Abstract
Dedifferentiation of hepatocellular carcinoma implies aggressive clinical behavior and is associated with an increasing number of genomic alterations, eg deletion of 13q. Genes directly or indirectly deregulated due to these genomic alterations are mainly unknown. Therefore this study compares array comparative genomic hybridization and whole genome gene expression data of 23 well, moderately, or poorly dedifferentiated hepatocellular carcinoma, using unsupervised hierarchical clustering. Dedifferentiated carcinoma clearly branched off from well and moderately differentiated carcinoma (P<0.001 χ2-test). Within the dedifferentiated group, 827 genes were upregulated and 33 genes were downregulated. Significance analysis of microarrays for hepatocellular carcinoma with and without deletion of 13q did not display deregulation of any gene located in the deleted region. However, 531 significantly upregulated genes were identified in these cases. A total of 6 genes (BIC, CPNE1, RBPMS, RFC4, RPSA, TOP2A) were among the 20 most significantly upregulated genes both in dedifferentiated carcinoma and in carcinoma with loss of 13q. These genes are involved in cell-cycle control and proliferation. Of 33 downregulated genes in the dedifferentiated subgroup, 4 metallothioneins had the lowest fold change, most probably mediated through inactivation of C/EBPα by the PI3K/AKT cascade. In conclusion dedifferentiation of hepatocellular carcinoma is associated with upregulation of genes involved in cell-cycle control and proliferation. Notably, a significant portion of these genes is also upregulated in carcinoma with deletion of 13q. As no downregulated genes were identified and microRNAs (mir-621, mir-16-1, mir-15a) are located within the deleted region of 13q and may be lost, we speculate that these miRNAs may induce the upregulation of critical cell-cycle control genes.
Similar content being viewed by others
Main
Sustained basic, clinical, and epidemiological studies have previously been performed to address the major problems caused by the fourth most malignant tumor in the world: hepatocellular carcinoma1 (http://www-dep.iarc.fr/). In many previous studies, this has led to the detection of various mutations of genes (eg p53, IGF2R, p16, PTEN) typically associated with the development of hepatocellular carcinoma.2 Besides these specific mutations, the carcinogenesis of hepatocellular carcinoma is known to be correlated with telomere shortening3 and chromosomal instability.4
Chromosomal instability is defined by microscopically detectable partial or complete gain or loss of chromosomes typically occurring in hepatocellular carcinoma and also in many other solid tumors.5, 6 The typical set of chromosomal imbalances found in comparative genomic hybridization studies includes gains of 1q, 6p, 8q, 17q, and 20q, and losses of 1p, 4q, 6q, 8p, 13q, 16q, and 17p detected in >90% of 780 cases analyzed up to now (see references 7, 8, 9, 10 for review:11). These aberrations have been correlated with etiological factors such as viral (hepatitis B, hepatitis C) infections.12, 13, 14 Clinical impact has been reported for losses of 8p and gain of 20q with respect to stage and prognosis.15 On the basis of genomic alterations detected by aCGH, hepatocellular carcinoma and its benign counterpart, hepatocellular adenoma, could be differentiated.16 It was also noted that dedifferentiation of hepatocellular carcinoma was closely correlated to losses of 4q and 13q.11 By fluorescence in situ hybridization, a stepwise increase in genomic instability, as well as a highly significant correlation of histological grading and grade of polyploidization, was seen.17 This is of clinical interest because morphological grading has been shown to correlate with survival.18, 19 Elucidation of the processes causing dedifferentiation of hepatocellular carcinoma is required because hepatocellular carcinoma dedifferentiated in more than half of patients within 7–34 months.20, 21
Investigations into the genomic and mRNA expression level have been attempted to provide insight into these processes, but a combined analysis has not yet been carried out. Using newly developed software tools for the analysis of genomic array data, we therefore aimed to identify genes deregulated during dedifferentiation of hepatocellular carcinoma, particularly those located in chromosomal regions known to be altered in poorly differentiated hepatocellular carcinoma.
Materials and methods
Clinical, serological, and pathomorphological data of 23 patients with hepatocellular carcinoma treated at Hannover Medical School between 1998 and 2002 are given in Table 1. Tumor specimens from these patients were analyzed with regard to mRNA expression in hepatocellular adenoma in an earlier study.22 Tumor material was obtained as surgical specimens taken for diagnostic purposes as permitted by the Ethics Committee of Hannover Medical School. Unfixed tissue samples of 0.5 cm in diameter were snap-frozen within 30 min after resection and stored at −80°C. A part of the tissue was cut out for RNA extraction; another piece was fixed in formalin and embedded in paraffin for histological examination. This ensured the closest proximity and homogeneity of the tissues used for different experiments.
Histological classification according to WHO criteria,23 was carried out after H&E and reticulin staining by two experienced pathologists (L.W., P.F.). A total of 4 well, 11 moderately, and 8 poorly differentiated hepatocellular carcinoma were detected with histological signs of partial or complete dedifferentiation (Figure 1).
Histological differentiation of hepatocellular carcinoma showing cytological and architectural features. (a) Well-differentiated hepatocellular carcinoma is defined by nuclei shape, which is very similar to normal hepatocytes. Additional staining (eg silver staining) may be required to ensure correct diagnosis. (b) Moderately differentiated carcinoma reveals a heterogeneous pattern with respect to cytology of nuclei and architectural pattern. (c) In moderate-to-poorly differentiated hepatocellular carcinoma, prominent nucleoli are seen in the nuclei; the shape of the nuclei and membrane is irregular. The trabecular pattern is also disturbed in a prominent way, making recognition of trabecular structures difficult. (d) In dedifferentiated hepatocellular carcinoma, nuclei have a bizarre and highly irregular shape, and lose their architectural pattern, making it difficult to separate these carcinomas from dedifferentiated carcinomas of other histogenetic origin.
For 22/23 cases of hepatocellular carcinoma, cytogenetic characterization was already performed using aCGH; detailed information is given in an earlier study.16 The findings described according to the international system of cytogenetic nomenclature ISCN24 are given in Supplementary Table 1. In brief, the most commonly gained region was 1q22.1–23.1 in 20 cases (91%). Gains of 8q were observed in 14 cases (64%), losses of 8p in 11 cases (50%), gains of 20q in 9 cases (41%), losses of 13q in 9 cases (41%), losses of 4q in 8 cases (36%), losses of 6q in 8 cases (36%), gains of 6p in 7 cases (32%), and losses of 17p in 7 cases (32%).
Methods
cDNA microarray analysis
cDNA microarray experiments were carried out as previously described.22 In brief, total RNA was isolated using Trizol (Invitrogen, Karlsruhe, Germany) followed by RNeasy Mini Kit (Qiagen, Hilden, Germany) and linearly amplified (MessageAmp aRNA Kit; Ambion, Huntingdon, UK). Aliquots of 1.5 μg of amplified RNA from the tumors and the human reference were reverse transcriptase labeled with Cy3 (reference) and Cy5 (tumor) (Amersham Biosciences, Little Chalfont, UK). Labeled RNA of tumors and references were comparatively hybridized overnight onto a cDNA chip containing the ‘Resgen clone set’ with >43 000 spots covering the entire human genome (Stanford Functional Genomics Facility, Stanford, CA, USA) according to the posted protocol (http://brownlab.Stanford.edu/protocols.html). Arrays were imaged using a GenePix 4000B scanner and dependent software (GenePix Pro 4.1; Axon Instruments). Data were exported to the Stanford Microarray Database (http://smd.stanford.edu). Fluorescence ratios were normalized for each array. Data were excluded if the regression correlation was <0.6, or the mean spot intensity/median background intensity was <1.5. The measured genes were subsequently ‘mean-centered’ (ie reported for each gene relative to the mean ratio across all samples, rather than relative to the human reference).
The hierarchical clustering algorithm based on Pearson correlation (centered) coefficients grouped the genes (as well as the samples) on the basis of similarity in their expression pattern. Normalized and log-transformed expression data served as input for significance analysis of microarrays25 with standard parameters. False discovery rates varied between 0.0 and 0.54.
RT–PCR
Quantitative RT-PCR was performed on the selected genes MT1F, MT1G, and TOP2A, to confirm the array results.
In a total volume of 120 μl, 1 μg of total RNA was reverse transcribed by M-MuLV reverse transcriptase using random hexamer primers (RevertAid First Strand cDNA Synthesis Kit; Fermentas, St-Leon-Rot, Germany). TOP2A cDNA samples (2 μl) were amplified on an iCycler iQ real-time detection system (Bio-Rad Laboratories, München, Germany) using a QuantiTect SYBR Green PCR kit (Qiagen, Hilden, Germany). TBP was used as the internal control gene. Aliquots of 5 μl of each cDNA sample were used for the analysis of MT1F and MT1G expression. This expression was performed using an iCycler iQ real-time detection system (Bio-Rad Laboratories). Primers and probes for MT1F and MT1G were synthesized by TibMolBiol (Berlin, Germany). Detailed PCR conditions and primer sequences for all genes used are shown in Table 2. β-Actin was used as the internal control gene. For calibration, the human reference (Stratagene, Amsterdam, Netherlands) was used. Relative quantification of gene expression was determined according to the ΔΔ-CT method using an iCycler iQ real-time detection system software version 3.1 (Bio-Rad Laboratories). All samples were analyzed in triplicate.
Results
cDNA Microarray Analysis
Gene expression analysis led to successful evaluation of all 23 cases; 7977 cDNA clones with the most varying expression among the tumor and reference samples were selected for further bioinformatical analysis.
Hierarchical Clustering
Unsupervised hierarchical clustering separated hepatocellular carcinoma into two groups of 11 and 12 cases. All well-differentiated carcinoma were seen in the first group. In the second group, all low-differentiated carcinoma occurred, with the exception of case N91. In the former group, six moderately differentiated carcinoma were seen, whereas five moderately differentiated carcinoma occurred in the latter (Figure 2).
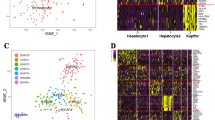
Unsupervised hierarchical clustering of global gene expression analysis. Upregulated genes are marked in red, downregulated genes in green. Two groups of hepatocellular carcinoma were obtained using this approach, with all well-differentiated carcinoma clustering in group A and all low-differentiated carcinoma clustering in group B with only one exception (N91). Moderately differentiated carcinomas were found in both clusters. Carcinomas of these two groups were analyzed in regard to histological grading and 13q−.
Statistical analysis was further performed by correlation with histological grading. For this parameter, well and moderately differentiated carcinoma were taken together (15 cases) and compared to low-differentiated carcinoma (8 cases). A highly significant correlation (P<0.001, χ2-test) was detectable in regard to the histological grading. For example, four metallothioneins (MT1E, MT1F, MT1M, MT2A) had the lowest fold change (0.09–0.18) in poorly differentiated hepatocellular carcinoma. Supervised cluster analyses for these genes branched off all dedifferentiated tumors from well and moderately differentiated samples (Figure 3).
Supervised cluster analyses for members of the metallothionein family branched off all dedifferentiated carcinoma from well and moderately differentiated tumors. Well-differentiated carcinomas are grouped together in one group, whereas low-differentiated carcinoma are in the other group. Upregulated genes are marked in red, downregulated genes in green.
Analysis of Gene Expression by Statistical Analysis of Microarrays
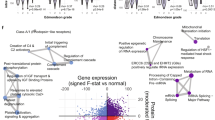
Using this approach for ‘histological grading’, the number of significantly dysregulated genes with a false discovery rate of <5% was 860. A total of 827 genes were upregulated in the group of dedifferentiated carcinoma; 33 genes were downregulated (Supplementary Tables 2 and 3). For loss of 13q, the number of dysregulated genes was 531.
For further analysis of these findings, we focused on 20 genes seen with the highest or lowest fold changes (Table 3). For histological grading, the fold change of the most upregulated 20 genes was 2.58–7.47. For the 20 downregulated genes, this change was 0.09–0.47. For the parameter loss of 13q, the top 20 upregulated genes had fold changes of 2.62–4.00. Only two genes (MDFIC and GALE) were downregulated, with fold changes of 0.38 and 0.40, respectively.
Six genes were found in both sets of upregulated genes, one from dedifferentiated carcinoma and one from carcinoma with 13q− (BIC, CPNE1, RBPMS, RFC4, RPSA, TOP2A). All of these genes mentioned are involved in cell-cycle control and proliferation. Supervised cluster analyses for these genes, branched off all dedifferentiated carcinoma from well and the moderately differentiated carcinoma. Furthermore, these analyses branch off the most carcinoma with 13q− from carcinoma without 13q− (Figure 4).
Supervised clustering including six genes (BIC, CPNE1, RBPMS, RFC4, RPSA, TOP2A) branched off all dedifferentiated hepatocellular carcinoma from well and moderately differentiated carcinoma, with only one moderately differentiated hepatocellular carcinoma sample (N1) occurring in the cluster of dedifferentiated carcinoma. Vice versa, no dedifferentiated carcinoma was seen in the groups of well or moderately differentiated carcinoma. Furthermore, these analyses branched off most of carcinoma with loss of 13q without 13q−. Upregulated genes are marked in red, downregulated genes in green.
In the sets of genes downregulated in poorly differentiated carcinoma, four metallothioneins (MT1E, MT1F, MT1M, MT2A) had the lowest fold change (0.09–0.18). All metallothioneins are allocated to chromosome band 16q13. A loss of 16q13 may have caused the downregulation in three samples (N1, N37, N83) according to the results of aCGH.
Real-Time RT-PCR for Relative Quantification of TOP2A, MT1F, and MT1G
Results obtained for mRNA expression analysis were proven exemplarily for genes TOP2A, MT1G, and MT1F. TOP2A was found upregulated in both sets, MT1G and MT1F were downregulated in the group of dedifferentiated hepatocellular carcinoma. RT–PCR confirmed the results of array analysis revealing a systematic increase of mRNA expression of TOP2A and a decrease for MT1G and MT1F as also found by statistical analysis of microarrays, summarized graphically in Figure 5.
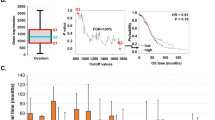
Graphical illustration of RT–PCR. Array-based mRNA expression analysis was confirmed by these experiments. The decreased gene expression on the mRNA level for MT1F and MT1G was also detectable by quantitative RT-PCR as demonstrated by significantly lower ΔΔ-CT values in low-differentiated carcinoma compared to well and moderately differentiated carcinoma. Vice versa, TOP2A was seen to be upregulated in low-differentiated carcinoma, as expected after array-based expression analysis.
Comparison of Cytogenetic Data with mRNA Expression
Because no up- or downregulated gene was located on 13q, and looking for additional factors that could be influenced by deletion of 13q, an Internet-based search using the Sanger Data Bank (www.microrna.sanger.ac.uk) was performed and identified miRNAs mir-621, mir-16-1, and mir-15a within the deleted region of 13q. As reported earlier, miRNAs can alter gene expression in a direct or nondirect way (for review see reference 25), and thus are possible candidates modulating gene expression.
Discussion
Array-based gene expression analyses on hepatocellular carcinoma have been performed to identify genes in regard to grading, staging, and etiological factors associated with clinical outcome.26, 27, 28, 29, 30, 31
As 50–75% of patients develop dedifferentiation within 7–34 months,20, 21 this has a major clinical impact due to the extremely aggressive clinical behavior of dedifferentiated hepatocellular carcinoma.18, 19 We aimed to identify genes involved in the dedifferentiation of hepatocellular carcinoma. We previously showed that chromosomal instability increases parallel to dedifferentiation of hepatocellular carcinoma.17 Moreover, the number of copy number alterations increases parallel to dedifferentiation of hepatocellular carcinoma. Hepatocellular adenoma have a significantly lower number of copy number alterations than carcinoma. Hepatocellular adenoma can also be distinguished from carinoma based on global gene expression profiling.22 Comparing aCGH and gene expression data, it was possible to define a set of seven upregulated genes localized on chromosome band 1q22. As histological dedifferentiation is accompanied by loss of 13q,11, 16 we focused on gene expression in different grades of hepatocellular carcinoma bearing 13q or not. We therefore aimed in this study to analyze whether well and poorly differentiated carcinoma can be distinguished based on gene expression profiling, in order to identify genes involved in the dedifferentiation of hepatocellular carcinoma.
Overall statistical analysis of microarrays analysis revealed 860 genes significantly dysregulated in dedifferentiated hepatocellular carcinoma and 533 genes related to loss of 13q. When comparing 20 genes most upregulated both during dedifferentiation and due to loss of 13q, 6 genes (BIC, CPNE1, RBPMS, RFC4, RPSA, TOP2A) were found. B-cell receptor-inducible gene (BIC) was originally identified as a retroviral integration site in avian leukosis virus,32 and its product, miRNA 155, accumulates in various B-cell neoplasias.33 Upregulation of BIC is important in lymphomagenesis due to interaction with C-MYC,34 which is a gene that also plays a fundamental role in hepatocarcinogenesis. CPNE1 encodes a calcium-dependent membrane-binding protein. It has a broad tissue distribution, and is thought to be involved in membrane trafficking. RBPMS is a member of the RRM family of RNA-binding proteins. The protein encoded by this gene has a single, putative RRM domain, and its function is probably binding to RNA.
RPSA belongs to a family of extracellular matrix glycoproteins called laminins. These proteins have been implicated in cell adhesion, differentiation, migration, signaling, and metastasis, processes frequently altered in cancer cells, determining their invasive and metastatic potential. RFC4, together with PCNA, is necessary for elongation of primed DNA templates by DNA polymerase-δ and DNA polymerase-ɛ. Topoisomerase 2α encodes a protein that controls and alters the topologic states of DNA during transcription. It is found in the nucleus and involved in chromosome condensation, chromatid separation, and the relief of torsional stress during the transcription and replication of DNA. It catalyzes the transient breaking and rejoining of two strands of duplex DNA, which allows the strands to pass through one another, thus altering the topology of DNA. Moreover, high Topoisomerase 2α expression appears to be linked with cellular dedifferentiation and potentially aggressive tumor phenotype in invasive breast cancer.35
For the genes mentioned above, more detailed information can be obtained by accessing the URLs in Supplementary Table 2.
Significantly downregulated genes were seen to be associated with dedifferentiation. Four metallothionein genes had the lowest fold change. Metallothioneins are a group of cysteine-rich intracellular proteins with high affinity for metal ions. Their enzymatic activity is not known. Metallothionein genes are located in chromosome band 16q13. In hepatocellular carcinoma, a decrease in the intensity of immunohistochemical staining has been reported for metallothioneins.36, 37 Metallothioneins were also found to be downregulated in hepatocellular carcinoma in a global mRNA expression analysis by Okabe et al,30 concordant with our results. In patients suffering from hepatocellular carcinoma, metallothioneins were measured in serum, and the concentration correlated inversely with the grade of differentiation of hepatocellular carcinoma.38, 39 In oxidative stress, eg due to infection with hepatitis-B virus or hepatitis-C virus leading to chronic inflammation, metallothioneins are reported to be cell-protective in mice.40 Conversely, MT-1 and MT-2 knockout mice are markedly more sensitive to chemically induced carcinogenesis.41, 42 Very recently, it has been shown that expression of metallothioneins is suppressed in hepatocellular carcinoma mediated through inactivation of C/EBPα by the PI3K/AKT cascade.43 Elements of this pathway (eg ErbB2, mTOR) can be modulated by various drugs44 that are probably candidates for targeted treatment of hepatocellular carcinoma.
Interestingly, none of the downregulated genes were identified within the deleted region of 13q. However, three known miRNAs (mir-621, mir-16-1, mir-15a) are located in the deleted chromosomal region of 13q and therefore may be lost. This could lead to a less effective inhibition of targets of these miRNA and to an increase of gene expression of putative oncogenes, particularly on the protein level. It may also be speculated that partial loss of miRNA positively upregulates critical cell-cycle control genes such as those listed above.
In conclusion, dedifferentiation of hepatocellular carcinoma is associated with upregulation of genes involved in cell-cycle control and proliferation. Notably, a significant portion of these genes is also upregulated in hepatocellular carcinoma with deletion of 13q. It is unclear how genes located in chromosome regions other than 13q are upregulated because downregulated genes were not identified within the deleted region of 13q. As three known miRNAs (mir-621, mir-16-1, mir-15a) are located in the deleted chromosomal region of 13q, we speculate that these miRNAs are involved in dedifferentiation and in the induction of chromosomal instability in hepatocellular carcinoma and should be investigated in further studies.
References
El-Serag HB . Hepatocellular carcinoma: an epidemiologic view. J Clin Gastroenterol 2002;35 (5 Suppl 2):72–78.
Thorgeirsson SS, Grisham JW . Molecular pathogenesis of human hepatocellular carcinoma. Nat Genet 2002;31:339–346.
Wiemann SU, Satyanarayana A, Buer J, et al. Contrasting effects of telomere shortening on organ homeostasis, tumor suppression, and survival during chronic liver damage. Oncogene 2005;24:1501–1509.
Plentz RR, Caselitz M, Bleck JS, et al. Hepatocellular telomere shortening correlates with chromosomal instability and the development of human hepatoma. Hepatology 2004;40:80–86.
Mitelman F . Catalog of Chromosome Aberrations in Cancer. 5th edn. Wiley-Liss: New York, 1994.
Rajagopalan H, Lengauer C . Aneuploidy and cancer. Nature 2004;432:338–341.
Marchio A, Meddeb M, Pineau P, et al. Recurrent chromosomal abnormalities in hepatocellular carcinoma detected by comparative genomic hybridization. Genes Chromosomes Cancer 1997;18:59–65.
Wong N, Lai P, Lee SW, et al. Assessment of genetic changes in hepatocellular carcinoma by comparative genomic hybridization analysis: relationship to disease stage, tumor size, and cirrhosis. Am J Pathol 1999;154:37–43.
Kusano N, Shiraishi K, Kubo K, et al. Genetic aberrations detected by comparative genomic hybridization in hepatocellular carcinomas: their relationship to clinicopathological features. Hepatology 1999;29:1858–1862.
Wilkens L, Bredt M, Flemming P, et al. Diagnostic impact of fluorescence in situ hybridization in the differentiation of hepatocellular adenoma and well-differentiated hepatocellular carcinoma. J Mol Diagn 2001;3:68–73.
Moinzadeh P, Breuhahn K, Stutzer H, et al. Chromosome alterations in human hepatocellular carcinomas correlate with aetiology and histological grade—results of an explorative CGH meta-analysis. Br J Cancer 2005;92:935–941.
Laurent-Puig P, Legoix P, Bluteau O, et al. Genetic alterations associated with hepatocellular carcinomas define distinct pathways of hepatocarcinogenesis. Gastroenterology 2001;120:1763–1773.
Hashimoto K, Mori N, Tamesa T, et al. Analysis of DNA copy number aberrations in hepatitis C virus-associated hepatocellular carcinomas by conventional CGH and array CGH. Mod Pathol 2004;17:617–622.
Sy SM, Wong N, Lai PB, et al. Regional over-representations on chromosomes 1q, 3q and 7q in the progression of hepatitis B virus-related hepatocellular carcinoma. Mod Pathol 2005;6:86–92.
Guan XY, Fang Y, Sham JS, et al. Recurrent chromosome alterations in hepatocellular carcinoma detected by comparative genomic hybridization. Genes Chromosomes Cancer 2000;29:110–116.
Steinemann D, Skawran B, Becker T, et al. Assessment of differentiation and progression of hepatic tumors using array- based comparative genomic hybridization. Clin Gastroenterol Hepatol 2006;4:1283–1291.
Wilkens L, Flemming P, Gebel M, et al. Induction of aneuploidy by increasing chromosomal instability during dedifferentiation of hepatocellular carcinoma. Proc Natl Acad Sci USA 2004;101:1309–1314.
Nzeako UC, Goodman ZD, Ishak KG . Comparison of tumor pathology with duration of survival of North American patients with hepatocellular carcinoma. Cancer 1995;76:579–588.
Lauwers GY, Terris B, Balis UJ, et al. Prognostic histologic indicators of curatively resected hepatocellular carcinomas: a multi-institutional analysis of 425 patients with definition of a histologic prognostic index. Am J Surg Pathol 2002;26:25–34.
Kenmochi K, Sugihara S, Kojiro M . Relationship of histologic grade of hepatocellular carcinoma (HCC) to tumor size, and demonstration of tumor cells of multiple different grades in single small HCC. Liver 1987;7:18–26.
Sugihara S, Nakashima O, Kojiro M, et al. The morphologic transition in hepatocellular carcinoma. A comparison of the individual histologic features disclosed by ultrasound-guided fine-needle biopsy with those of autopsy. Cancer 1992;70:1488–1492.
Skawran B, Steinemann D, Weigmann A, et al. Gene expression profiling in hepatocelluar carcinoma: coordinated up-regulation of genes localised in amplified chromosome regions. Mod Pathol 2008;21:505–516.
Ishak KG, Anthony PP, Sobin LH . Histological typing of tumours of the liver. World Health Organization International Histological Classification of Tumours. 2nd edn. Springer-Verlag: Berlin, 1994.
Shaffer LG, Tommerup N, eds. ISCN 2005: An International System for Human Cytogenetic Nomenclature. S. Karger: Basel, 2005.
Tusher VG, Tibshirani R, Chu G . Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA 2001;98:5116–5121.
Neo SY, Leow CK, Vega VB, et al. Identification of discriminators of hepatoma by gene expression profiling using a minimal dataset approach. Hepatology 2004;39:944–953.
Wu CG, Salvay DM, Forgues M, et al. Distinctive gene expression profiles associated with Hepatitis B virus x protein. Oncogene 2001;20:3674–3682.
Smith MW, Yue ZN, Geiss GK, et al. Identification of novel tumor markers in hepatitis C virus-associated hepatocellular carcinoma. Cancer Res 2003;63:859–864.
Shirota Y, Kaneko S, Honda M, et al. Identification of differentially expressed genes in hepatocellular carcinoma with cDNA microarrays. Hepatology 2001;33:832–840.
Okabe H, Satoh S, Kato T, et al. Genome-wide analysis of gene expression in human hepatocellular carcinomas using cDNA microarray: identification of genes involved in viral carcinogenesis and tumor progression. Cancer Res 2001;61:2129–2137.
Iizuka N, Oka M, Yamada-Okabe H, et al. Molecular signature in three types of hepatocellular carcinoma with different viral origin by oligonucleotide microarray. Int J Oncol 2004;24:565–574.
Clurman BE, Hayward WS . Multiple proto-oncogene activations in avian leukosis virus-induced lymphomas: evidence for stage-specific events. Mol Cell Biol 1989;9:2657–2664.
Eis PS, Tam W, Sun L, et al. Accumulation of miR-155 and BIC RNA in human B cell lymphomas. Proc Natl Acad Sci USA 2005;102:3627–3632.
Tam W, Hughes SH, Hayward WS, et al. Avian bic, a gene isolated from a common retroviral site in avian leukosis virus-induced lymphomas that encodes a noncoding RNA, cooperates with c-myc in lymphomagenesis and erythroleukemogenesis. J Virol 2002;76:4275–4286.
Nakopoulou L, Lazarisa AC, Kavantzasa N, et al. DNA topoisomerase II-alpha immunoreactivity as a marker of tumor aggressiveness in invasive breast cancer. Pathobiology 2000;68:137–143.
Theocharis SE, Margeli AP, Klijanienko JT, et al. Metallothionein expression in human neoplasia. Histopathology 2004;45:103–118.
Deng DX, Chakrabarti S, Waalkes MP, et al. Metallothionein and apoptosis in primary human hepatocellular carcinoma and metastatic adenocarcinoma. Histopathology 1998;32:340–347.
Endo T, Yoshikawa M, Ebara M, et al. Immunohistochemical metallothionein expression in hepatocellular carcinoma: relation to tumor progression and chemoresistance to platinum agents. J Gastroenterol 2004;39:1196–1201.
Ebara M, Fukuda H, Hatano R, et al. Relationship between copper, zinc and metallothionein in hepatocellular carcinoma and its surrounding liver parenchyma. J Hepatol 2000;33:415–422.
Quaife CJ, Cherne RL, Newcomb TG, et al. Metallothionein overexpression suppresses hepatic hyperplasia induced by hepatitis B surface antigen. Toxicol Appl Pharmacol 1999;155:107–116.
Suzuki JS, Nishimura N, Zhang B, et al. Metallothionein deficiency enhances skin carcinogenesis induced by 7,12-dimethylbenz[a]anthracene and 12-O-tetradecanoylphorbol-13-acetate in metallothionein-null mice. Carcinogenesis 2003;24:1123–1132.
Waalkes MP, Liu J, Kasprzak KS, et al. Hypersusceptibility to cisplatin carcinogenicity in metallothionein-I/II double knockout mice: production of hepatocellular carcinoma at clinically relevant doses. Int J Cancer 2006;119:28–32.
Datta J, Majumder S, Kutay H, et al. Metallothionein expression is suppressed in primary human hepatocellular carcinomas and is mediated through inactivation of CCAAT/enhancer binding protein alpha by phosphatidylinositol 3-kinase signaling cascade. Cancer Res 2007;67:2736–2746.
Pang RW, Poon RT . From molecular biology to targeted therapies for hepatocellular carcinoma: the future is now. Oncology 2007;72 (Suppl 1):30–44.
Acknowledgements
This work was supported by grants from the Deutsche Forschungsgemeinschaft (KFO 119) and the Deutsche Krebshilfe (106200).
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/conflict of interest
The authors have no conflict of interest to disclose.
Supplementary Information accompanies the paper on Modern Pathology website (http://www.nature.com/modpathol)
Rights and permissions
About this article
Cite this article
Skawran, B., Steinemann, D., Becker, T. et al. Loss of 13q is associated with genes involved in cell cycle and proliferation in dedifferentiated hepatocellular carcinoma. Mod Pathol 21, 1479–1489 (2008). https://doi.org/10.1038/modpathol.2008.147
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2008.147
Keywords
This article is cited by
-
Exploring the Activities of RBPMS Proteins in Myocardial Biology
Pediatric Cardiology (2019)
-
MiRNA-621 sensitizes breast cancer to chemotherapy by suppressing FBXO11 and enhancing p53 activity
Oncogene (2016)
-
The role and clinical implications of microRNAs in hepatocellular carcinoma
Science China Life Sciences (2012)
-
Molecular profile and copy number analysis of sporadic colorectal cancer in Taiwan
Journal of Biomedical Science (2011)
-
Hepatitis B virus X protein downregulates expression of the miR-16 family in malignant hepatocytes in vitro
British Journal of Cancer (2011)