel.
Mutational Status of Paired Breast Lesions
In 14 specimens, mutation status of columnar cell lesions could be compared with mutation status of concurrent in situ or invasive carcinoma. Interestingly, only five demonstrated the same genotype in matched samples of columnar cell change and carcinoma (5/14=35%, four wild-type patients 20–22, 11R; 1 PIK3CA H1047R-patient 4; Figure 2 and Table 1). Interestingly, five patients had mutations in columnar cell lesion with wild-type ductal carcinoma in situ or invasive carcinoma (patients 5–7 PIK3CA H1047R; patients 11L–12 PIK3CA E542K, PIK3CA C420R; Table 1); two patients had different point mutations in columnar cell lesion and carcinoma (patients 8, 11L; Figure 2 and Table 1). Only three cases had wild-type columnar cell lesion and mutated carcinoma (patients 13–15; Table 1).
Of the few pairs of ductal carcinoma in situ and invasive carcinoma tested, two of the three were discordant for mutation status (patients 11L and 14, both of these patients had different mutations in each of the several different lesions). Of the three wild-type metastatic carcinomas, two were associated with wild-type invasive carcinoma, while one was associated with ductal carcinoma in situ harboring the PIK3CA exon 9 E545K mutation (patient 8), without invasive carcinoma available for analysis.
Discussion
The PIK3CA mutation prevalence of 25–30% in invasive as well as in situ breast carcinoma is fairly well established in the literature, and is perhaps slightly higher in estrogen receptor-positive subgroups.12, 13, 14, 15, 30, 31, 32, 33, 34, 35, 36, 37, 38, 39, 40, 41, 42, 43 However, the mutation prevalence in breast columnar cell lesions, as well as the evolution of activating point mutations across different breast lesions in a single patient, has not been well studied to date. In a cohort of 24 columnar cell lesions, encompassing columnar cell change, columnar cell hyperplasia, and flat epithelial atypia, we found PIK3CA mutations in 13 samples. In addition, we found little concordance between mutation status of columnar cell lesions and accompanying carcinoma.
The 54% PIK3CA mutation frequency in columnar cell lesions in our study is higher than the mutation frequency of invasive carcinoma, both in our study (3/8, 37%), and higher than reported in the literature, including recent studies using very sensitive techniques, similar to those employed in this study.14, 31, 32, 44, 45, 46 However, our group previously identified PIK3CA or AKT1 mutations in 65% of benign or atypical breast papillary lesions,16 suggesting that a variety of proliferative lesions of the breast may harbor a high frequency of PIK3CA pathway mutations. In contrast, Li et al13 studied a spectrum of proliferative and atypical breast lesions and found relatively fewer mutations. They screened usual ductal hyperplasia, Ductal Intraepithelial Neoplasia 1A (flat epithelial atypia), and Ductal Intraepithelial Neoplasia 1B (atypical ductal hyperplasia), as well as ductal carcinoma in situ and invasive carcinoma using direct sequencing, with laser capture microdissection. They found mutations in only 1/20 (5%) cases of Ductal Intraepithelial Neoplasia 1A and 2/32 (6%) cases of Ductal Intraepithelial Neoplasia 1B, and none in 16 cases of usual ductal hyperplasia.13 Although the frequency of mutations identified by Li et al13 in ductal carcinoma in situ and invasive carcinoma was comparable to other reports in the literature (24.5 and 26.8%, respectively), direct sequencing is known to have lower sensitivity for small samples, as compared with Sequenom mass-array, SNaPshot PCR, and other methods.15, 45, 47, 48
Progression from pre-neoplastic lesions to in situ to invasive carcinoma has been associated with accumulation of genetic changes, as is well established in the progression of colonic adenomas to colon carcinoma.49, 50 In breast pathology, comparative genomic hybridization and a number of other techniques have substantiated the known precursor status of ductal carcinoma in situ.23, 51, 52 These techniques are now being employed to investigate columnar cell change and other proliferative breast lesions as putative precursors to breast carcinoma.22, 23, 24, 25, 26, 27, 28 Previously, Moinfar et al22 analyzed loss of heterozygosity in Ductal Intraepithelial Neoplasia-flat lesions, and found significant alterations, including changes in common with concurrent carcinoma; however, 9 of the 22 studied cases were ‘pleomorphic’ Ductal Intraepithelial Neoplasia-flat, equivalent to clinging carcinoma (high nuclear grade flat ductal carcinoma in situ). In a subsequent study using array comparative genomic hybridization Moinfar's group showed by that Ductal Intraepithelial Neoplasia low grade harbored changes common to lobular neoplasia, low nuclear grade ductal carcinoma in situ, and invasive carcinoma (loss of 16q, gain of 1q).28 Simpson et al24 found chromosomal alterations, as analyzed by comparative genomic hybridization, across the spectrum of columnar cell lesions (including some with architectural complexity that would alternatively be classified as atypical ductal hyperplasia by some experts). They noted a greater number of changes in lesions with cytologic atypia, hyperplasia, and/or architectural complexity. Further, they studied a large number of lesions from three patients and found recurring genetic changes in lesions of different severity, ranging from columnar cell change to low-grade ductal carcinoma in situ in two of three patients. The authors suggested that columnar cell lesions might be an important non-obligate precursor of breast carcinoma, even the ‘missing link.’18, 24 In further support of that hypothesis, Dabbs et al25 evaluated loss of heterozygosity of 10 microsatellite markers, and also demonstrated an accumulation of alterations across the spectrum from columnar cell lesions, to atypical ductal hyperplasia, to ductal carcinoma in situ and invasive carcinoma. Recently, Ellsworth et al27 studied allelic imbalance (loss of heterozygosity) in columnar cell lesions and atypical ductal hyperplasia that were not associated with carcinoma. They found levels of genomic alterations intermediate between normal breast and invasive/in situ carcinoma, importantly without recurrent alterations and 16q and 17q. They concluded that these molecular profiles do not support a role for columnar cell lesions and atypical ductal hyperplasia as obligate precursors.27
Similar to those previous studies, we found a high percentage of genetic alterations in the form of activating PIK3CA mutations in breast columnar cell lesions. However, we found a very low concordance with mutational status of accompanying carcinoma. In fact, only 5/14 cases were concordant, and four of these cases were wild type. More commonly, we found that mutated columnar cell lesion was accompanied by wild-type carcinoma, or carcinoma with a different activating point mutation (9/14). This combination of genotypes does not follow the canonical genetic stepwise progression model from precursor with few key early mutations, to carcinoma after accumulation of additional deleterious mutations. Certainly, these data remain compatible with the hypothesis that columnar cell lesions might be non-obligate or heterogeneous precursors, such that only a small fraction of columnar cell lesions might progress to carcinoma. This and several alternative hypotheses deserve further study, including columnar cell lesions as a marker of susceptibility, columnar cell lesions as incidental unrelated proliferations without precursor potential, etc.
PIK3CA or AKT1 mutations are thought to have key roles in the biology of breast cancer, in that PIK3CA/AKT pathway activation may account for resistance to trastuzumab or small molecule tyrosine kinase inhibitors in some Her2-positive tumors.32, 53, 54, 55, 56 Further, in estrogen receptor-positive breast cancers, a PIK3CA mutation-gene signature has been elucidated by expression analysis, paradoxically imparting a relatively favorable prognosis.39, 40, 43, 44, 57 Nevertheless, whether PIK3CA/AKT1 mutations have a role in breast carcinogenesis, perhaps as an early primary or ‘driver’ mutation remains unclear.50 The high frequency of PIK3CA/AKT1 mutations in columnar cell lesions and benign breast papillomas suggests that PIK3CA/AKT1 mutations may be related to the development of breast proliferative lesions. If columnar cell lesions represent non-obligate precursors of in situ and invasive breast carcinoma, our data suggest the possibility that PIK3CA/AKT1 mutations may not necessarily confer increased risk of progression, as the PIK3CA/AKT1 mutation status of columnar cell lesions and concurrent carcinoma were frequently discordant. Thus, it appears that mutant clones may be bypassed, selected against, or mutations may be lost, during progression to carcinoma, allowing speculation that PIK3CA/AKT1 mutations might be less beneficial in later stage lesions. Similarly, an elegant repressible mouse model system has shown that mammary tumors induced by overexpression of PIK3CA H1047R frequently recur independent of the PIK3CA mutant transgene, with many recurrent tumors harboring additional genomic alterations, including Met or myc perturbations, among others.58
In summary, we have demonstrated a high prevalence of PIK3CA mutations in breast columnar cell lesions, as well as a lack of genotypic concordance of columnar cell lesions with paired carcinoma. Although these findings need validation in a larger study, they raise a number of interesting questions as to the biologic/precursor potential of breast columnar cell lesions, and the role of PIK3CA/AKT1 mutations in breast carcinogenesis.
References
Karakas B, Bachman KE, Park BH . Mutation of the PIK3CA oncogene in human cancers. Br J Cancer 2006;94:455–459.
Castaneda CA, Cortes-Funes H, Gomez HL, et al. The phosphatidyl inositol 3-kinase/AKT signaling pathway in breast cancer. Cancer Metastasis Rev 2010;29:751–759.
Zhao L, Vogt PK . Class I PI3K in oncogenic cellular transformation. Oncogene 2008;27:5486–5496.
Engelman JA, Luo J, Cantley LC . The evolution of phosphatidylinositol 3-kinases as regulators of growth and metabolism. Nat Rev Genet 2006;7:606–619.
Wymann MP, Marone R . Phosphoinositide 3-kinase in disease: timing, location, and scaffolding. Curr Opin Cell Biol 2005;17:141–149.
McAuliffe PF, Meric-Bernstam F, Mills GB, et al. Deciphering the role of PI3K/Akt/mTOR pathway in breast cancer biology and pathogenesis. Clin Breast Cancer 2010;10 (Suppl 3):S59–S65.
Engelman JA . Targeting PI3K signalling in cancer: opportunities, challenges and limitations. Nat Rev Cancer 2009;9:550–562.
Samuels Y, Wang Z, Bardelli A, et al. High frequency of mutations of the PIK3CA gene in human cancers. Science 2004;304:554.
Bachman KE, Argani P, Samuels Y, et al. The PIK3CA gene is mutated with high frequency in human breast cancers. Cancer Biol Ther 2004;3:772–775.
Troxell ML . PIK3CA/AKT1 mutations in breast carcinoma: a comprehensive review of experimental and clinical studies. J Clin Exp Pathol 2012; doi: 10.4172/2161-0681.S1-002.
Carpten JD, Faber AL, Horn C, et al. A transforming mutation in the pleckstrin homology domain of AKT1 in cancer. Nature 2007;448:439–444.
Dunlap J, Le C, Shukla A, et al. Phosphatidylinositol-3-kinase and AKT1 mutations occur early in breast carcinoma. Breast Cancer Res Treat 2010;120:409–418.
Li H, Zhu R, Wang L, et al. PIK3CA mutations mostly begin to develop in ductal carcinoma of the breast. Exp Mol Pathol 2010;88:150–155.
Miron A, Varadi M, Carrasco D, et al. PIK3CA mutations in in situ and invasive breast carcinomas. Cancer Res 2010;70:5674–5678.
Kalinsky K, Heguy A, Bhanot UK, et al. PIK3CA mutations rarely demonstrate genotypic intratumoral heterogeneity and are selected for in breast cancer progression. Breast Cancer Res Treat 2011;129:635–643.
Troxell ML, Levine J, Beadling C, et al. High prevalence of PIK3CA/AKT pathway mutations in papillary neoplasms of the breast. Mod Pathol 2010;23:27–37.
Schnitt SJ, Vincent-Salomon A . Columnar cell lesions of the breast. Adv Anat Pathol 2003;10:113–124.
Abdel-Fatah TM, Powe DG, Hodi Z, et al. High frequency of coexistence of columnar cell lesions, lobular neoplasia, and low grade ductal carcinoma in situ with invasive tubular carcinoma and invasive lobular carcinoma. Am J Surg Pathol 2007;31:417–426.
Moinfar F . Flat ductal intraepithelial neoplasia of the breast: a review of diagnostic criteria, differential diagnoses, molecular-genetic findings, and clinical relevance--it is time to appreciate the Azzopardi concept!. Arch Pathol Lab Med 2009;133:879–892.
Lerwill MF . Flat epithelial atypia of the breast. Arch Pathol Lab Med 2008;132:615–621.
Tavassoli F, Hoefler H, Rosai J, et al. Flat epithelial atypia. In: Tavassoli F, Devilee P (eds). Tumous of the Breast and Female Genital Organs Vol. IARC Press: Lyon, 2003, pp 65–66.
Moinfar F, Man YG, Bratthauer GL, et al. Genetic abnormalities in mammary ductal intraepithelial neoplasia-flat type (″clinging ductal carcinoma in situ″): a simulator of normal mammary epithelium. Cancer 2000;88:2072–2081.
Reis-Filho JS, Lakhani SR . The diagnosis and management of pre-invasive breast disease: genetic alterations in pre-invasive lesions. Breast Cancer Res 2003;5:313–319.
Simpson PT, Gale T, Reis-Filho JS, et al. Columnar cell lesions of the breast: the missing link in breast cancer progression? A morphological and molecular analysis. Am J Surg Pathol 2005;29:734–746.
Dabbs DJ, Carter G, Fudge M, et al. Molecular alterations in columnar cell lesions of the breast. Mod Pathol 2006;19:344–349.
Abdel-Fatah TM, Powe DG, Hodi Z, et al. Morphologic and molecular evolutionary pathways of low nuclear grade invasive breast cancers and their putative precursor lesions: further evidence to support the concept of low nuclear grade breast neoplasia family. Am J Surg Pathol 2008;32:513–523.
Ellsworth RE, Ellsworth DL, Weyandt JD, et al. Chromosomal alterations in pure nonneoplastic breast lesions: implications for breast cancer progression. Ann Surg Oncol 2010;17:1688–1694.
Stacher E, Boldt V, Leibl S, et al. Chromosomal aberrations as detected by array comparative genomic hybridization in early low-grade intraepithelial neoplasias of the breast. Histopathology 2011;59:549–555.
Beadling C, Heinrich MC, Warrick A, et al. Multiplex mutation screening by mass spectrometry evaluation of 820 cases from a personalized cancer medicine registry. J Mol Diagn 2011;13:504–513.
Campbell IG, Russell SE, Choong DY, et al. Mutation of the PIK3CA gene in ovarian and breast cancer. Cancer Res 2004;64:7678–7681.
Li SY, Rong M, Grieu F, et al. PIK3CA mutations in breast cancer are associated with poor outcome. Breast Cancer Res Treat 2006;96:91–95.
Stemke-Hale K, Gonzalez-Angulo AM, Lluch A, et al. An integrative genomic and proteomic analysis of PIK3CA, PTEN, and AKT mutations in breast cancer. Cancer Res 2008;68:6084–6091.
Hennessy BT, Gonzalez-Angulo AM, Stemke-Hale K, et al. Characterization of a naturally occurring breast cancer subset enriched in epithelial-to-mesenchymal transition and stem cell characteristics. Cancer Res 2009;69:4116–4124.
Buttitta F, Felicioni L, Barassi F, et al. PIK3CA mutation and histological type in breast carcinoma: high frequency of mutations in lobular carcinoma. J Pathol 2006;208:350–355.
Liang X, Lau QC, Salto-Tellez M, et al. Mutational hotspot in exon 20 of PIK3CA in breast cancer among Singapore Chinese. Cancer Biol Ther 2006;5:544–548.
Maruyama N, Miyoshi Y, Taguchi T, et al. Clinicopathologic analysis of breast cancers with PIK3CA mutations in Japanese women. Clin Cancer Res 2007;13:408–414.
Perez-Tenorio G, Alkhori L, Olsson B, et al. PIK3CA mutations and PTEN loss correlate with similar prognostic factors and are not mutually exclusive in breast cancer. Clin Cancer Res 2007;13:3577–3584.
Benvenuti S, Frattini M, Arena S, et al. PIK3CA cancer mutations display gender and tissue specificity patterns. Hum Mutat 2008;29:284–288.
Loi S, Haibe-Kains B, Majjaj S, et al. PIK3CA mutations associated with gene signature of low mTORC1 signaling and better outcomes in estrogen receptor-positive breast cancer. Proc Natl Acad Sci USA 2010;107:10208–10213.
Ellis MJ, Lin L, Crowder R, et al. Phosphatidyl-inositol-3-kinase alpha catalytic subunit mutation and response to neoadjuvant endocrine therapy for estrogen receptor positive breast cancer. Breast Cancer Res Treat 2010;119:379–390.
Bozhanov SS, Angelova SG, Krasteva ME, et al. Alterations in p53, BRCA1, ATM, PIK3CA, and HER2 genes and their effect in modifying clinicopathological characteristics and overall survival of Bulgarian patients with breast cancer. J Cancer Res Clin Oncol 2010;136:1657–1669.
Boyault S, Drouet Y, Navarro C, et al. Mutational characterization of individual breast tumors: TP53 and PI3K pathway genes are frequently and distinctively mutated in different subtypes. Breast Cancer Res Treat 2011;132:29–39.
Cizkova M, Cizeron-Clairac G, Vacher S, et al. Gene expression profiling reveals new aspects of PIK3CA mutation in ERalpha-positive breast cancer: major implication of the Wnt signaling pathway. PLoS One 2010;5:e15647.
Kalinsky K, Jacks LM, Heguy A, et al. PIK3CA mutation associates with improved outcome in breast cancer. Clin Cancer Res 2009;15:5049–5059.
Dupont Jensen J, Laenkholm AV, Knoop A, et al. PIK3CA mutations may be discordant between primary and corresponding metastatic disease in breast cancer. Clin Cancer Res 2011;17:667–677.
Vorkas PA, Poumpouridou N, Agelaki S, et al. PIK3CA hotspot mutation scanning by a novel and highly sensitive high-resolution small amplicon melting analysis method. J Mol Diagn 2010;12:697–704.
Arcila ME, Oxnard GR, Nafa K, et al. Rebiopsy of lung cancer patients with acquired resistance to EGFR inhibitors and enhanced detection of the T790M mutation using a locked nucleic acid-based assay. Clin Cancer Res 2011;17:1169–1180.
Gonzalez-Angulo AM, Ferrer-Lozano J, Stemke-Hale K, et al. PI3K pathway mutations and PTEN levels in primary and metastatic breast cancer. Mol Cancer Ther 2011;10:1093–1101.
Vogelstein B, Fearon ER, Hamilton SR, et al. Genetic alterations during colorectal-tumor development. N Engl J Med 1988;319:525–532.
Wood LD, Parsons DW, Jones S, et al. The genomic landscapes of human breast and colorectal cancers. Science 2007;318:1108–1113.
Ma XJ, Salunga R, Tuggle JT, et al. Gene expression profiles of human breast cancer progression. Proc Natl Acad Sci USA 2003;100:5974–5979.
Arpino G, Laucirica R, Elledge RM . Premalignant and in situ breast disease: biology and clinical implications. Ann Intern Med 2005;143:446–457.
Berns K, Horlings HM, Hennessy BT, et al. A functional genetic approach identifies the PI3K pathway as a major determinant of trastuzumab resistance in breast cancer. Cancer Cell 2007;12:395–402.
Park BH, Davidson NE . PI3 kinase activation and response to Trastuzumab Therapy: what′s neu with herceptin resistance? Cancer Cell 2007;12:297–299.
Nahta R, Yu D, Hung MC, et al. Mechanisms of disease: understanding resistance to HER2-targeted therapy in human breast cancer. Nat Clin Pract Oncol 2006;3:269–280.
O′Brien NA, Browne BC, Chow L, et al. Activated phosphoinositide 3-kinase/AKT signaling confers resistance to trastuzumab but not lapatinib. Mol Cancer Ther 2010;9:1489–1502.
Di Cosimo S, Baselga J . Phosphoinositide 3-kinase mutations in breast cancer: a ″good″ activating mutation? Clin Cancer Res 2009;15:5017–5019.
Liu P, Cheng H, Santiago S, et al. Oncogenic PIK3CA-driven mammary tumors frequently recur via PI3K pathway-dependent and PI3K pathway-independent mechanisms. Nat Med 2011;17:1116–1120.
Acknowledgements
We acknowledge the excellent administrative assistance of Eileen Maisen, Erin Popelka, and Tabatha Pulliam, and computer technical assistance of Alex Bolinder (Figure 2). This research was supported by a grant from Susan G Komen for the Cure, KG100112 (MLT), and from the California Breast Cancer Research Program, 17IB-0038 (RBW).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
This project was presented in abstract form at the San Antonio Breast Cancer Symposium, 8 December 2011.
Supplementary Information accompanies the paper on Modern Pathology website
Supplementary information
Rights and permissions
About this article
Cite this article
Troxell, M., Brunner, A., Neff, T. et al. Phosphatidylinositol-3-kinase pathway mutations are common in breast columnar cell lesions. Mod Pathol 25, 930–937 (2012). https://doi.org/10.1038/modpathol.2012.55
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2012.55
Keywords
This article is cited by
-
Machine learning-based image analysis for accelerating the diagnosis of complicated preneoplastic and neoplastic ductal lesions in breast biopsy tissues
Breast Cancer Research and Treatment (2021)
-
AKT1 E17K mutation profiling in breast cancer: prevalence, concurrent oncogenic alterations, and blood-based detection
BMC Cancer (2016)
-
Appraisal of the technologies and review of the genomic landscape of ductal carcinoma in situ of the breast
Breast Cancer Research (2015)
-
Cell-lineage heterogeneity and driver mutation recurrence in pre-invasive breast neoplasia
Genome Medicine (2015)
-
Chromosomal copy number alterations for associations of ductal carcinoma in situ with invasive breast cancer
Breast Cancer Research (2015)