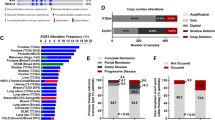
Abstract
6q12-22 is the second most commonly deleted genomic region in prostate cancer. Mapping studies have described a minimally deleted area at 6q15, containing MAP3K7/TAK1, which was recently shown to have tumor suppressive properties. To determine prevalence and clinical significance of MAP3K7 alterations in prostate cancer, a tissue microarray containing 4699 prostate cancer samples was analyzed by fluorescence in situ hybridization. Heterozygous MAP3K7 deletions were found in 18.48% of 2289 interpretable prostate cancers. MAP3K7 deletions were significantly associated with advanced tumor stage (P<0.0001), high Gleason grade (P<0.0001), lymph node metastasis (P<0.0108) and early biochemical recurrence (P<0.0001). MAP3K7 alterations were typically limited to the loss of one allele as homozygous deletions were virtually absent and sequencing analyses revealed no evidence for MAP3K7 mutations in 15 deleted and in 14 non-deleted cancers. There was a striking inverse association of MAP3K7 deletions and TMPRSS2:ERG fusion status with 26.7% 6q deletions in 1125 ERG-negative and 11.1% 6q deletions in 1198 ERG-positive cancers (P<0.0001). However, the strong prognostic role of 6q deletions was retained in both ERG-positive and ERG-negative cancers (P<0.0001 each). In summary, our study identifies MAP3K7 deletion as a prominent feature in ERG-negative prostate cancer with strong association to tumor aggressiveness. MAP3K7 alterations are typically limited to one allele of the gene. Together with the demonstrated tumor suppressive function in cell line experiments and lacking evidence for inactivation through hypermethylation, these results indicate MAP3K7 as a gene for which haploinsufficency is substantially tumorigenic.
Similar content being viewed by others
Main
In prostate cancer a variety of chromosomal deletions occur frequently, whereas gains of chromosomal material and high-level amplification occur rarely in this tumor.1, 2 Deletions of 6q12-22 have been described to occur in 22–62% of prostate cancers.1, 2, 3, 4, 5 6q12-22 deletions rank second in the list of frequent prostate cancer deletions after 8p11-21.1, 2
Published studies mapping the 6q12-22 deletion suggest a small commonly deleted region on 6q15 with a length of 3–4 megabases (Mb). This area contains several genes with a known or suspected tumor suppressing function including MAP3K7, for which a tumor suppressive role for prostate epithelial cells has recently been demonstrated.6 MAP3K7 is ubiquitary expressed and involved in various biological processes such as cell growth, differentiation and apoptosis. MAP3K7 exerts these effects by interacting with a number of different signaling pathways and activator molecules.7, 8, 9, 10 For example, MAP3K7 can be activated by TGF-β and BMP.8, 11, 12 MAP3K7 is one of the first kinases in the p38 and JNK signaling pathways.8, 13, 14 MAP3K7 also has an important role in the balance between the BMP- and WNT-signaling pathways.7, 15
The clinical effects of MAP3K7 alterations are unclear. Some studies associated this alteration with unfavorable phenotype,4 metastasis16 or decreased tumor recurrence.3, 5 Others could not find clinically relevant associations or described associations with more benign tumor features.5, 17 These studies included patient cohorts of 24–95 cases. To identify possible clinico-pathological associations of MAP3K7 deletions in prostate cancer, we analyzed a prostate cancer tissue microarray containing samples of 4699 patients. We also performed sequencing analyses and functional studies, to learn more on possible other mechanisms of MAP3K7 inactivation and its consequences. The data pinpoint MAP3K7 as a tumor suppressor gene with substantial clinical relevance in prostate cancer.
Materials and methods
Patients
A set of prostate cancer tissue microarrays was used in this study containing one tissue core each from 4699 consecutive radical prostatectomy specimens from patients undergoing surgery between 1992 and 2008 at the Department of Urology, and the Martini Clinics at the University Medical Center Hamburg-Eppendorf. During this time span, all prostatatectomy specimens were completely embedded and sectioned according to a standardized protocol.18 The tissue microarray manufacturing process was described earlier in detail.19, 20 In short, one 0.6 mm core was taken from a representative tissue block from each patient. The tissues were distributed among 10 tissue microarray blocks, each containing 129–522 cores. The clinical and pathological data associated with these patients are given in Supplementary Table 1. Clinical follow-up data were available for 4203 of the 4699 arrayed tumors. Median follow-up was 46.7 months ranging from 1 to 219 months. None of the patients received neo-adjuvant endocrine therapy. Additional (salvage) therapy was only initiated after biochemical relapse, the clinical end point of this study. Prostate-specific antigen (PSA) values were measured following surgery and recurrence was defined as a postoperative PSA of 0.2 ng/ml and increasing at first of appearance. Immunohistochemical data on ERG and Ki67 were available from previous studies.21, 22
Fluorescence In Situ Hybridization
Four micron tissue microarray sections were used for dual color fluorescence in situ hybridization (FISH). Before hybridization, sections were deparaffinized and proteolytically pretreated with a commercial kit (paraffin pretreatment reagent kit; Abbott Molecular, Wiesbaden, Germany), followed by dehydration in 70, 80 and 96% ethanol, air drying and denaturation for 10 min at 72 °C in 70% formamide-2 × SSC solution. For MAP3K7 deletion analysis, a dual color FISH probe set was assembled. For MAP3K7 detection, two contiguous BAC probes (BAC RP3-470J8, BAC RP11-501P02; Imergenes, Germany) containing the MAP3K7 gene and an adjacent up- and downstream region measuring 338 kilobases (Kb) in total were spectrum green-labeled. A commercial spectrum orange-labeled centromere 6 probe (#06J36-06; Abbott) was used as a reference. Hybridization was done over night at 37 °C in a humidified chamber, slides were washed and counterstained with 0.2 μmol/L 4′-6-diamidino-2-phenylindole in an antifade solution. For MAP3K7 deletion analysis, each tissue spot was carefully evaluated and the predominant signal counts were recorded for each FISH probe. Tissue samples were excluded from the analysis if a careful morphological and immunohistochemical (basal cell marker 34βE12; AMACR) suggested absence of clear-cut tumor cells in adjacent tissue microarray sections. Information from this section was also used to identify small cancer areas on tissue microarray spots. Tissue spots were also excluded from analysis if there was evidence for insufficient hybridization such as lack of MAP3K7 signals in all (tumor and normal) cells or lack of MAP3K7 signals in all tumor cells but absence of normal cells containing unequivocal MAP3K7 signals. Heterozygous deletion of MAP3K7 was defined as presence of fewer MAP3K7 signals than centromere 6 probe signals in >50% tumor nuclei. This threshold was selected because a good concordance was achieved between deletions identified in array CGH and by FISH using this approach in earlier studies analyzing PTEN.22 Representative cases with and without MAP3K7 deletion are shown in Figure 1.
MAP3K7 Mutational Analysis
Tissue specimens from 30 patients were selected if at least 70% tumor cells were present in the tumor area of the paraffin tissue block. Two punches (0.6 mm in diameter, 5 mm long) were taken from tissue blocks in one large tumor-containing area using a hollow needle. Paraffin was removed with xylene and absolute ethanol. After digestion with proteinase K at 58 °C over night, DNA was isolated using a commercial kit (QIAmp DNA FFPE Kit, Qiagen, Hilden, Germany). All 17 MAP3K7 exons were amplified by PCR using the AmpliTaqGold polymerase (Applied Biosystems, Darmstadt, Germany). Primer sequences are given in Supplementary Table S2. PCR cycling conditions included an initial denaturation step at 95 °C for 10 min, followed by 40 cycles of 95 °C denaturation for 20 s, 58 °C or 44 °C (exon 3) annealing for 30 s or 20 s (exon 3) and 72°C extension for 40 s, and a final extension step at 72 °C for 7 min. The quality of PCR products was confirmed by capillary electrophoresis on a QIAxelsystem (Qiagen). Sequencing was done using the Big Dye Terminator Kit (Applied Biosystems) and electrophoretic analysis on the Genetic Analyzer 3100 (Applied Biosystems). Sequencing primers are given in Supplementary Table S3. Sequence variations were always validated in DNA derived from a second, independent PCR amplification.
SNP Array Analysis
A total of 72 snap-frozen prostate cancer samples with at least 70% tumor cell content and 5 prostate cancer cell lines (LNCaP, PC-3, BPH-1, DU-145, VCaP) were selected for SNP array analysis. DNA was isolated using a commercial kit (QIAamp DNA Mini Kit, Qiagen). Affymetrix SNP V6.0 arrays were used for copy number analysis. Fragmentation, labeling and hybridization of the DNA to the SNP arrays were carried out exactly as described in the Affymetrix V6.0 SNP array manual. A home-made genomic browser (FISH Oracle) was used to map all 6q12-22 deletions to the human genome reference sequence (Archive EnsEMBL release 54 – May 2009) and to define the minimally overlapping region of deletion.23
Cell Culture, Constructs and Lentivirus Production
The LNCaP, BPH-1, PC-3, DU-145, RWPE-1 and VCaP prostate cell lines were obtained and cultured according to the supplier’s instructions (ATCC and Sigma). All cell lines have been periodically tested (every 3 months) for cell morphology, growth rate and gene expression. Following expression constructs were used: pWZL-Neo-Myr-Flag-MAP3K7,24 PSG5L-HA-RB1, PSG5L-HA-PTEN, pLNGY. For depletion experiments, shRNA-expressing vectors based on lentiviral pLKO.1 construct and part of the RNAi Consortium (TRC) vector collection were purchased from Sigma-Aldrich. The shRNAs used included the mature sense sequences for MAP3K7: 5′-CAGTGTGTCTTGTGATGGAAT-3′, PTEN: 5′-CCACAGCTAGAACTTATCAAA-3′, RB1: 5′-GTGCGCTCTTGAGGTTGTAAT-3′ and green fluorescent protein (GFP) 5′-CAACAAGATGAAGAGCACCAA-3′. Lentivirus supernatants were prepared after co-transfection into HEK-293T cells of a lentivirus vector plasmid with pVSV-G (expressing the VSV envelope protein) pREV and pRRE and (expressing lentivirus helper functions), as described previously.25 Prostate cancer cells were transduced with lentiviruses expressing shRNAs directed against either luciferase or GFP (as a control) or MAP3K7. Transduced target cells were selected with puromycin (1.5 μg/ml).
Western Blot Analysis
Cells were collected in SDS-PAGE loading buffer. Proteins were resolved on SDS gels, and then transferred to nitrocellulose membranes by western blotting. The following antibodies were used: anti-a-tubulin (Sigma-Aldrich, Germany), anti-MAP3K7 (Sigma-Aldrich).
RNA-Isolation and Taqman PCR
Total RNA was extracted using Trizol and RNeasy system (Macherey-Nagel, Germany). RNA was reverse transcribed using the High Capacity cDNA Archive Kit (Applied Biosystems). Real-time reverse transcriptase-PCR (RT-PCR) was performed as described previously.26 For all other genes Assay-on-Demand primer/probe sets supplied by Applied Biosystems were used (Assay IDs are available upon request). Relative expression was calculated by normalization to a selected housekeeper mRNA (GAPDH) by the ΔΔCt method.27
Colony Formation Assay
BPH-1 and DU-145 prostate cells were plated at about 2 × 105 in six-well plates. Cells were transfected with 4 μg of indicated cDNAs using Lipofectamine 2000 (Life Technologies, NY, USA). Thirty-six hours after transfection cells were cultured in medium containing puromycin (1.5 μg/ml). Medium was replaced every 2–3 days with fresh medium containing the selection drug. Drug-resistant colonies appearing about 2 weeks later were fixed with methanol, stained with Giemsa and counted.
Statistics
For statistical analysis, the JMP 8.0 software (SAS Institute, NC, USA) was used. Contingency tables were calculated to study association between MAP3K7 deletion and clinico-pathological variables, and the χ2(Likelihood) test was used to find significant relationships. Kaplan–Meier curves were generated for PSA recurrence-free survival. The log-Rank test was applied to test the significance of differences between stratified survival functions. Cox proportional hazards regression analysis was performed to test the statistical independence and significance between pathological, molecular and clinical variables.
Results
Technical Issue
A total of 2289 of 4699 (48.7%) prostate cancer probes were successfully analyzed in this study. Analysis failed in 2410 tumors either because of lack of tissue spots in the tissue microarray section, absence of unequivocal cancer cells or because of faint or lacking FISH signals for centromere 6, MAP3K7 or both.
MAP3K7 Deletions
MAP3K7 deletions were found in 18.5% (423 of 2289) of all prostate cancers (Figure 1). All of these were heterozygous MAP3K7 deletions. Homozygous MAP3K7 deletions were not found. The relationship between MAP3K7 deletions and tumor phenotype and clinical parameters is summarized in Table 1. MAP3K7 deletions were significantly linked to advanced tumor stage (P<0.0001), high Gleason grade (P<0.0001), high preoperative PSA serum level (P=0.0017) and presence of lymph node metastasis (P=0.0108). However, presence of MAP3K7 deletion was unrelated to the surgical margin status (P=0.13). MAP3K7 deletions were also significantly associated with early PSA recurrence (Figure 2). However, multivariate analysis including Gleason grade, pT category and surgical margins did not suggest MAP3K7 as an independent prognostic factor (Table 2).
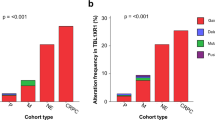
(a) Association between MAP3K7 deletion and biochemical recurrence in prostate cancer. (b)Association of MAP3K7 deletion status to ERG fusion status (P<0001). Association between MAP3K7 deletion in ERG fusion-negative (c) and -positive (d) tumors and biochemical recurrence in prostate cancer. Normal: two MAP3K7 gene copies, Heterozygous: heterozygous deletion with one MAP3K7 gene copy.
Association of MAP3K7 to Other Molecular Markers
Results of MAP3K7 and ERG expression status could be compared in 2323 cancer samples. MAP3K7 deletion was strongly associated with ERG fusion negative tumors (26.7% vs 11.1%, P<0.0001). The prognostic relevance of MAP3K7 deletions hold for the ERG-positive and ERG-negative subgroups (Figure 2). Data on MAP3K7 status and Ki67 labeling index (Ki67Li) were available from 1918 prostate cancers. Ki67 LI was significantly higher in MAP3K7 deleted (average Ki67LI 6.95) than in undeleted cancers (av. Ki67LI 5.23, P<0.0001) if all cancers were jointly analyzed. In a separate analysis of tumors with identical Gleason grade, this association remained statistically significant in tumors with Gleason grade 3+4 and 4+3, but not in Gleason 3+3 and ≥4+4 cancers (Supplementary table S4).
Mutation Analysis of MAP3K7
All 17 exons of MAP3K7 were successfully analyzed in 29 out of selected 30 prostate cancers including 15 MAP3K7 deleted and 14 non-deleted prostate cancers (validated by FISH). Mutations of MAP3K7 were not found in any of these cases.
SNP Array Analysis
Deletions involving 6q were found in 22 of the 77 analyzed prostate cancer samples. The largest deletion spanned more than 43 Mb. The smallest common area of deletion marked a region of 2.1 Mb that were commonly deleted in 21 of 22 tumors with 6q15 deletions. This region contains the MAP3K7 gene and 11 other genes (Figure 3).
Genomic 6q15 deletions detected in 22/77 prostate cancer samples in SNP-array analysis. Left part shows an overview of position and size of genomic deletion relative to 6q region in 22 prostate cancers. Right, detailed view of genomic deleted region relative to 6q15 region. The position of BAC clones (RP3-470J8, RP11-501P02) relative to the MAP3K7 locus is indicated.
Expression of MAP3K7 and its Tumor Suppressor Function In Vitro
SNP array analysis had revealed a 6q15 deletion spanning 4.3 Mb only in LNCaP, whereas BPH-1 and DU-145 had normal 6q15 copy numbers. Accordingly, mRNA expression analysis by RT-PCR had revealed a reduced expression of all analyzed genes involved in the 6q15 deletion including MAP3K7 in LNCaP cells as compared with BPH-1 (Supplementary Figure S1). To study the effects of both shRNA-mediated MAP3K7 downregulation and ectopic overexpression on cell growth colony formation assay experiments were performed. The extent of MAP3K7 downregulation by Lentivirus-mediated gene knockdown was validated both on the mRNA and protein level. Known essential (PTEN) and non-essential (RB1) tumor suppressor genes were included for control. Overexpression of MAP3K7 resulted in strong growth inhibition in BPH-1 and DU-145 cells comparable to the effects seen with both PTEN and RB1 (Figure 4a,c). Knockdown of MAP3K7 did not significantly interfere with cell growth (Figure 4b, d). Taken together, these data are consistent with a role of MAP3K7 as a tumor suppressor in prostate cancer.
(a, c) Effect of MAP3K7 overexpression on cell growth. DU-145 (a) and BPH-1 (c) cells were transfected with expression vector encoding MAP3K7. The control group consisted of EYFP-expressing lentiviral vector pLKO PLNGY (control) and the known tumor suppressor genes RB1 and PTEN. (b, d) Effect of MAP3K7 depletion on cell growth. DU-145 (b) BPH-1 (d) cells were transfected with a MAP3K7-targeting shRNA vector. The control group consisted of shGFP expressing lentiviral vector pLKO (shGFP) and shRNA constructs against the known RB1 and PTEN (PTEN and RB1-KO). Cells were selected with puromycin and cultured for 2 weeks, and colonies were fixed with methanol, stained with Giemsa and counted.
Discussion
These data show that MAP3K7 alterations are typically monoallelic and strongly linked to adverse clinical outcome and ERG negativity in prostate cancer.
Data from our SNP array copy number analysis demonstrated a minimal commonly deleted region at 6q15 in prostate cancer containing 12 genes. This region is in the range of most previous studies reporting a small, commonly deleted area at 6q15 measuring between 0.8 and 4 Mb and containing 5–23 genes.2, 3, 4, 5 Liu et al4 identified an 817 Kb commonly deleted region that was affected in 20 of 21 tumors. The five genes located within this area (MDN1, CASP8AP2, GJA10, BACH2 and MAP3K7) are also part of the minimal commonly deleted region founding our study. All of these five genes have a known or suggested tumor suppressor function,3, 5, 6, 28, 29, 30 but accumulating evidence suggested MAP3K7 as the 6q15 tumor suppressor in prostate cancer.2, 3, 4, 5, 6 MAP3K7 was the only one of the five genes, which was found expressed at significantly lower levels in tumor cells as compared with normal prostate.4 Moreover, a correlation between 6q15 deletion and decreased MAP3K7 expression was demonstrated in prostate cancers.2 Data from our functional analyses are in line with a tumor suppressive function of MAP3K7 in prostate cancer. Moreover, Wu et al recently demonstrated a tumor suppressive function of MAP3K7 /TAK1 in a shTAK1 prostate cancer mouse model system.6, 31 Also considering the biological role of MAP3K7 as a regulator of cell growth, differentiation and apoptosis,7, 8, 9 the evidence for MAP3K7 representing the relevant tumor suppressor gene on 6q15 is very strong.
The rate of heterozygous 6q15 deletions (18.5%) detected by FISH in our series of primary prostate cancers treated by prostatectomy is in the range of most previous studies reporting between 13 and 62% 6q15 deletion in localized and metastatic prostate cancers.1, 2, 3, 4, 5, 16, 17 Most of these studies used LOH,16, 17 classical metaphase-based CGH,1 or array CGH.2, 3, 5 Only one study used FISH with a probe specific for MAP3K7 and found a deletion in 32% of 95 tumors.4 FISH represents the gold standard for the analysis of gene copy number changes. FISH enables a cell by cell analysis, making the method independent of a possible admixture of non-neoplastic cells and genomic cancer heterogeneity.
To fully exploit the strength of the FISH approach, stringent criteria were applied for assuring presence of cancer cells in each analyzed tissue spot and also for defining MAP3K7 deletions in our study. Although prostate cancer is notorious for its heterogeneity, it is unlikely, that relevant heterogeneity becomes visible within an area of 0.6 mm cancer tissue analyzed per tissue micorarray spot. We therefore assume that—at least in the vast majority of cases—MAP3K7 deletions should be present in either all or none of the tumor cells of one tissue microarray spot. Because some cells always lose some FISH signals due to nucleus truncation during tissue sectioning, a cutoff of 50% of tumor cells having less MAP3K7 than centromere 6 signals was selected in this project to define MAP3K7-deleted tumors with unequivocal clonally expansion of deleted tumor cells. The validity of our approach is supported by the observation of most deleted cases having 80% or more cells with fewer MAP3K7 than centromere 6 signals while cancers without deletion usually did not have >15% cancer cells with fewer MAP3K7 than centromere 6 signals. In an earlier study we had successfully validated this cutoff level by comparing PTEN deletions detected by FISH and CGH arrays and demonstrating a 100% concordance.22
The evaluation of a possible relationship between MAP3K7 alterations and tumor phenotype or clinical outcome was a key element of our study. The large number of tumors included in our study enabled us to identify 423 tumors with MAP3K7 deletions. The strong link of MAP3K7 deletions with morphological parameters of high malignancy and early PSA recurrence suggest that a lost or reduced MAP3K7 gene product confers substantial malignant potential to prostate cancer cells. These findings are concordant with two earlier studies, suggesting a link between MAP3K7 deletion, or loss of heterozygosis (LOH) at 6q and high Gleason grade or metastatic tumors in studies on 95 and 51 prostate cancers.4, 16 One study on 52 tumors could not find clinically relevant associations.17 There were also two reports linking 6q15 deletions to a decreased risk of tumor recurrence in 24 and 55 patients.3, 5
Our search for mechanisms leading to a complete inactivation of MAP3K7 in clinical prostate cancer samples did neither reveal MAP3K7 mutations nor any homozygous deletions. Whole genome sequencing studies of 7 common32 and 11 early-onset prostate cancers (manuscript submitted) had also not suggested inactivating rearrangements of the MAP3K7 gene. Moreover, promoter hypermethylation of MAP3K7 was not found by next-generation sequencing in 11 prostate cancers (Christoph Plass, personal communication). The complete absence of homozygous MAP3K7 inactivation is remarkable as a parallel study of PTEN deletions had revealed that 60% of the deletions were homozygous in the same patient cohort. Many genes including several tumor suppressor genes are essential for normal and neoplastic cells and cannot be completely inactivated or only in case of activation of a parallel pathway.33, 34 The list of such ‘essential’ tumor suppressor genes includes PTEN, PinX1 and NKX3.1.34, 35, 36, 37 For example, PTEN can only be completely inactivated in tumor cells if a p53-mediated senescence is bypassed.34 As our functional data did not show evidence for at least a low MAP3K7 expression being required of cell survival, it can be speculated that at least one of the immediately adjacent 6q15 genes might be essential for prostate cells.
The strong association of heterozygous MAP3K7 deletions with clinical features of prostate cancer in combination with absence of known mechanisms inactivating the second allele may suggest that MAP3K7 haploinsufficiency exerts a significant biological effect in prostate cancer cells.
Numerous genes have recently been described to contribute to cancer development and progression in case of haploinsufficiency. These include, for example, TP53, which is thought to be the most frequently mutated gene in cancer38, 39 and, PTEN40 and NKX3.141 in prostate cancer. It also seems possible, that the strong biologic/clinical effects of some of the large deletions in prostate cancer (and other tumors) are exerted by a combination of reduced expression of multiple genes.
6q15 deletions were about 2.5 times more frequent in ERG-negative as compared with ERG-positive cancers. This association was not due to a higher fraction of advanced tumors in the subset of ERG fusion-positive cancers, as stage and grade distribution was comparable in both subsets (Supplementary Table S5). These data indicate that a reduced function or inactivation of a gene on 6q15 might provide a selection advantage to ERG-negative tumor cells. Alternatively, it could be possible that 6q15 deletion prevents tumor cells from developing ERG fusions. Results of whole genome sequencing studies by us and others have demonstrated that mutations of various chromatin-regulating genes are involved in prostate cancer, demonstrating, that this gene category has a pivotal role in prostate cancer.2, 32, 42, 43 It could therefore be speculated, that untimely activation or inactivation of chromatin modifiers may substantially impact a cells ability to develop rearrangements of activated genes. However, none of the 6q15 genes has a known role as a chromatin modifier. The identical prognostic impact of MAP3K7 deletions to ERG-positive and -negative tumors demonstrates at least, that the impact of this deletion on tumor cell properties driving aggressiveness is unaffected by ERG.
In summary, our data indicate that MAP3K7 deletions are strongly linked to adverse features of prostate cancer and to the subset of ERG fusion-negative cancers. The absence of homozygous MAP3K7 deletion, MAP3K7 mutations and promoter hypermethylation in all examined tumors suggests that MAP3K7 haploinsufficiency may contribute to cancer development and progression.
References
Sun J, Liu W, Adams TS et al DNA copy number alterations in prostate cancers: a combined analysis of published CGH studies. Prostate 2007;67:692–700.
Taylor BS, Schultz N, Hieronymus H et al Integrative genomic profiling of human prostate cancer. Cancer Cell 2010;18:11–22.
Ishkanian AS, Mallof CA, Ho J et al High-resolution array CGH identifies novel regions of genomic alteration in intermediate-risk prostate cancer. Prostate 2009;69:1091–1100.
Liu W, Chang BL, Cramer S et al Deletion of a small consensus region at 6q15, including the MAP3K7 gene, is significantly associated with high-grade prostate cancers. Clin Cancer Res 2007;13:5028–5033.
Lapointe J, Li C, Giacomini CP et al Genomic profiling reveals alternative genetic pathways of prostate tumorigenesis. Cancer Res 2007;67:8504–8510.
Wu M, Shi L, Cimic A et al Suppression of Tak1 Promotes Prostate Tumorigenesis. Cancer Res 2012;72:2833–2843.
Delaney JR, Mlodzik M . TGF-beta activated kinase-1: new insights into the diverse roles of TAK1 in development and immunity. Cell Cycle 2006;5:2852–2855.
Edlund S, Bu S, Schuster N et al Transforming growth factor-beta1 (TGF-beta)-induced apoptosis of prostate cancer cells involves Smad7-dependent activation of p38 by TGF-beta-activated kinase 1 and mitogen-activated protein kinase kinase 3. Mol Biol Cell 2003;14:529–544.
Verheyen EM . Opposing effects of Wnt and MAPK on BMP/Smad signal duration. Dev Cell 2007;13:755–756.
Yang YM, Bost F, Charbono W et al C-Jun NH(2)-terminal kinase mediates proliferation and tumor growth of human prostate carcinoma. Clin Cancer Res 2003;9:391–401.
Gunnell LM, Jonason JH, Loiselle AE et al TAK1 regulates cartilage and joint development via the MAPK and BMP signaling pathways. J Bone Miner Res 2010;25:1784–1797.
Yamaguchi K, Shirakabe K, Shibuya H et al Identification of a member of the MAPKKK family as a potential mediator of TGF-beta signal transduction. Science 1995;270:2008–2011.
Davis RJ . Signal transduction by the JNK group of MAP kinases. Cell 2000;103:239–252.
Cuadrado A, Nebreda AR . Mechanisms and functions of p38 MAPK signalling. Biochem J 2010;429:403–417.
Katoh M . Networking of WNT, FGF, Notch, BMP, and Hedgehog signaling pathways during carcinogenesis. Stem Cell Rev 2007;3:30–38.
Saric T, Brkanac Z, Troyer DA et al Genetic pattern of prostate cancer progression. Int J Cancer 1999;81:219–224.
Cooney KA, Wetzel JC, Consolino CM et al Identification and characterization of proximal 6q deletions in prostate cancer. Cancer Res 1996;56:4150–4153.
McNeal JE, Redwine EA, Freiha FS et al Zonal distribution of prostatic adenocarcinoma. Correlation with histologic pattern and direction of spread. Am J Surg Pathol 1988;12:897–906.
Minner S, Jessen B, Stiedenroth L et al Low level HER2 overexpression is associated with rapid tumor cell proliferation and poor prognosis in prostate cancer. Clin Cancer Res 2010;16:1553–1560.
Bubendorf L, Kononen J, Koivisto P et al Survey of gene amplifications during prostate cancer progression by high-throughout fluorescence in situ hybridization on tissue microarrays. Cancer Res 1999;59:803–806.
Minner S, Enodien M, Sirma H et al ERG status is unrelated to PSA recurrence in radically operated prostate cancer in the absence of antihormonal therapy. Clin Cancer Res 2011;17:5878–5888.
Krohn ADT, Burkhardt L, Mayer PS et al Genomic deletion of PTEN is associated with tumor progression and early PSA recurrence in ERG fusion positive and fusion negative prostate cancer. Am J Pathol 2012;181:401–412.
Mader M, Simon R, Steinbiss S et al FISH Oracle: a web server for flexible visualization of DNA copy number data in a genomic context. J Clin Bioinforma 2011;1:20.
Boehm JS, Zhao JJ, Yao J et al Integrative genomic approaches identify IKBKE as a breast cancer oncogene. Cell 2007;129:1065–1079.
Cuchet D, Sykes A, Nicolas A et al PML isoforms I and II participate in PML-dependent restriction of HSV-1 replication. J Cell Sci 2011;124:280–291.
Livak KJ, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001;25:402–408.
Zitzer H, Wente W, Brenner MB et al Sterol regulatory element-binding protein 1 mediates liver X receptor-beta-induced increases in insulin secretion and insulin messenger ribonucleic acid levels. Endocrinology 2006;147:3898–3905.
Vieira SA, Deininger MW, Sorour A et al Transcription factor BACH2 is transcriptionally regulated by the BCR/ABL oncogene. Genes Chromosomes Cancer 2001;32:353–363.
Miasari M, Puthalakath H, Silke J . Ubiquitylation and cancer development. Curr Cancer Drug Targets 2008;8:118–123.
Leithe E, Sirnes S, Omori Y et al Downregulation of gap junctions in cancer cells. Crit Rev Oncog 2006;12:225–256.
Xin L, Lawson DA, Witte ON . The Sca-1 cell surface marker enriches for a prostate-regenerating cell subpopulation that can initiate prostate tumorigenesis. Proc Natl Acad Sci USA 2005;102:6942–6947.
Berger MF, Lawrence MS, Demichelis F et al The genomic complexity of primary human prostate cancer. Nature 2011;470:214–220.
Berger AH, Pandolfi PP . Haplo-insufficiency: a driving force in cancer. J Pathol 2011;223:137–146.
Chen Z, Trotman LC, Shaffer D et al Crucial role of p53-dependent cellular senescence in suppression of Pten-deficient tumorigenesis. Nature 2005;436:725–730.
Chen Z, Carracedo A, Lin HK et al Differential p53-independent outcomes of p19(Arf) loss in oncogenesis. Sci Signal 2009;2: ra44.
Zhou XZ, Huang P, Shi R et al The telomerase inhibitor PinX1 is a major haploinsufficient tumor suppressor essential for chromosome stability in mice. J Clin Invest 2011;121:1266–1282.
Mogal AP, van der Meer R, Crooke PS et al Haploinsufficient prostate tumor suppression by Nkx3.1: a role for chromatin accessibility in dosage-sensitive gene regulation. J Biol Chem 2007;282:25790–25800.
Greenblatt MS, Bennett WP, Hollstein M et al Mutations in the p53 tumor suppressor gene: clues to cancer etiology and molecular pathogenesis. Cancer Res 1994;54:4855–4878.
Coles C, Condie A, Chetty U et al p53 mutations in breast cancer. Cancer Res 1992;52:5291–5298.
Kwabi-Addo B, Giri D, Schmidt K et al Haploinsufficiency of the Pten tumor suppressor gene promotes prostate cancer progression. Proc Natl Acad Sci USA 2001;98:11563–11568.
Abdulkadir SA, Magee JA, Peters TJ et al Conditional loss of Nkx3.1 in adult mice induces prostatic intraepithelial neoplasia. Mol Cell Biol 2002;22:1495–1503.
Kumar A, Shendure J, Nelson PS . Genome interrupted: sequencing of prostate cancer reveals the importance of chromosomal rearrangements. Genome Med 2011;3:23.
Wilson BG, Roberts CW . SWI/SNF nucleosome remodellers and cancer. Nat Rev Cancer 2011;11:481–492.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Modern Pathology website
Rights and permissions
About this article
Cite this article
Kluth, M., Hesse, J., Heinl, A. et al. Genomic deletion of MAP3K7 at 6q12-22 is associated with early PSA recurrence in prostate cancer and absence of TMPRSS2:ERG fusions. Mod Pathol 26, 975–983 (2013). https://doi.org/10.1038/modpathol.2012.236
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2012.236
Keywords
This article is cited by
-
TP63–TRIM29 axis regulates enhancer methylation and chromosomal instability in prostate cancer
Epigenetics & Chromatin (2024)
-
6q deletion is frequent but unrelated to patient prognosis in breast cancer
Breast Cancer (2022)
-
TGF-β1 promotes epithelial-to-mesenchymal transition and stemness of prostate cancer cells by inducing PCBP1 degradation and alternative splicing of CD44
Cellular and Molecular Life Sciences (2021)
-
Up regulation of the Hippo signalling effector YAP1 is linked to early biochemical recurrence in prostate cancers
Scientific Reports (2020)
-
Secreted Frizzled-Related Protein 4 (SFRP4) Is an Independent Prognostic Marker in Prostate Cancers Lacking TMPRSS2: ERG Fusions
Pathology & Oncology Research (2020)