Abstract
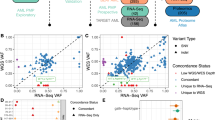
Risk stratification of acute myeloid leukemia (AML) patients needs improvement. Several AML risk classification models based on somatic mutations or gene-expression profiling have been proposed. However, systematic and independent validation of these models is required for future clinical implementation. We performed whole-transcriptome RNA-sequencing and panel-based deep DNA sequencing of 23 genes in 274 intensively treated AML patients (Clinseq-AML). We also utilized the The Cancer Genome Atlas (TCGA)-AML study (N=142) as a second validation cohort. We evaluated six previously proposed molecular-based models for AML risk stratification and two revised risk classification systems combining molecular- and clinical data. Risk groups stratified by five out of six models showed different overall survival in cytogenetic normal-AML patients in the Clinseq-AML cohort (P-value<0.05; concordance index >0.5). Risk classification systems integrating mutational or gene-expression data were found to add prognostic value to the current European Leukemia Net (ELN) risk classification. The prognostic value varied between models and across cohorts, highlighting the importance of independent validation to establish evidence of efficacy and general applicability. All but one model replicated in the Clinseq-AML cohort, indicating the potential for molecular-based AML risk models. Risk classification based on a combination of molecular and clinical data holds promise for improved AML patient stratification in the future.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Grimwade D, Mrozek K . Diagnostic and prognostic value of cytogenetics in acute myeloid leukemia. Hematol Oncol Clin North Am 2011; 25: 1135–1161, vii.
Grimwade D, Hills RK, Moorman AV, Walker H, Chatters S, Goldstone AH et al. Refinement of cytogenetic classification in acute myeloid leukemia: determination of prognostic significance of rare recurring chromosomal abnormalities among 5876 younger adult patients treated in the United Kingdom Medical Research Council trials. Blood 2010; 116: 354–365.
Lazarevic V, Horstedt AS, Johansson B, Antunovic P, Billstrom R, Derolf A et al. Incidence and prognostic significance of karyotypic subgroups in older patients with acute myeloid leukemia: the Swedish population-based experience. Blood Cancer J 2014; 4: e188.
Marcucci G, Haferlach T, Dohner H . Molecular genetics of adult acute myeloid leukemia: prognostic and therapeutic implications. J Clin Oncol 2011; 29: 475–486.
Dohner H, Estey EH, Amadori S, Appelbaum FR, Buchner T, Burnett AK et al. Diagnosis and management of acute myeloid leukemia in adults: recommendations from an international expert panel, on behalf of the European LeukemiaNet. Blood 2010; 115: 453–474.
Patel JP, Gonen M, Figueroa ME, Fernandez H, Sun Z, Racevskis J et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N Engl J Med 2012; 366: 1079–1089.
Papaemmanuil E, Gerstung M, Bullinger L, Gaidzik VI, Paschka P, Roberts ND et al. Genomic classification and prognosis in acute myeloid leukemia. N Engl J Med 2016; 374: 2209–2221.
Bullinger L, Dohner K, Bair E, Frohling S, Schlenk RF, Tibshirani R et al. Use of gene-expression profiling to identify prognostic subclasses in adult acute myeloid leukemia. N Engl J Med 2004; 350: 1605–1616.
Metzeler KH, Hummel M, Bloomfield CD, Spiekermann K, Braess J, Sauerland MC et al. An 86-probe-set gene-expression signature predicts survival in cytogenetically normal acute myeloid leukemia. Blood 2008; 112: 4193–4201.
Li Z, Herold T, He C, Valk PJ, Chen P, Jurinovic V et al. Identification of a 24-gene prognostic signature that improves the European LeukemiaNet risk classification of acute myeloid leukemia: an international collaborative study. J Clin Oncol 2013; 31: 1172–1181.
Marcucci G, Yan P, Maharry K, Frankhouser D, Nicolet D, Metzeler KH et al. Epigenetics meets genetics in acute myeloid leukemia: clinical impact of a novel seven-gene score. J Clin Oncol 2014; 32: 548–556.
Eppert K, Takenaka K, Lechman ER, Waldron L, Nilsson B, van Galen P et al. Stem cell gene expression programs influence clinical outcome in human leukemia. Nat Med 2011; 17: 1086–1093.
Wahlin A, Billstrom R, Bjor O, Ahlgren T, Hedenus M, Hoglund M et al. Results of risk-adapted therapy in acute myeloid leukaemia. A long-term population-based follow-up study. Eur J Haematol 2009; 83: 99–107.
Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N Engl J Med 2013; 368: 2059–2074.
Harrell FE Jr, Lee KL, Mark DB . Multivariable prognostic models: issues in developing models, evaluating assumptions and adequacy, and measuring and reducing errors. Stat Med 1996; 15: 361–387.
Pencina MJ, D'Agostino RB . Overall C as a measure of discrimination in survival analysis: model specific population value and confidence interval estimation. Stat Med 2004; 23: 2109–2123.
Cox DR . Regression models and life-tables. Breakthroughs in statistics. Springer: New York, NY, USA, 1992; pp 527–541.
McGeechan K, Macaskill P, Irwig L, Liew G, Wong TY . Assessing new biomarkers and predictive models for use in clinical practice: a clinician's guide. Arch Intern Med 2008; 168: 2304–2310.
Schroder MS, Culhane AC, Quackenbush J, Haibe-Kains B . survcomp: an R/Bioconductor package for performance assessment and comparison of survival models. Bioinformatics 2011; 27: 3206–3208.
Therneau TM, Grambsch PM Modeling survival data: extending the Cox model: Springer Science & Business Media 2000.
Shanmugam R, Gade P, Wilson-Weekes A, Sayar H, Suvannasankha A, Goswami C et al. A noncanonical Flt3ITD/NF-kappaB signaling pathway represses DAPK1 in acute myeloid leukemia. Clin Cancer Res 2012; 18: 360–369.
Gerstung M, Pellagatti A, Malcovati L, Giagounidis A, Porta MG, Jadersten M et al. Combining gene mutation with gene expression data improves outcome prediction in myelodysplastic syndromes. Nat Commun 2015; 6: 5901.
Radmacher MD, Marcucci G, Ruppert AS, Mrozek K, Whitman SP, Vardiman JW et al. Independent confirmation of a prognostic gene-expression signature in adult acute myeloid leukemia with a normal karyotype: a Cancer and Leukemia Group B study. Blood 2006; 108: 1677–1683.
Acknowledgements
We acknowledge funding from Swedish Cancer Society (Cancerfonden), Swedish e-Science Research Centre (SERC)–‘e-Science for Cancer Prevention and Control (ecpc)’, the Swedish Research Council (Vetenskapsrådet), the Strategic Research Programme in Cancer (StratCan) in Karolinska Institutet and the Stockholm County Council.
Author contributions
MW, JL and MR developed the concept and designed the study, analyzed data and wrote the paper. DK analyzed the data. CN collected and assembled the data. ASM analyzed the data. SL developed the concept and designed the research and wrote the paper. HG developed the concept and designed the research.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website .
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, M., Lindberg, J., Klevebring, D. et al. Validation of risk stratification models in acute myeloid leukemia using sequencing-based molecular profiling. Leukemia 31, 2029–2036 (2017). https://doi.org/10.1038/leu.2017.48
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2017.48
This article is cited by
-
A chest CT-based nomogram for predicting survival in acute myeloid leukemia
BMC Cancer (2024)
-
The exon-junction complex helicase eIF4A3 holds therapeutic potential in acute myeloid leukemia
Leukemia (2024)
-
Identifying an AML Prognostic Model Using 10 Marker Genes from Single-Cell Transcriptome and Bulk Transcriptome Analysis
Biochemical Genetics (2024)
-
Prediction model for drug response of acute myeloid leukemia patients
npj Precision Oncology (2023)
-
Perturbed epigenetic transcriptional regulation in AML with IDH mutations causes increased susceptibility to NK cells
Leukemia (2023)