Abstract
Alterations in methylation of CpG dinucleotides at the 5 position of deoxycytidine residues (5mC) are a hallmark of cancer cells, including testicular germ cell tumors. Virtually all testicular germ cell tumors are believed to be derived from intratubular germ cell neoplasia unclassified (IGCNU), which is thought to arise from primordial germ cells. Prior studies revealed that seminomas contain reduced levels of global DNA methylation as compared with nonseminomatous germ cell tumors. Smiraglia et al have proposed a model whereby seminomas arise from IGCNU cells derived from primordial germ cells that have undergone 5mC erasure, and nonseminomas arise from IGCNU cells derived from primordial germ cells that have already undergone de novo methylation after the original erasure of methylation and contain normal 5mC levels. Yet the methylation status of IGCNU has not been determined previously. We used immunohistochemical staining against 5mC to evaluate global methylation in IGCNU and associated invasive testicular germ cell tumors. Strikingly, staining for 5mC was undetectable (or markedly reduced) in the majority of IGCNU and seminomas, yet there was robust staining in nonseminomatous germ cell tumors. The lack of staining for 5mC in IGCNU and seminomas was also found in mixed germ cell tumors containing both seminomatous and nonseminomatous components. Lack of 5mC staining was not related to a lack of the maintenance methyltransferase (DNA methyltransferase 1) protein. We conclude that testicular germ cell tumors are derived in most cases from IGCNU cells that have undergone developmentally programmed 5mC erasure and that the degree of subsequent de novo methylation is most closely related to the differentiation state of the neoplastic cells. That is, IGCNU cells and seminoma cells remain unmethylated, whereas all other histological types appear to arise after de novo methylation.
Similar content being viewed by others
Main
Germ cell tumors are the most common form of testicular neoplasm and can be divided into three main groups infantile/prepubertal, adolescent/young adult and spermatocytic seminoma.1 The adolescent/young adult group, consisting of seminomatous and nonseminomatous germ cell tumors in postpubertal patients, is by far the most common form. Epidemiological studies of postpubertal testicular germ cell tumors suggest that the initiating events may occur during the prenatal period.1 Although genetic changes, such as gain of the short arm of chromosome 12, are very common in testicular germ cell tumors,1, 2 epigenetic changes may also be important in initiating these tumors.
Somatic epigenetic alterations, such as changes in methylation of deoxycytidine residues at the 5 position in CpG dinucleotides (5mC), are a hallmark of cancer.3, 4, 5, 6, 7 Somatic hypermethylation of deoxycytidine within CpG islands in the promoter regions of genes is associated with transcriptional silencing, whereas hypomethylation of CpG islands may result in gene activation. Cancer cell DNA also frequently contains ‘global’ hypomethylation,8, 9, 10, 11, 12 which generally occurs in repetitive sequences, and reduced global levels of DNA methylation have been associated with genomic instability.6, 13
In terms of postpubertal testicular germ cell tumors, prior studies14, 15 uncovered marked differences in DNA methylation patterns in seminomas vs nonseminomatous germ cell tumors. Peltomaki15 examined X chromosomes by Southern blotting after methylation-sensitive restriction enzyme digestion. Although nonseminomatous germ cell tumors usually showed a pattern of hypermethylation, similar to that seen in the inactive female X chromosome, seminomas were usually relatively hypomethylated with a pattern similar to the normally active X chromosome. Two autosomal loci showed a similar trend: seminomas were relatively hypomethylated, whereas nonseminomas were either normally methylated or hypermethylated.15 Smiraglia et al14 used a more global restriction landmark genomic scanning approach to show that seminomas contain much lower levels of 5mC than nonseminomatous germ cell tumors. Others have examined the CpG islands of specific genes (non-X linked) and have generally concluded that nonseminomas tend to be similar to many adult non-germ cell tumors in that they often contain hypermethylation in these regions, whereas seminomas generally lack methylation of these genes.16, 17 In terms of imprinting, which is generally mediated by parental allele-specific deoxycytidine methylation of specific CpG residues, several groups have shown that most postpubertal testicular germ cell tumors show a lack of imprinting at both the H19 and IGF2 loci.18, 19 As primordial germ cells are known to undergo a profound hypomethylation and loss of imprinting during their maturation,20, 21, 22, 23 these studies of global methylation status and imprinting provide strong support for the supposition that testicular germ cell tumors arise from abnormally retained primordial germ cells.1, 14, 17, 24, 25, 26, 27, 28 However, little is known regarding the timing of events of epigenetic changes occurring early during development of testicular germ cell tumors.
Virtually all postpubertal testicular germ cell tumors are thought to arise from intratubular germ cell neoplasia unclassified (IGCNU).1, 29 Smiraglia et al14 have proposed a model whereby seminomas arise from IGCNU cells derived from primordial germ cells that have developed to the point whereby they have undergone 5mC erasure. However, for nonseminomatous germ cell tumors, these authors proposed that they arise from IGCNU cells derived from primordial germ cells that had progressed past this point in their maturation in which they have undergone reprogramming by de novo methylation to contain normal 5mC levels. However, due to limitations of the approaches utilized in previous studies, which entailed DNA isolation from bulk tumor tissues, the methylation status of IGCNU could not be determined.14
Immunohistochemical assessment of 5mC content has been validated as an accurate and potentially advantageous method to evaluate global deoxycytidine DNA methylation compared to more technically demanding methodologies such as in vitro radiolabeled methyl group incorporation assays.10, 30 The primary goal of the present study was to use immunohistochemistry to determine directly relative global DNA methylation levels in IGCNU and to compare those levels with those in pure seminomas, pure nonseminomatous germ cell tumors of various histological types, and mixed seminoma and nonseminomatous germ cell tumors. A secondary goal was to begin to address the possible mechanism of reduced global methylation in seminoma and IGCNU by comparing 5mC levels in these tumors to the expression pattern of the maintenance deoxycytidine DNA methyltransferase, DNMT1. We discuss the present findings in terms of current models of the progenitor cell derivation of testicular germ cell tumor development.
Materials and methods
Patients
This study was approved by our institutional review board. H&E slides and paraffin material, from a total of 55 testicular and extratesticular germ cell tumors diagnosed between 1995 and 2005, were retrieved from the surgical pathology archives of The Johns Hopkins Hospital, USA (48 patients) and Osaka University Hospital, Japan (7 patients). All original diagnoses were confirmed by two pathologists on the study.
Immunohistochemistry
Representative paraffin sections were immunostained using a mouse monoclonal antibody for 5mC (Calbiochem, EMD Chemicals Inc., San Diego, CA, USA) and a rabbit polyclonal antibody for DNMT1. The DNMT1 antibody used was an affinity-purified IgG preparation as described.31 Antigen retrieval consisted of steaming for 40 min in DAKO Target Retrieval Solution (Dako Inc., Carpinteria, CA, USA) for DNMT1 (1:16 000) and 20 min steam in citrate buffer (pH 6.0) followed by treatment in 3.5 N HCl for 15 min at room temperature for 5mC (1:4000). Primary antibody incubations were conducted at room temperature for 45 min. 5mC was detected using the PowerVision+™ immunohistochemistry Detection System from ImmunoVision Technologies Co (Norwell, MA, USA), and DNMT1 was detected using the DakoCytomation EnVision®+ System-HRP (Dako). Histochemical localization was accomplished using 3,3′-diaminobenzidine tetrahydrochloride as the chromogen. After immunohistochemical staining, tissue sections were counterstained with hematoxylin.
To verify antibody specificities the following controls were used: for 5mC we compared staining of the colonic carcinoma cell line HCT116 to a derivative of this cell line in which the DNMT1 and DNMT3b genes were subjected to targeted disruption.32 The clone we used (the original clone 8) was reported to contain approximately 50% of the 5mC levels of the wild-type HCT116 cells as determined by reversed-phase high-pressure liquid chromatography.32 To determine whether we could detect such a decrease we subjected the cell lines to formalin fixation and paraffin embedding to simulate tissue sections. Visual inspection after staining for 5mC showed clearly reduced levels in the knockout cell line (data not shown). In addition, we performed quantitative image analysis using newly developed software33 and found that the overall level of the knockout cells was 62% of the parental cells, indicating a good correlation between the two methods (data not show). Further, to determine whether accessibility of the 5mC antibody or secondary reagents was altered in seminoma cells, we treated tissue sections of a negatively staining seminoma with the prokaryotic CpG-specific methylase from Spiroplasma, M. SssI methylase, which methylates unmethylated deoxycytidines within CpG dinucleotides. M. SssI treatment resulted in a marked increase in staining (data not shown), which was fully dependent upon the addition of the methyl donor, S-adenosyl-methionine (data not shown). This shows that our primary and secondary reagents can access methylated DNA within seminoma cells if indeed 5mC is present.
The distribution and intensity of nuclear staining were visually assessed for each marker as a percentage of cells staining in each of four (0, 1+, 2+ and 3+) intensity categories. 5mC and DNMT1 scores (Σ(intensity score × percentage); possible score range is 0–300) were then calculated. A score was calculated for each histological component in tumors of ‘mixed forms’ in addition to associated IGCNU when present.
Statistic Analysis
Findings were analyzed using the Stata 9.2. (StataCorp, College Station, TX, USA) software package. Equality of population medians among groups was tested using Kruskal–Wallis test for nonparametric one-way analysis of variance by ranks. Pairwise correlation coefficients were calculated to test relationship among variables.
Results
Cohort Characteristics
The histological types of the 55 germ cell tumors are indicated in Table 1. The tumors included 42 germ cell tumors of one histological type (pure forms): 25 seminomas, 11 teratomas, 4 yolk sac tumors and 2 embryonal carcinomas. The remaining 13 germ cell tumors were tumors of more than one histological type (mixed forms). The latter included eight mixed-form tumors containing seminoma and one or more nonseminomatous components, and five mixed tumors containing more than one nonseminomatous component (ie, embryonal carcinoma, teratoma, yolk sac tumor and/or choriocarcinoma). Associated IGCNU was identified in 23 tumors. A total of 40 tumors were primary testicular germ cell tumors. Seven were primary extratesticular germ cell tumors including one suprasellar ‘germinoma’ of the pineal gland, one parasacral and five mediastinal tumors. Seven tumors were metastatic from prior testicular primaries. The location of the remaining lesion was not available. Patient median age was 32 years (range: 0.5–64 years). 5mC and DNMT1 expression in IGCNU, seminomas or nonseminomatous germ cell tumors were not associated with patient age (P≥0.12).
Global Methylation Status (Immunohistochemical 5mC Content)
The 5mC content results are summarized in Table 1. 5mC content was strikingly lower in IGCNU (median 5mC score 0; mean, 18.3; s.d., 64.9) and seminoma (median 5mC score 10; mean, 80.3, s.d., 112) compared to nonseminomatous germ cell tumors (median 5mC score 300; mean, 289.3; s.d., 32.5) (P=0.0001, Kruskal–Wallis). Of 33 seminomas, 14 had no immunohistochemical evidence of 5mC staining compared to the 41 nonseminomatous germ cell tumors that all had strong positive staining (Figure 1a and b).
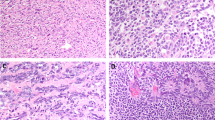
5mC immunostaining in male germ cell neoplasms. (a) Absence of 5mC staining in seminoma ( × 400 magnification). Thick arrow shows seminoma cell nucleus staining with counterstain only. Thin arrows show lymphocytes with strong 5mC staining. (b) Strong and diffuse 5mC staining in a yolk sac tumor ( × 200 magnification).
5mC staining was absent in 20 of 23 IGCNU studied (Figure 2). Of the three IGCNU cases staining positively, only one IGCNU lesion demonstrated strong and diffuse 5mC staining whereas the remaining two lesions had only weak and focal staining. IGCNU 5mC status was strongly correlated with the 5mC status in associated invasive seminomas (pairwise r=0.73, P=0.0004) but not with that of the invasive nonseminomatous germ cell tumors (pairwise P=0.46). In general, diffuse and strong 5mC staining was observed in associated non-neoplastic seminiferous tubules including non-neoplastic spermatogonia. Finally, no significant difference in 5mC levels in IGCNU lesions was seen when grouped according to associated germ cell tumor types (pure seminomas, pure nonseminomatous germ cell tumors and mixed) (P=0.71, Kruskal–Wallis). A summary of 5mC results is graphically depicted in Figure 3.
Combined box/dot plot of 5mC immunohistochemical reactivity scores in 55 male germ cell tumors according to histological type. IGCNU, intratubular germ cell neoplasia unclassified. Other histological types are as indicated. Box shows the interquartile range and the whiskers are 1.5 × the interquartile range. Outliers beyond these ranges are indicated.
DNMT1 Expression
In marked contrast to 5mC levels, all IGCNU and germ cell tumors stained positively for DNMT1 (Table 2). No significant overall difference in DNMT1 expression was detected between IGCNU (median DNMT1 score 200; mean, 204.7; s.d., 54.8), seminomatous (median DNMT1 score 200, mean, 223.5; s.d., 59.0) and nonseminomatous germ cell tumor groups (median DNMT1 score 230; mean, 212; s.d., 91.7; P=0.12, Kruskal–Wallis). However, DNMT1 scores were significantly higher in the embryonal carcinoma subset of nonseminomatous germ cell tumors (median DNMT1 score 300; mean, 284; s.d., 26) compared to seminoma and IGCNU (P=0.037, Kruskal–Wallis). No statistically significant correlation was present between DNMT1 status in IGCNU and associated seminomas (r=0.14; P=0.57, pairwise) nor associated nonseminomatous germ cell tumors (r=0.09; P=0.84, pairwise).
Relationship between 5mC and DNMT1 Status
In the nonseminomatous germ cell tumor group, DNMT1 staining correlated with 5mC status (r=0.46; P=0.035, pairwise). By contrast, no correlation between 5mC and DNMT1 status was present in the seminomas (r=0.32; P=0.07, pairwise) or IGCNU (r=0.02; P=0.9, pairwise).
5mC and DNMT1 Status in Mixed Germ Cell Neoplasm Group
In the subset of tumors containing mixed seminoma and nonseminomatous germ cell tumor components, strikingly lower 5mC was again seen in IGCNU elements (median 5mC score 0; mean, 17.1; s.d., 37.3) and seminomatous elements (median 5mC score 0; mean, 47.5; s.d., 96.6) compared to nonseminomatous germ cell tumor components (median 5mC score 300; mean, 292; s.d., 14.8, P=0.0003, Kruskal–Wallis). 5mC staining was absent in the seminomatous component of five of eight mixed germ cell tumors compared to none of the corresponding nonseminomatous germ cell tumor components (P=0.0004, Kruskal–Wallis; Figure 4). No statistically significant difference in DNMT1 scores was demonstrated between the seminomas and concurrent nonseminomatous germ cell tumor components of mixed tumors (P=0.07, Kruskal–Wallis). 5mC status strongly correlated with DNMT1 expression in the nonseminomatous germ cell tumor components (CC, 1.0; P<0.0001, pairwise) but not in the IGCNU or seminomatous components (P>0.6, pairwise) of the mixed germ cell tumors.
5mC immunostaining in mixed seminomatous and nonseminomatous germ cell tumors. Low power ( × 100) view showing positive 5mC staining in teratomatous component (left half) and negative 5mC staining in seminomatous component (right half). Thin arrow shows teratomatous epithelium, and thick arrow shows seminomas cells.
Discussion
In the current study we used immunohistochemical staining against 5mC to evaluate global methylation in IGCNU and associated invasive testicular germ cell tumors. Our most significant findings were that staining for 5mC was undetectable (or markedly reduced) in the vast majority of IGCNU and in most seminomas, yet there was robust 5mC staining in nonseminomatous germ cell tumors. The lack of staining for 5mC in IGCNU and seminoma was also found in mixed germ cell tumors in which there were both seminomatous and nonseminomatous components. Thus, 5mC staining closely correlated with morphological features of these tumors. This in situ analysis was critical for our finding of profound hypomethylation in IGCNU and seminoma, as we were able to evaluate the relative methylation status of both the neoplastic cells and the nontumor host cells (eg, tumor-infiltrating lymphocytes). The non-neoplastic cells have clearly confounded previous estimates of methylation levels performed by other techniques. Importantly, our staining results are quite consistent with previous work showing markedly lower levels of 5mC in seminoma as compared to nonseminomatous germ cell tumors.15, 16, 17
These results support previous models suggesting that in the majority of cases IGCNU cells are derived from primordial germ cells that have matured/differentiated to the point at which the majority of their 5mC marks have been erased1, 14, 17, 24, 25, 26, 27, 28 (Figure 5), but have not yet matured to the point where the normal developmentally programmed de novo methylation has occurred. Since IGCNU cells appear unmethylated, even when associated with pure nonseminomatous germ cell tumors that are apparently fully methylated, these results suggest that in most cases, nonseminomatous germ cell tumors do not appear to be derived from IGCNU cells that have undergone developmentally programmed de novo methylation.14 Our data show that the absence of detectable 5mC signal is most closely related to the differentiation state of the neoplastic cells (IGCNU and invasive germ cell tumor cells), regardless of the presence of associated seminoma, nonseminomatous germ cell tumor or mixed germ cell tumor components. Thus, we propose a slightly modified model in which seminoma cells, which are morphologically and phenotypically highly similar to IGCNU cells, retain virtual absence of detectable methylation in most cases (Figure 5). And, IGCNU or seminoma cells that differentiate into any other male testicular germ cell tumor type undergo de novo CpG methylation in an apparent attempt to recapitulate the process of normal development (Figure 5). This model allows for both ‘linear’ development of IGCNU to seminoma and then to nonseminomatous germ cell tumors, as well as, ‘nonlinear’ development of IGCNU directly into nonseminomatous germ cell tumors. The fact that a rare IGCNU (1 out of 23 studied lesions) showed diffuse strong 5mC staining suggests that de novo methylation can occur in IGCNU, or rare IGCNU lesions can be derived from methylated progenitor cells as originally suggested by Smiraglia et al.14 Additional studies are needed to determine whether the reason for lack of 5mC in these cells is a failure in de novo methylation or an active demethylation activity in these cells. The mechanisms underlying the frequently observed global hypomethylation in cancers remain poorly understood.34, 35 However, it appears that the mechanisms are different in testicular germ cell tumors compared with somatic tumors as testicular germ cell tumors arise from primordial germ cells that undergo erasure of methylation marks as part of their normal maturation process into mature gametes,20, 21, 22 and cells from which somatic cancers arise are presumably fully methylated. De novo methylation is catalyzed in germs cells by the de novo methyltransferases DNA methyltransferases 3A (DNMT3A) and its cofactor DNMT3-like DNMT3L.23 These established patterns are maintained by DNMT1.23, 36 In the current study the presence of apparently normal levels of DNMT1 in seminoma, nonseminomatous germ cell tumors and IGCNU would suggest a mechanism other than lower expression of DNMT1 to be responsible for our observed overall differences in 5mC levels in these tumors. In the current study, the higher levels of DNMT1 in a subset of nonseminomatous germ cell tumors, namely embryonal carcinoma, and the observed correlation of 5mC and DNMT1 expression only in the nonseminomatous germ cell tumors could point to a role for DNMT1 expression levels in controlling overall 5mC levels in that subset of lesions. In terms of lack of de novo methylation, future studies to examine the levels of the de novo methyltransferases will be important to determine if a lack of these enzymes in IGCNU cells and seminoma cells may be responsible for the lack of their methylation.
Model for global methylation in seminomatous and nonseminomatous germ cell tumors. Primordial germ cells (PGCs) undergo physiological erasure of 5mC during normal development and become PGCs with methylation erasure. These cells normally go on to de novo methylation to become PGCs with de novo methylation marks, and then to spermatogonial stem cells, which are committed to sperm cell differentiation. After birth, these cells continually give rise to maturing and mature sperm cells (gametes). In the development of IGCNU there is abnormal retention of PGCs with methylation erasure that may persist after birth (not shown). These cells are prone to undergo transformation into IGCNU cells in response to environmental insults (lower right). In most cases, it is these cells that give rise to seminomas, which remain unmethylated. In mixed germ cell tumors in which there are seminomatous and nonseminomatous components, the seminoma cells undergo de novo methylation as they differentiate into other neoplastic cell types (eg, yolk sac tumor, teratoma, embryonal carcinoma). In rare cases IGCNU cells may be derived from PGCs that have already undergone de novo methylation in which these cells are prone to undergo development into nonseminomatous germ cell tumors. Gray boxes show neoplastic cells. Dashed boxes indicate transient and/or infrequent nature of the methylated state. Dashed small arrows indicate infrequent events. NS-GCT, nonseminomatous germ cell tumors.
In our model, it is not clear whether methylation occurs first in seminoma cells or rare IGCNU lesions and then differentiation takes place (perhaps as a result of the methylation events) or whether differentiation occurs first. This can be tested in part by forced expression of de novo methyltransferases in seminomas cells37 in culture to determine whether this facilitates differentiation into other germ cell tumor cell types.
In summary, we found strikingly low staining for 5mC in precursor IGCNU lesions associated with testicular germ cell tumors. This marked hypomethylation was present in the majority of IGCNU, regardless of whether these lesions were associated with seminoma, which were similarly hypomethylated, or with nonseminomatous germ cell tumors, which appeared to contain normal levels of methylation. We interpret our findings as supportive of previous modified models of germ cell tumor development in which IGCNU cells arise (in most cases) from a primordial germ cell that had previously undergone normal developmental erasure of methylation marks. Seminomas cells, which are phenotypically highly similar to IGCNU cells, generally retain this lack of methylation whereas nonseminomatous germ cell tumors do not. Additional studies are needed to further explore the molecular basis of the dynamics of these proposed methylation changes and to determine whether abnormal retention of primordial germ cells lacking 5mC predisposes to genetic instability, IGCNU and germ cell tumor development.
References
Reuter VE . Origins and molecular biology of testicular germ cell tumors. Mod Pathol 2005;18 (Suppl 2):S51–S60.
Korkola JE, Heck S, Olshen AB, et al. In vivo differentiation and genomic evolution in adult male germ cell tumors. Genes Chromosomes Cancer 2008;47:43–55.
Baylin SB, Ohm JE . Epigenetic gene silencing in cancer—a mechanism for early oncogenic pathway addiction? Nat Rev Cancer 2006;6:107–116.
Herman JG, Baylin SB . Gene silencing in cancer in association with promoter hypermethylation. N Engl J Med 2003;349:2042–2054.
Ng HH, Bird A . DNA methylation and chromatin modification. Curr Opin Genet Dev 1999;9:158–163.
Laird PW, Jaenisch R . The role of DNA methylation in cancer genetic and epigenetics. Annu Rev Genet 1996;30:441–464.
Baylin SB, Herman JG, Graff JR, et al. Alterations in DNA methylation: a fundamental aspect of neoplasia. Adv Cancer Res 1998;72:141–196.
Soares J, Pinto AE, Cunha CV, et al. Global DNA hypomethylation in breast carcinoma: correlation with prognostic factors and tumor progression. Cancer 1999;85:112–118.
Galusca B, Dumollard JM, Lassandre S, et al. Global DNA methylation evaluation: potential complementary marker in differential diagnosis of thyroid neoplasia. Virchows Arch 2005;447:18–23.
Piyathilake CJ, Bell WC, Jones J, et al. Pattern of nonspecific (or global) DNA methylation in oral carcinogenesis. Head Neck 2005;27:1061–1067.
Piyathilake CJ, Frost AR, Bell WC, et al. Altered global methylation of DNA: an epigenetic difference in susceptibility for lung cancer is associated with its progression. Hum Pathol 2001;32:856–862.
Feinberg AP, Ohlsson R, Henikoff S . The epigenetic progenitor origin of human cancer. Nat Rev Genet 2006;7:21–33.
Eden A, Gaudet F, Waghmare A, et al. Chromosomal instability and tumors promoted by DNA hypomethylation. Science 2003;300:455.
Smiraglia DJ, Szymanska J, Kraggerud SM, et al. Distinct epigenetic phenotypes in seminomatous and nonseminomatous testicular germ cell tumors. Oncogene 2002;21:3909–3916.
Peltomaki P . DNA methylation changes in human testicular cancer. Biochim Biophys Acta 1991;1096:187–196.
Koul S, Houldsworth J, Mansukhani MM, et al. Characteristic promoter hypermethylation signatures in male germ cell tumors. Mol Cancer 2002;1:8.
Okamoto K, Kawakami T . Epigenetic profile of testicular germ cell tumours. Int J Androl 2007;30:385–392; discussion 392.
Mishina M, Ogawa O, Kinoshita H, et al. Equivalent parental distribution of frequently lost alleles and biallelic expression of the H19 gene in human testicular germ cell tumors. Jpn J Cancer Res 1996;87:816–823.
van Gurp RJ, Oosterhuis JW, Kalscheuer V, et al. Biallelic expression of the H19 and IGF2 genes in human testicular germ cell tumors. J Natl Cancer Inst 1994;86:1070–1075.
Trasler JM . Gamete imprinting: setting epigenetic patterns for the next generation. Reprod Fertil Dev 2006;18:63–69.
Schaefer CB, Ooi SKT, Bestor TH, et al. Epigenetic decisions in mammalian germ cells. Science 2007;316:398–399.
Guenatri M, Bourc’his D . In vitro methods of male germ cell specification and differentiation. Med Sci (Paris) 2007;23:619–625.
Reik W . Stability and flexibility of epigenetic gene regulation in mammalian development. Nature 2007;447:425–432.
Clark AT . The stem cell identity of testicular cancer. Stem Cell Rev 2007;3:49–59.
Skakkebaek NE, Rajpert-De Meyts E, Jorgensen N, et al. Germ cell cancer and disorders of spermatogenesis: an environmental connection? APMIS 1998;106:3–11; discussion 12.
Chaganti RS, Houldsworth J . Genetics and biology of adult human male germ cell tumors. Cancer Res 2000;60:1475–1482.
Chaganti RS, Houldsworth J . The cytogenetic theory of the pathogenesis of human adult male germ cell tumors. Review article. APMIS 1998;106:80–83; discussion 83–84.
Houldsworth J, Korkola JE, Bosl GJ, et al. Biology and genetics of adult male germ cell tumors. J Clin Oncol 2006;24:5512–5518.
Dieckmann KP, Skakkebaek NE . Carcinoma in situ of the testis: review of biological and clinical features. Int J Cancer 1999;83:815–822.
Piyathilake CJ, Johanning GL, Frost AR, et al. Immunohistochemical evaluation of global DNA methylation: Comparison with in vitro radiolabeled methyl incorporation assay. Biotech Histochem 2000;75:251–258.
De Marzo AM, Marchi VL, Yang ES, et al. Abnormal regulation of DNA methyltransferase expression during colorectal carcinogenesis. Cancer Res 1999;59:3855–3860.
Rhee I, Bachman KE, Park BH, et al. DNMT1 and DNMT3b cooperate to silence genes in human cancer cells. Nature 2002;416:552–556.
Cornish T, De Marzo AM, Gurel B, et al. FRIDA an open source framework for image dataset analysis. Advancing practice, instruction, and innovation through informatics annual meeting. 19–23 October 2007, Pittsburgh, PA.
Ehrlich M . DNA methylation in cancer: too much, but also too little. Oncogene 2002;21:5400–5413.
Feinberg AP, Tycko B . The history of cancer epigenetics. Nat Rev Cancer 2004;4:143–153.
Goll MG, Bestor TH . Eukaryotic cytosine methyltransferases. Annu Rev Biochem 2005;74:481–514.
de Jong J, Stoop H, Gillis AJ, et al. Further characterization of the first seminoma cell line TCam-2. Genes Chromosomes Cancer 2007;47:185–196.
Acknowledgements
We thank Dr Bert Vogelstein and Dr Stephen Baylin from The Johns Hopkins University School of Medicine for providing the DNMT3a/DNMT1 double knockout cells.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Netto, G., Nakai, Y., Nakayama, M. et al. Global DNA hypomethylation in intratubular germ cell neoplasia and seminoma, but not in nonseminomatous male germ cell tumors. Mod Pathol 21, 1337–1344 (2008). https://doi.org/10.1038/modpathol.2008.127
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2008.127
Keywords
This article is cited by
-
Testicular cancer in mice: interplay between stem cells and endocrine insults
Stem Cell Research & Therapy (2022)
-
Gene expression microarray analysis of adult testicular germ cell tumor: a comparison between pure-type seminomas and seminoma components in mixed tumors
Virchows Archiv (2021)
-
Cell-to-cell variation of chromosomal number in the adult testicular germ cell tumors: a comparison of chromosomal instability among histological components and its putative role in tumor progression
Virchows Archiv (2019)
-
Differences in DNA methylation profiles by histologic subtype of paediatric germ cell tumours: a report from the Children’s Oncology Group
British Journal of Cancer (2018)
-
Incorporating DNA Methyltransferase Inhibitors (DNMTis) in the Treatment of Genitourinary Malignancies: A Systematic Review
Targeted Oncology (2018)