Abstract
Deregulation of the miR-15a/16-1 cluster has a key role in the pathogenesis of chronic lymphocytic leukemia (CLL), a clinically heterogeneous disease with indolent and aggressive forms. The miR-15a/16-1 locus is located at 13q14, the most frequently deleted region in CLL. Starting from functional investigations of a rare SNP upstream the miR cluster, we identified a novel allele-specific mechanism that exploits a cryptic activator region to recruit the RNA polymerase III for miR-15a/16-1 transcription. This regulation of the miR-15a/16- locus is independent of the DLEU2 host gene, which is often transcribed monoallellically by RPII. We found that normally one allele of miR-15a/16-1 is transcribed by RNAPII, the other one by RNAPIII. In our subset of CLL patients harboring 13q14 deletions, exclusive RNA polymerase III (RPIII)-driven transcription of the miR-15a/16-1 was the consequence of loss of the RPII-regulated allele and correlated with high expression of the poor prognostic marker ZAP70 (P=0.019). Thus, our findings point to a novel biological process, characterized by double allele-specific transcriptional regulation of the miR-15a/16-1 locus by alternative mechanisms. Differential usage of these mechanisms may distinguish at onset aggressive from indolent forms of CLL. This provides a basis for the clinical heterogeneity of the CLL patients carrying 13q14 deletions.
Similar content being viewed by others
Introduction
Chronic lymphocytic leukemia (CLL), the most common form of adult leukemia in the Western world,1 is characterized by a heterogeneous clinical course, essentially dividable into indolent and aggressive forms.2 Deletion of 13q14 is the most frequent genetic aberration associated with CLL3, 4, 5, 6 and represents an early disease-associated event.7, 8, 9 Although generally considered a favorable prognostic factor,10, 11 CLL patients bearing this deletion show heterogeneous clinical outcomes, that can be partially stratified according to 13q deletion subtype.12, 13
In vitro and in vivo studies14, 15, 16, 17 conclusively demonstrate that the miR-15a/16-1 cluster, located at 13q14, controls cell cycle and apoptosis in B cells and that deregulation of these miRNAs contributes to the pathogenesis of CLL. The 13q14 deletion is one of the main causes of the downregulation of the miR-15a and miR-16-1 in CLL, although it should be considered that miR-15a/16-1 expression shows allelic imbalances,18, 19 and that TP53 mutations or deletions affect the transcription and/or maturation of the primiR.4, 15, 20 This cluster is located within the DLEU2 gene, whose expression is controlled by epigenetic mechanisms entailing a monoallelic expression in lymphoid cells from both CLL patients and healthy controls.19 This occurs for the majority of the genes located at this region, suggesting that the miR-15a/16-1 cluster could be regulated in the same way.
In the present work, we studied the region immediately upstream the miR-15a/16-1 cluster. We first investigated a functional SNP, rs115069827, whose minor allele abrogates the maturation of primiR-15a/16-1. We then demonstrated that the region surrounding the SNP functions as an activator of transcription by both RNA polymerase II and RPIII. The RPIII-driven transcription of primiR-15a/16-1 from this cryptic activator is allele-specific and, in our CLL patients, results dominant in the cases with 13q14 deletions that are characterized by high expression of ZAP70, a predictor of poor prognosis.
Materials and methods
Primary cells, cell lines and cell culture
CLL samples were obtained from patients enrolled in the CLL Research Consortium and ‘Policlinico Agostino Gemelli’, ‘Università Cattolica del Sacro Cuore’, Rome, on written informed consent in accordance with the Declaration of Helsinki. The study protocols were approved by the Institutional Review Boards of The Ohio State University and the ‘Policlinico Agostino Gemelli’. The participating institutions provided the clinical data associated with each patient at the time of sample collection. Among patients, there was a rare case of monozygotic twin sisters that share a germline single nucleotide variation in ARLTS1.21 The samples were analyzed to determine expression of ZAP70, immunoglobulin heavy chain variable mutational status and karyotype, as previously described.22 Peripheral blood mononuclear cells (PBMC) were isolated by density gradient centrifugation with Ficoll-PaquePlus (Amersham Biosciences, Uppsala, Sweden). Cell lines WaC3CD5 (WAC),23 MEC-2, MEC-1,24 HEK293T, K562 (from ATCC) and CLL cells were cultured in RPMI1640, supplemented with 10% fetal bovine serum of Australian origin, 2 mmol/l L-glutamine, 100 units/ml of penicillin and 100 units/ml streptomycin in a humidified incubator containing 5% CO2 at 37 °C. All the cell lines were authenticated by short tandem repeat analysis (Promega, Fitchburg, WI, USA, PowerPlex16HS) comparing the results with the on-line databases (www.lgcstandards-atcc.org/en/STR_Database.aspx?slp=1; www.dsmz.de/fp/cgi-bin/str.html; http://strdb.cogcell.org/) (Supplementary Table S5). The STR profile of WaC3CD5 was not in the databases. MEC-1 and MEC-2 cell lines, which originate from the same patient and share the same STR profile, were authenticated by karyotype analysis (Supplementary Table S5). The densities of cultured cells were maintained between 1 and 3 million/ml. Epstein–Barr virus infection of CLL cells was conducted as described by Doyle et al.25
Cell treatments, transfection and luciferase assay
The RNAPII inhibitor, α-amanitin (A2263 Sigma, St Louis, MO, USA) was dissolved in bi-distilled sterile water and filtered by 0.2 μm filter. The experiments were performed by using growing doses of a-amanitin, that is, 1, 2, 3, 5, 10 μg/ml on each point of 2000.000 cells/ml or 2 and 10 μg/ml on each point of 500 000–1000 000 cells/ml. After 16 h, the cells were lysed in 700 μl of Qiazol (Qiagen, Venlo, Netherlands) for the RNA extraction.
Transfection of DNA vectors (200–500 ng/50 000 cells) in HEK293T was performed with Lipofectamine2000 (Invitrogen, Carlsbad, CA, USA) in accordance with the manufacturer’s procedures, whereas transfection of MEC-1, MEC-2, WaC3CD5 and K562 was performed using the Amaxa nucleofector (Lonza, Basel, Switzerland) according to the manufacturer’s protocols (plasmids:3000 ng/1 000 000 cells; small interfering RNA 40 pmol/1 000 000 cells). siPOLR3A (sc-90684), siPOLR3G (sc-43507) and small interfering RNA -A as control (sc-37007) were purchased from Santa Cruz Biotechnology (Santa Cruz, CA, USA). The efficiency of transfection was determined by cytofluorometry, messenger RNA expression and\or fluorescence microscopy. After transfection of luciferase reporter vectors (pGL3Ren-187_A and pGL3Ren-187_G), the luciferase assay was conducted according to manufacturer’s dual luciferase assay protocol (Promega).
Cell staining for flow cytometry
Cell staining was performed as already reported.26, 27 Briefly, 5 × 105 cells per sample were washed, resuspended in 100 μl of surface antibody mix (1 μl/sample of anti CD5-PE and 1 μl/sample of anti-CD19-PECy5) and incubated for 30 min at 4 °C in the dark. Cells were washed and stored at 4 °C in the dark until acquisition. Phycoerythrin-conjugated anti-CD5 (CD5-PE) and PE-cyanin 5-conjugated anti-CD19 (CD19-PECy5) were purchased from Becton Dickinson (BD, Franklin Lakes, NJ, USA). A FACSCantoII flow cytometer, running with FACSDiVa software (BD), was used for cell analysis; 10 000 events for each sample were recorded. CompBeads (BD) as well as the antibodies used in the assay, as single stained tubes, were utilized to assess fluorescence compensation. Non-specific fluorescence was evaluated using fluorescence minus one controls.27 Data were analyzed using the softwares FACSDiVa (BD) and FlowJo (TreeStar, Ashland, OR, USA).
DNA extraction, sequencing and copy number variation
Genomic DNA was isolated from cell lines and PBMC from CLL samples by standard treatment with Hirt’s solution in the presence of 200 μg/ml of proteinase K, followed by phenol/chloroform/isoamyl extraction and ethanol precipitation. The DNA regions of interest were amplified with primers detailed in Supplementary Table S4 and sequenced using the BigDye Terminator v3.1 Cycle Sequencing Kit (Life Technologies, Waltham, MA, USA) according to the manufacturer’s specification, with an ABI3130xl Genetic Analyzer (Life Technologies). Copy number variation was performed by real-time PCR using the TaqMan technology. Briefly, 10 ng of DNA was amplified using specific primers and fluorescent probes and a standard internal reference (TaqMan RNase P assay, Life Technologies) Each sample was analyzed in triplicate on 7900HT Real-Time PCR system. Each well was normalized to RNase P to obtain a ΔCt (2−(FAM dye Ct−VIC dye Ct)), and then the average ΔCt for each triplicate was calculated. All samples were then normalized to a calibrator (eight DNA samples from healthy patients) to determine ΔΔCt. As the percentage of B-CLL cells could vary among patients, samples where considered carrying 13q deletion if loss of genetic material at the 13q chromosome region was ⩾25% compared with normal control.
RNA extraction and quantitative real-time PCR
RNA was extracted using the miRNeasy MicroKit (Qiagen), treated with the RNase-Free DNase Set (Qiagen) according to the instructions of the manufacturer and quantified with Nanodrop2000 (Thermo-Scientific, Waltham, MA, USA). RNA quality was analyzed using the Experion RNA HighSens Analysis Kit (Bio-Rad, Hercules, CA, USA). RNA quality indicator was on a median average of 8.15. Gene sequences were from NCBI (http://www.ncbi.nlm.nih.gov/), RNA sequences for 18S, DLEU2, were used into the Universal ProbeLibrary (UPL) Assay Design Center software (https://www.rocheappliedscience.com/sis/rtpcr/upl/index.jsp?id=uplct_030000) to identify primers and UPL probes (Supplementary Table S4). Total RNA (50–200 ng) was retrotranscribed with High Capacity cDNA Reverse Transcription Kit (Life Technologies). The primiR exogenous expression was assessed with the SYBR green technology (Qiagen), while the endogenous expression of the primiR was performed with TaqMan assay (Life Technologies). To test DNAse treatment efficiency, cDNA synthesis was performed without the enzyme reverse transcriptase (noRT-PCR).
MicroRNAs quantization was performed using stem loop RT primers designed28 with a modification to include the UPL #21 sequence binding site.29 UPL probe #21 was from UPL database (Roche Diagnostics, Basel, Switzerland). Total RNA (50 ng) was retrotranscribed with TaqMan Micro-RNA Reverse Transcription Kit (Life Technologies). Reactions were incubated 30 min at 16 °C, followed by pulsed RT of 60 cycles at 30 °C for 30 s, 42 °C for 30 s and 50 °C for 1 s.30 Real-time PCRs were performed in an Applied Biosystems 7900 instrument. MiRNA and messenger RNA levels were measured using Ct (threshold cycle). Target amount, normalized to an endogenous references 18S/RNU44 and relative to a calibrator, is given by 2−ΔΔCt and/or 2−ΔCt methods (Life Technologies).
RNA-binding protein immunoprecipitation
RIP was performed with Magna RIP kit (Millipore, Billerica, MA, USA) according to the manufacturer’s instruction. Cells transfected with vectors pCDF1070_A or pCDF1070_G were lysed and the RNA-associated proteins were immunoprecipitated (IP) with anti-DROSHA (Novus Biologicals, Littleton, CO, USA, NBP1-03349). The precipitated RNA was retrotranscribed and measured by quantitative real-time PCR (SYBR green, Qiagen) and referenced to the non-IP control (Input). The Magna RIP kit does not include a DNase step before RT-PCR, thus, given that our experimental conditions required plasmid transfection, the evaluation of the IP primiR-15a/16-1 was calculated taking into account plasmid contamination: IP_(DROSHA_primiR)=(2-(Ct IP_DROSHA_primir-Ct INPUT_DROSHA_primir))/(2-(Ct noRT_DROSHA_PrimiR-Ct INPUT_DROSHA_primir)).
Chromatin immunoprecipitation (ChIP)
ChIP was carried out with the Magna ChIP-A/G kit (Millipore) according to the manufacturer’s instruction. Briefly, transfected K562 cells, treated with formaldehyde, were sonicated after nuclear protein extraction. Nuclear extracts were IP with anti-POL II (CTD4H8, Santa Cruz sc-47701), as positive control, and anti-POL III (RPC32, Santa Cruz sc-28712). Specific DNA enrichment was measured by real-time PCR (SYBR green, Qiagen) and referenced to the non-IP control (Input). The experiments were performed in duplicate.
Results
A single nucleotide variation upstream the miR-15a/16-1 cluster in CLL patients
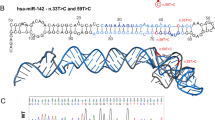
Because of the importance of the miR-15a/16-1 cluster in CLL, we sequenced ∼1000 nucleotides around this genomic region in 249 DNA samples from PBMCs of CLL patients. We found A–G heterozygosis (rs115069827) in DNA from two CLL patients, CLL479 and CLL585 (Table 1, Supplementary Figure S1a). This polymorphism is located within the region immediately upstream of miR-15a (Figure 1). To complete the genetic analysis of these samples, we confirmed germline heterozygosity of CLL479 and CLL585 by analyzing DNA from buccal swab of the two patients.
Schematic representation of the DLEU2/miR-15a/16-1 locus. (a) Genomic structure of the DLEU2/miR-15a/16-1 locus from the reference NC000013 genomic sequence (NCBI). Telomeric (Tel) and centromeric (Cen) position, genomic positions and structure of DLEU2, FLJ31945, DLEU1, miR-15a/16-1, KCNRG and TRIM13, exons (black bars), start site and direction (arrows), the SNP rs115069827 (red triangle), the second promoter of DLEU2 responsive to E2F1 and CMYC transcription factors (gray box),58, 59 the minimal deleted region (gray arrow), U59 DLEU2 probe for copy number variation and gene expression analysis (black triangles) are shown. (b) Sequence of the genomic fragment (1070 bp) analyzed for nucleotide variants in 249 CLL patients and cloned for exogenous expression analysis. Underlined the 187 bp sequence investigated for transcription activity, copy number variation and RNA expression; the rs115069827 SNP is gray labeled; in bold and italic the stem loops of the miR-15a and miR-16-1.
The genomic region upstream the miR-15a/16-1 cluster acts as its transcriptional activator and is not affected by the SNP rs115069827
To determine whether the A–G SNP affected miR-15a/16-1 expression, we studied the genomic region encompassing the variant allele. At first we speculated that this region could act as transcriptional activator of the candidate miRNAs and that the G and A variants differently influence miRNAs expression. To test this, PCR-amplified fragments (187 bp), comprising either the A or the G variants, were obtained from CLL479 and subcloned into a modified version of the pGL3 basic vector including Renilla luciferase as internal control for transfection efficiency. HEK293 and MEC-02 cells were transiently transfected with the generated constructs (pGL3Ren-187_A and pGL3Ren-187_G). The results indicated strong promoter activity of the subcloned fragments, with no differences between the two variants (Figure 2a). To assess the primiR expression, we then cloned a broad region (1070 bp), including the entire cluster and the putative promoter with either the A or the G variant (Figure 2b), in the lentiviral expression vector lacking the constitutive promoter (pCDF1070_A and pCDF1070_G). The vectors were transiently transfected into K562 and WAC cells. We noted a threefold induction of exogenous primiR-15a/16-1 expression compared with the controls (empty vector), irrespective of the variant. The results were more convincing in the WAC cell line, in which only exogenous primiR was detectable, as these cells lack both alleles of the miR-15a/16-1 cluster locus. The primiR was normalized on GFP expression, to exclude differences in transfection efficiency, and the effective DNase treatment was tested by PCR in the RNA samples (noRT, Supplementary Figure S2).
The genomic region upstream the miR-15a and 16-1 cluster acts as promoter of transcription activity. (a) Relative luciferase activity of the 187 bp genomic fragments harboring the two variants of the SNP rs115069827 (pGL3Ren-187_A and 187_G) after 24 h of transfection in HEK293 and MEC-02 cell lines. The pGL3Ren empty vector was used as control. (b) Induction of primiR-15a/16-1 expression in K562 and WaC3CD5 (WAC) cell lines after 24 h transfection of pCDF-1070A and pCDF-1070G vectors. RNAs were previously treated with DNase to avoid genomic/plasmid contamination that was further investigated by qPCR for the GFP plasmid DNA fragment (data not shown). The primiR-15a/16-1 expression was normalized on the expression of the internal reporter gene GFP. The pCDF vector used (SBI) is lacking of the CMV promoter. The pCDF empty vector was used as control. P value >0.05 was considered not significant, between 0.01 to 0.05 (*) significant, between 0.001 to 0.01 (**) very significant and <0.001 (***) extremely significant.
These findings suggest that the regulation of the primiR-15a/16-1 can occur independently of its host gene DLEU2. To corroborate these results, we correlated the expression of miR-15a and miR-16 with that of DLEU2 in two subgroups of patients with different CLL karyotype and prognosis (13q deletion and 11q deletion).31 Respective Spearman correlation factors (ρ) for miR-15a and miR-16 were 0.55 (P=0.0095) and 0.47 (P=0.031) for all the CLL cases; 0.83 (P=0.0008) and 0.86 (P=0.0003), for the CLLs with the 13q deletion; conversely, −0.166 (P=0.67) and −0.3 (P=0.43) for the CLLs with 11q deletion (Supplementary Figure S3). This evidence further suggests that the miR-15a/16-1 has its own transcriptional activator in the immediate 5′ flanking region and that, according to CLL karyotype, its regulation is related or unrelated to DLEU2 expression.
The SNP rs115069827 affects the maturation of miR-15a
As the SNP rs115069827 does not affect the strength of the newly identified transcriptional activator, we investigated its role in the processing of the miR-15a. We evaluated the expression of the primiR and the miR-15a in PBMCs from CLL479 and CLL585 (both heterozygous for rs115069827_SNP), from 5 CLL cases with normal karyotype and from five healthy donors. Among these samples, CLL479 showed the highest primiR expression level, whereas the expressions of the mature forms of miR-15a and miR-16 were significantly downregulated, in line with that of the other CLLs (Figure 3a), according to the published data.4 In matched samples, the median ratios between miR and primiR expression in the CLLs and in the normal samples were respectively 0.002 and 0.028 (P=0.017) for miR-15a, versus 1.72 and 8.34 for miR-16 (Figure 3b, Supplementary Table S1); hinting at a strong relevance of impaired miR-15a/16-1 processing in CLL. The ratio for miR-15a was even lower (0.00047) in CLL479, suggesting that in this specific case, defective maturation was the main cause of miR-15a downregulation. In this analysis, the expression of miR-16 was not considered, as it is also transcribed by a different genomic locus. Of note, CLL585, also positive for the SNP, showed a miR-15a/primiR ratio similar to that of CLL479 (0.00076) (Supplementary Table S1). CLL585 carries a homozygous deletion of the miR-15a/16-1 locus in B cells and hence, the detected miR and primiR must have originated from non-B cells. Taken together, these data support the involvement of rs115069827SNP in the maturation steps of miR-15a, in both leukemic and healthy cells. To investigate this, we evaluated the expression of the mature form of the miR-15a in K562 and WAC cells transfected with either pCDF-1070_A or _G. In both cell lines, the expression of the miR-15a from the vector with the G variant was lower than that from the vector with the A variant (Figure 3c), in spite of the allele type independent mode of expression of the exogenous primiR-15a/16-1 (Figure 2b). For miR-16-1, similar results were obtained in B cells derived from the transgenic CLL mouse model Eμ-TCL132 and infected with lentivirus particles transducing the 1070 bp region with either the A or the G allele (Supplementary Figure S4). SNPs in the flanking non-structured RNA sequences of microRNAs can change the hairpin structures of the primiR, interfering with the binding of proteins necessary for their processing to the mature forms.5, 33, 34, 35, 36, 37 Given that in mice the DEAD-box helicase subunits of the Drosha complex (p68 and p72) are required for the processing of primiR-15a/16-1 to mature miR-16-1,38 we evaluated the efficiency of the interaction between the DROSHA complex and the primiR-15a/16-1 in K562 cells transfected with the pCDF-1070_A or _G vectors. By RIP analysis, the G allele showed reduced DROSHA binding capacity when compared with the A allele (Figure 3d).
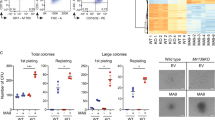
The minor allele of the SNP rs115069827 affect the miR-15a maturation by impairing the primiR/Drosha binding complex. (a) Expression analysis of primiR-15a/16-1 and mature miR-15a in 7 CLL and five healthy donor (N) PBMC samples. Data were normalized to RNU6B and RNU44 expression. (b) Graphics represent the relative efficiency of maturation of miR-15a calculated by the equation 2−ΔCtmiR-15a/ 2−ΔCtprimiR-15a/16-1. (c) qRT–PCR of the mature form of miR-15a in K562 and WAC cell lines after 24 h of transfection with the pCDF-1070_A/G vectors. (d) primiRNA 15a/16-1 quantification of the IP RNA with the anti-DROSHA antibody in K562 cells transfected with the pCDF-1070_A/G vectors (RIP). Data were normalized on the primiRNA 15a/16-1 expression of the RNA not IP (INPUT). Transfection efficiency was counted by GFP-positive cells determined by fluorescence-activated cell sorting analysis. P value >0.05 was considered not significant, between 0.01 to 0.05 (*) significant, between 0.001 to 0.01 (**) very significant and <0.001 (***) extremely significant.
RPIII is involved in the transcription of the primiR-15a/16-1
During the course of our experiments, we found that the 107 bp region immediately upstream the miRNAs stem loops acts as transcriptional activator of miR-15a/16-1 locus and is transcribed as well. This is explainable considering either RPIII occupancy at this region or a mechanism of transcription independent of the RPI/RPII/RPIII, as recently described.39 We first investigated the involvement of RPIII. We tested the primiR-15a/16-1 expression in K562 cells transfected with either the pCDF_1070A/G vectors or the control after treatment with the RPII inhibitor α-amanitin. The left side of Figure 4a shows that the RPII takes part in the transcription of the primiR-15a/16-1 derived from the pCDF_1070A/G vectors. However, as these pCDF vectors hold an RPII promoter (EF1, upstream the GFP reporter gene) we could not exclude its influence at this region and/or a cooperation between RPII and RPIII transcription.40, 41 Therefore, we performed a similar experiment after subcloning the 1070 bp fragments within the pCR2.1 vector, to obtain pPCR2.1-1070 A/G vectors. We chose this plasmid as it does not contain any eukaryotic promoters and/or regulatory sequence. Treatment with α-amanitin significantly reduced the endogenous ACTB pre-messenger RNA used as control in the transfected cells (Supplementary Figure S5a), but did not reduce the exogenous primiR-15a/16-1 expression (Figure 4a), suggesting the existence of an RPII-independent mechanism of transcription. Controls show that plasmid contamination, although not completely removed, was very low and irrelevant to the experimental results (Supplementary Figure S5b). Similar results were observed by measuring the expression of the miR-15a (Supplementary Figure S5c and d). To further support RPIII involvement in primiR-15a/16-1 transcription, we silenced two polymerase III subunits (POLR3A and POLR3G) in K562 cells, to inhibit the RPIII activity. POLR3G is a specific subunit of the RPIII complex, but its silencing does not prevent RPIII elongation,42 while POLR3A is specifically implicated in the elongation of the RPIII transcripts.43 At 48 h from silencing, the cells were transfected with the pCR2.1_1070_A vector to induce primiR transcription by RPIII. Silencing of the POLR3A gene reduced the expression of the primiR and of tRNAtyr (transcribed by RPIII), while the expression of the ACTB pre-messenger RNA, transcribed by RPII, was not affected; (Figure 4b). Moreover, by ChIP analysis, we also observed slight binding of RPIII to the new transcription activator region, apparently independent of the SNP allele (Figure 4c). Finally, we performed ChIP analysis with the RPIII RPC32 subunit on the MEC-1 B-CLL cell line24 transfected with the pCDF_1070_A vector and treated or not with α-amanitin. Given that the pCDF_1070 s vectors drive primiR-15a/16-1 expression by RPII (Figure 4a), we tested RPIII occupancy after inhibition of RPII transcription. We registered twofold primiR-15a/16-1 DNA enrichment only in cells treated with α-amanitin (Figure 4d). A similar experiment was conducted in K562 cells by measuring the endogenous primiR-15a/16-1 expression. K562 cells harbor a monoallelic 13q deletion and show RPII-dependent transcription of primiR-15a/16-1 (Supplementary Figure S6d). The ChIP analysis shows a primiR-15a/16-1 DNA enrichment only in K562 cells treated with α-amanitin (Figure 4e). These results confirm RPIII involvement in a different cell line and support the participation of RPIII in primiR-15a/16-1 transcription only when RPII activity is prevented or inhibited.
RPIII is involved in the transcription of primiR-15a/16-1. (a) PrimiR-15a/16-1 expression in K562 cells transfected with the pCDF1070_A/G (left side) and pCR2.1_1070_A/G vectors (right side) with and without α-amanitin treatment (2 μg/ml). (b) PrimiR-15a/16-1, preACTB, tRNAtyr, POLR3A and POLR3G relative expression (normalized on 18S expression) in K562 cells silenced for polymerase III subunits (POLR3A, POLR3G) and then transfected with the pCR2.1_1070_A vector. (c) ChIP analysis for RPIII occupancy in the primiR-15a/16-1 and tRNAtyr (as control) DNA regions by qPCR in K562 cells transfected with pCR2.1_1070_A/G vectors. Controls are represented by the pCR2.1 Empty vector (e) and the non-specific unconjugated rabbit polyclonal antibody immunoglobulin G IP. Pre 15-16 and Pri 15-16 are two different genomic DNA fragments of the miR-15a/16-1 cluster region (Supplementary Table S4). (d) ChIP analysis for RPIII occupancy in the primiR-15a/16-1 DNA region by qPCR in MEC-1 cells transfected with pCDF_1070_A vector with and without α-amanitin treatment. Controls are represented by the non-specific unconjugated rabbit polyclonal antibody immunoglobulin G IP and as there is not primiR-15a/16-1 DNA enrichment in the non α-amanitin-treated MEC-1 cells, the amplification of the DNA IP with RPIII could be considered a specific signal. (e) ChIP analysis for RPIII occupancy in the primiR-15a/16-1 and tRNAtyr (as control) DNA regions by PCR in nontransfected K562 cell line. P value >0.05 was considered not significant, between 0.01 to 0.05 (*) significant, between 0.001 to 0.01 (**) very significant and <0.001 (***) extremely significant.
Allele-specific transcription of primiR-15a/16-1
To assess the existence of an RPII-independent mechanism at the candidate region in CLL patients, we evaluated the expression of the primiR-15a/16-1 in PBMCs from CLL patients upon α-amanitin treatment. To evaluate one allele at the time, we chose a subset of 31 cases (48% of the total analyzed patients) in which the 13q region showed macrodeletions (by FISH) and/or microdeletions (by copy number variation analysis of the DLEU2/miR-15a/16-1 region, Supplementary Table S2, Supplementary Figure S1b). Regulation of primiR-15a/16-1 was respectively scored as RPII dependent, in the samples that upon treatment shared a trend similar to that of preACTB and/or preGAPDH and opposite to that of the tRNAtyr gene, or RPIII-dependent, in the samples where primiR expression was either unchanged or increased after α-amanitin treatment (Supplementary Figure S6). Twenty out of 31 samples were informative on the basis of technical controls. The results showed that in 8/20 cases (40%), the regulation of primiR-15a/16-1 was RPIII-dependent, while in the other 12 cases (60%) was driven by RPII (Table 2), suggesting that the two alleles of this locus are differentially regulated. As we could not distinguish whether the RPIII involvement arises from pathologic and/or physiologic mechanisms, we decided to treat PBMCs from the patient CLL585 with α-amanitin. This patient has a bi-allelic deletion of the miR-15a/16-1 locus in B-CLL cells, but it still shows heterozygosis of the SNP rs115069827 at both DNA and RNA levels, likely arisen from the non-B cells. Therefore, α-amanitin is predicted to affect only the ‘normal’ non-B PBMC population. By sequencing, we demonstrated selective and progressive inhibition of the expression of the A allele in an α-amanitin dose-dependent manner, while the expression of the G allele was unchanged (Figure 5). Overall, these data support allele-specific regulation of the primiR-15a/16-1: one allele is most likely controlled by the DLEU2 promoter/s, whereas transcription of the other allele occurs through the recruitment of RPIII to the newly identified transcription activator region.
Allele-specific expression of the primiR-15a/16-1 in CLL585. (a) PBMC from CLL585 were treated with α-amanitin at 2 and 10 μg/ml for 17 h. The arrow indicates the variant A–G that shows dose-dependent downregulation of the minor allele G after the α-amanitin treatment. (b) primiR-15a/16-1 relative expression in CLL585 normalized on the 18S expression (RNAPI) after α-amanitin treatment.
Allele-specific transcription of primiR-15a/16-1 is associated with ZAP70 expression in CLL
In the tested cases, we noted that the newly identified RPIII-dependent mechanism predominated in CLLs with high expression of ZAP70 (Fisher’s exact test P=0.019). Indeed, most of the samples with high ZAP70 (6 out of 8) showed transcription of the primiR-15a/16-1 by RPIII, conversely in the cases negative for ZAP70, primiR-15a/16-1 transcription was mostly regulated by RPII (10 out of 12 CLL patients) (Table 2, Supplementary Figure S6). Notably, we had informative monozygotic twins with different clinical course of the disease: CLL3tw, aggressive, and CLL4tw, indolent (Supplementary Table S3). CLL cells from these twins were immortalized and treated with α-amanitin. Upon treatment, we noted transcription of the primiR-15a/16-1 respectively mediated by RPIII in cells from CLL3tw (aggressive CLL) and by RPII in cells from CLL4tw (indolent CLL) (Supplementary Figure S6b, Table 2). To evaluate at molecular level the differences between B-CLL cells in which the mir-15a/16-1 cluster is transcribed by RPII versus RPIII, we measured the levels of miR-15a, primiR-15a/16-1 as well as TP53, BCL2 and Cyclin E proteins, direct targets of miR-15a, in our B-purified CLL samples. Although we did not find changes in the expression of primiR-15a/16-1 and/or miR-15a (data not shown), we registered differences in the protein levels of BCL2, TP53 and Cyclin E with decreasing trend in the CLL samples in which the primiR-15a/16-1 was transcribed by RPIII (Supplementary Figure S7); suggesting that RNPII/RNPIII regulation of miR-15a/16-1 affects the target genes. Considering ZAP70 status, a similar trend was observed only for TP53, while for BCL2 the protein levels were differently distributed between the two groups (F-test, P=0.0014).
In our survey (training set n=36, validation set n=18) there were samples with 13q deletions that did not include the miR-15a/16-1 locus. Almost all of these cases showed high expression of ZAP70 (Fisher’s exact test: training set P<0.001; validation set P=0.006; Supplementary Table S2) and, in those whose cells were available, despite the presence of both alleles of miR-15a/16-1, we detected a RPIII-dependent primiR regulation. Therefore, we performed copy number variation analysis of a more telomeric DNA region (∼30 kb, U59 DLEU2 region), that encompasses the second DLEU2 promoter. We found monoallelic deletion of this region, resulting in the loss of the primiR regulation by the RPII-dependent DLEU2 promoter.
The molecular model of the DLEU2/miR-15a/16-1 locus we propose is recapitulated in Figure 6, that illustrates the genetic characteristics and their associations with ZAP70 status.
Model of transcriptional regulation of miR-15a/16-1 cluster. On the left, transcriptional regulation of miR-15a/16-1 in normal cells is modeled. The miR-15a/miR-16-1 expression is driven by two different mechanisms specific for the two alleles. Allele A shows the transcription of DLEU2 gene and therefore, the miR-15a/16-1 cluster by an RPII-dependent mechanism. Conversely, the allele B shows DLEU2 expression epigenetically repressed and consequently the miR-15a/16-1 expression is driven by an RPII-independent mechanism. Immediately upstream, the miR-15a, the newly identified transcriptional activator region engages the RPIII to start the transcription of the miR-15a/16-1 cluster. 13q deletions in CLL patients (right side) lead to the expression of miR-15a/16-1 by RPII and/or RPIII, depending on the deleted allele. Deletion of the region encompassing the miR stem loops on allele A occurs in ZAP70− and ZAP70+ CLL, whereas the deletion of allele B mostly occurs in CLL patients with low ZAP70 expression. In our model, the expression of miR-15a/16-1 cluster in ZAP70+ CLLs is driven by RPIII by two different deletions of the allele A: deletion of the stem loop cluster transcribed by RPII or deletion of the regulatory elements of the active DLEU2 gene.
Discussion
The minimal deleted region of 13q14 starts about 20 kb upstream to the cluster encompassing the second DLEU2 promoter3, 44 and most genes localized in this critical region are downregulated by a factor >2 in the CLLs with monoallelic 13q14 deletions. The miR-15a/16-1 cluster is located within the DLEU2 gene and, monoallelic DLEU2 expression has so far been demonstrated in 20–40% of B and T cells from healthy donors,19 suggesting that there could normally be monoallelic expression of the miR cluster as well. As miR-15a and miR-16-1 are strongly implicated in the pathogenesis of the disease,14, 15, 16, 17 we aim to clarify the transcriptional regulation of the miR-15a/16-1 cluster in CLL. Here, we demonstrate that the transcription of the cluster is regulated simultaneously by alternative mechanisms that use either RPII or RPIII in allele-specific manner. The loss of the transcriptional regulation of primiR-15a/16-1 by RPII is associated with high expression of ZAP70, a poor prognostic feature of CLL patients.
We reached these results through characterization of the SNP rs115069827, identified in two CLL patients at about 100 bp upstream the miR-15a stem loop, able to abrogate the maturation of the primiR-15a/16-1. This SNP is epidemiologically irrelevant for CLL, because of its rarity, but allowed us to demonstrate the existence of allele-specific effects on the transcription and, consequently, the expression of miR-15a. In fact, the chromosome region immediately upstream the miR cluster, once isolated from other eukaryotic regulatory elements, was able to promote RNA transcription by an RPII-independent, allele-exclusive and RPIII-engaging mechanism. This novel mechanism is consistent with the results of an independent study demonstrating that the primiR-15a/16-1 RNA may or may not present the characteristic RPII 5′cap structure.45 RPIII-dependent transcription of the miR-15a/16-1 cluster and defective miR processing are unmasked in an allele-specific manner by the 13q14 deletion.
RPII and RPIII share several transcriptional elements,40, 46, 47 but it is well known that there are important differences in their transcriptional regulation.48 Increase in RPIII activity has been observed in cancers.49 Moreover, RPIII activity is linked to the transcription factors MYC, P53 and RB1,50, 51, 52 whose deregulation have been associated with poor prognosis in several cancers, including CLL.53, 54, 55, 12 These genes control cell cycle and apoptosis in concert with miR-15a/16-1.50, 51, 52, 56, 57 Therefore, we sought to determine the clinical correlations associated with RPII versus RPIII regulation of the miR cluster in our case series. Although the follow up did not allow direct prognostic correlations, we identified a significant association between RPIII-dependent regulation of primiR-15a/16-1 and high expression of ZAP70, a predictor of poor prognosis in CLL.22 Notably, we analyzed a rare case of familial CLL in monozygotic twins that differed in ZAP70 status and clinical features (aggressive versus indolent disease).22 Transcription of primiR-15a/16-1 was driven by RPIII in the aggressive ZAP70-positive CLL patient, but by RPII in the indolent and ZAP70-negative case. We also identified a significant correlation between type of genomic deletion and ZAP70 expression status: almost all the CLLs cases with 13q deletions that retained two copies of the miR-15a/16-1 cluster, but lacked a more telomeric region, showed high ZAP70 expression. Higher miR-15a expression has previously correlated with ZAP70 positivity in CLLs.5 We found RPIII-dependent regulation of the primiR in almost all ZAP70-positive cells, but no difference in miR-15a expression between the RPII- and RPIII-dependent groups was observed, probably because the majority of the patients in our subset had an indolent disease at the time of analysis. However, differences in the levels of P53, BCL2 and Cyclin E proteins, key targets of miR-15a and miR-16-1, were observed between CLL cells with RPIII- versus RPII-mediated transcription of the primiR-15a/16-1.
In conclusion, both alleles of the miR-15a/16-1 cluster are normally transcribed, one by RPII together with DLEU2, which is often expressed monoallelically, the other one by RPIII, independently of the host gene. Being the DLEU2/miR-15a/miR-16-1 locus finely regulated by cell cycle-related transcription factors,15, 44, 58, 59 loss of regulation by RPII due to 13q14 deletions should alter the expression of miR-15a/miR-16-1 and, consequently, cell cycle and apoptosis. Evidences shows that co-expression of DLEU2 and of miR-15a/16-1 by RPII is repressed by CMYC and activated by E2F1 and P53.15, 44, 58, 59 In our model, this applies only to the RPII-dependent allele, while the transcription of the primiR-15a/16-1 from the RPIII-dependent allele could be activated by CMYC and repressed by P53.49 Exclusive RPIII regulation of miR-15a/16-1 could affect P53/miR-34/ZAP70 signaling,15 resulting, over time, in ZAP70-positive B-CLL cells. Two genetic characteristics associate with high ZAP70: (i), primiR expression driven by RPIII; (ii), type of 13q deletion that retains both alleles of the miR-15a/16-1 locus.
The double allele-specific regulation of a miR-15a/16-1 driven by either RPII or RPIII may provide the basis for understanding the clinical heterogeneity of the CLL patients carrying 13q14 deletions, and exclusive RPIII-driven transcription of this miR cluster may differentiate at the onset aggressive from indolent forms of CLL.
References
Chiorazzi N, Rai KR, Ferrarini M . Chronic lymphocytic leukemia. N Engl J Med 2005; 352: 804–815.
Bosch F, Montserrat E . Refining prognostic factors in chronic lymphocytic leukemia. Rev Clin Exp Hematol 2002; 6: 335–349, discussion 449-350.
Edelmann J, Holzmann K, Miller F, Winkler D, Buhler A, Zenz T et al. High-resolution genomic profiling of chronic lymphocytic leukemia reveals new recurrent genomic alterations. Blood 2012; 120: 4783–4794.
Calin GA, Dumitru CD, Shimizu M, Bichi R, Zupo S, Noch E et al. Frequent deletions and down-regulation of micro- RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc Natl Acad Sci USA 2002; 99: 15524–15529.
Calin GA, Ferracin M, Cimmino A, Di Leva G, Shimizu M, Wojcik SE et al. A microRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N Engl J Med 2005; 353: 1793–1801.
Landau DA, Carter SL, Stojanov P, McKenna A, Stevenson K, Lawrence MS et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 2013; 152: 714–726.
Rawstron AC . Monoclonal B-cell lymphocytosis. Hematology Am Soc Hematol Educ Program 2009, 430–439.
Rossi D, Sozzi E, Puma A, De Paoli L, Rasi S, Spina V et al. The prognosis of clinical monoclonal B cell lymphocytosis differs from prognosis of Rai 0 chronic lymphocytic leukaemia and is recapitulated by biological risk factors. Br J Haematol 2009; 146: 64–75.
Rawstron AC, Bennett FL, O'Connor SJ, Kwok M, Fenton JA, Plummer M et al. Monoclonal B-cell lymphocytosis and chronic lymphocytic leukemia. N Engl J Med 2008; 359: 575–583.
Krober A, Seiler T, Benner A, Bullinger L, Bruckle E, Lichter P et al. V(H) mutation status, CD38 expression level, genomic aberrations, and survival in chronic lymphocytic leukemia. Blood 2002; 100: 1410–1416.
Dohner H, Stilgenbauer S, Benner A, Leupolt E, Krober A, Bullinger L et al. Genomic aberrations and survival in chronic lymphocytic leukemia. N Engl J Med 2000; 343: 1910–1916.
Ouillette P, Collins R, Shakhan S, Li J, Li C, Shedden K et al. The prognostic significance of various 13q14 deletions in chronic lymphocytic leukemia. Clin Cancer Res 2011; 17: 6778–6790.
Ouillette P, Erba H, Kujawski L, Kaminski M, Shedden K, Malek SN . Integrated genomic profiling of chronic lymphocytic leukemia identifies subtypes of deletion 13q14. Cancer Res 2008; 68: 1012–1021.
Klein U, Lia M, Crespo M, Siegel R, Shen Q, Mo T et al. The DLEU2/miR-15a/16-1 cluster controls B cell proliferation and its deletion leads to chronic lymphocytic leukemia. Cancer Cell 2010; 17: 28–40.
Fabbri M, Bottoni A, Shimizu M, Spizzo R, Nicoloso MS, Rossi S et al. Association of a microRNA/TP53 feedback circuitry with pathogenesis and outcome of B-cell chronic lymphocytic leukemia. JAMA 2011; 305: 59–67.
Aqeilan RI, Calin GA, Croce CM . miR-15a and miR-16-1 in cancer: discovery, function and future perspectives. Cell Death Differ 2010; 17: 215–220.
Calin GA, Cimmino A, Fabbri M, Ferracin M, Wojcik SE, Shimizu M et al. MiR-15a and miR-16-1 cluster functions in human leukemia. Proc Natl Acad Sci USA 2008; 105: 5166–5171.
Mertens D, Philippen A, Ruppel M, Allegra D, Bhattacharya N, Tschuch C et al. Chronic lymphocytic leukemia and 13q14: miRs and more. Leuk Lymphoma 2009; 50: 502–505.
Mertens D, Wolf S, Tschuch C, Mund C, Kienle D, Ohl S et al. Allelic silencing at the tumor-suppressor locus 13q14.3 suggests an epigenetic tumor-suppressor mechanism. Proc Natl Acad Sci USA 2006; 103: 7741–7746.
Suzuki HI, Yamagata K, Sugimoto K, Iwamoto T, Kato S, Miyazono K . Modulation of microRNA processing by p53. Nature 2009; 460: 529–533.
Calin GA, Trapasso F, Shimizu M, Dumitru CD, Yendamuri S, Godwin AK et al. Familial cancer associated with a polymorphism in ARLTS1. N Engl J Med 2005; 352: 1667–1676.
Rassenti LZ, Huynh L, Toy TL, Chen L, Keating MJ, Gribben JG et al. ZAP-70 compared with immunoglobulin heavy-chain gene mutation status as a predictor of disease progression in chronic lymphocytic leukemia. N Engl J Med 2004; 351: 893–901.
Wendel-Hansen V, Sallstrom J, De Campos-Lima PO, Kjellstrom G, Sandlund A, Siegbahn A et al. Epstein-Barr virus (EBV) can immortalize B-cll cells activated by cytokines. Leukemia 1994; 8: 476–484.
Stacchini A, Aragno M, Vallario A, Alfarano A, Circosta P, Gottardi D et al. MEC1 and MEC2: two new cell lines derived from B-chronic lymphocytic leukaemia in prolymphocytoid transformation. Leuk Res 1999; 23: 127–136.
Doyle MG, Catovsky D, Crawford DH . Infection of leukaemic B lymphocytes by Epstein Barr virus. Leukemia 1993; 7: 1858–1864.
Lachmann R, Lanuti P, Miscia S . OMIP-011: characterization of circulating endothelial cells (CECs) in peripheral blood. Cytometry A 2012; 81: 549–551.
Lanuti P, Ciccocioppo F, Bonanni L, Marchisio M, Lachmann R, Tabet N et al. Amyloid-specific T-cells differentiate Alzheimer's disease from Lewy body dementia. Neurobiol Aging 2012; 33: 2599–2611.
Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT et al. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res 2005; 33: e179.
Varkonyi-Gasic E, Wu R, Wood M, Walton EF, Hellens RP . Protocol: a highly sensitive RT-PCR method for detection and quantification of microRNAs. Plant Methods 2007; 3: 12.
Tang F, Hajkova P, Barton SC, Lao K, Surani MA . MicroRNA expression profiling of single whole embryonic stem cells. Nucleic Acids Res 2006; 34: e9.
Visone R, Rassenti LZ, Veronese A, Taccioli C, Costinean S, Aguda BD et al. Karyotype-specific microRNA signature in chronic lymphocytic leukemia. Blood 2009; 114: 3872–3879.
Bichi R, Shinton SA, Martin ES, Koval A, Calin GA, Cesari R et al. Human chronic lymphocytic leukemia modeled in mouse by targeted TCL1 expression. Proc Natl Acad Sci USA 2002; 99: 6955–6960.
Zeng Y, Cullen BR . Efficient processing of primary microRNA hairpins by Drosha requires flanking nonstructured RNA sequences. J Biol Chem 2005; 280: 27595–27603.
Wu M, Jolicoeur N, Li Z, Zhang L, Fortin Y, L'Abbe D et al. Genetic variations of microRNAs in human cancer and their effects on the expression of miRNAs. Carcinogenesis 2008; 29: 1710–1716.
Iwai N, Naraba H . Polymorphisms in human pre-miRNAs. Biochem Biophys Res Commun 2005; 331: 1439–1444.
Han SJ, Marshall V, Barsov E, Quinones O, Ray A, Labo N et al. Kaposi's sarcoma-associated herpesvirus microRNA single-nucleotide polymorphisms identified in clinical samples can affect microRNA processing, level of expression, and silencing activity. J Virol 2013; 87: 12237–12248.
Ryan BM, Robles AI, Harris CC . Genetic variation in microRNA networks: the implications for cancer research. Nat Rev Cancer 2010; 10: 389–402.
Fukuda T, Yamagata K, Fujiyama S, Matsumoto T, Koshida I, Yoshimura K et al. DEAD-box RNA helicase subunits of the Drosha complex are required for processing of rRNA and a subset of microRNAs. Nat Cell Biol 2007; 9: 604–611.
Schramm L, Hernandez N . Recruitment of RNA polymerase III to its target promoters. Genes Dev 2002; 16: 2593–2620.
Raha D, Wang Z, Moqtaderi Z, Wu L, Zhong G, Gerstein M et al. Close association of RNA polymerase II and many transcription factors with Pol III genes. Proc Natl Acad Sci USA 2010; 107: 3639–3644.
Monteys AM, Spengler RM, Wan J, Tecedor L, Lennox KA, Xing Y et al. Structure and activity of putative intronic miRNA promoters. RNA 2010; 16: 495–505.
Wang Z, Roeder RG . Three human RNA polymerase III-specific subunits form a subcomplex with a selective function in specific transcription initiation. Genes Dev 1997; 11: 1315–1326.
Sepehri S, Hernandez N . The largest subunit of human RNA polymerase III is closely related to the largest subunit of yeast and trypanosome RNA polymerase III. Genome Res 1997; 7: 1006–1019.
Lerner M, Harada M, Loven J, Castro J, Davis Z, Oscier D et al. DLEU2, frequently deleted in malignancy, functions as a critical host gene of the cell cycle inhibitory microRNAs miR-15a and miR-16-1. Exp Cell Res 2009; 315: 2941–2952.
Lee Y, Kim M, Han J, Yeom KH, Lee S, Baek SH et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J 2004; 23: 4051–4060.
James Faresse N, Canella D, Praz V, Michaud J, Romascano D, Hernandez N . Genomic study of RNA polymerase II and III SNAPc-bound promoters reveals a gene transcribed by both enzymes and a broad use of common activators. PLoS Genet 2012; 8: e1003028.
Listerman I, Bledau AS, Grishina I, Neugebauer KM . Extragenic accumulation of RNA polymerase II enhances transcription by RNA polymerase III. PLoS Genet 2007; 3: e212.
Cramer P, Armache KJ, Baumli S, Benkert S, Brueckner F, Buchen C et al. Structure of eukaryotic RNA polymerases. Annu Rev Biophys 2008; 37: 337–352.
Marshall L, White RJ . Non-coding RNA production by RNA polymerase III is implicated in cancer. Nat Rev Cancer 2008; 8: 911–914.
Felton-Edkins ZA, Kenneth NS, Brown TR, Daly NL, Gomez-Roman N, Grandori C et al. Direct regulation of RNA polymerase III transcription by RB, p53 and c-Myc. Cell Cycle 2003; 2: 181–184.
Gomez-Roman N, Grandori C, Eisenman RN, White RJ . Direct activation of RNA polymerase III transcription by c-Myc. Nature 2003; 421: 290–294.
Crighton D, Woiwode A, Zhang C, Mandavia N, Morton JP, Warnock LJ et al. p53 represses RNA polymerase III transcription by targeting TBP and inhibiting promoter occupancy by TFIIIB. EMBO J 2003; 22: 2810–2820.
Pospisilova S, Gonzalez D, Malcikova J, Trbusek M, Rossi D, Kater AP et al. ERIC recommendations on TP53 mutation analysis in chronic lymphocytic leukemia. Leukemia 2012; 26: 1458–1461.
Rinaldi A, Mian M, Kwee I, Rossi D, Deambrogi C, Mensah AA et al. Genome-wide DNA profiling better defines the prognosis of chronic lymphocytic leukaemia. Br J Haematol 2011; 154: 590–599.
Put N, Van Roosbroeck K, Konings P, Meeus P, Brusselmans C, Rack K et al. Chronic lymphocytic leukemia and prolymphocytic leukemia with MYC translocations: a subgroup with an aggressive disease course. Ann Hematol 2012; 91: 863–873.
Cairns CA, White RJ . p53 is a general repressor of RNA polymerase III transcription. EMBO J 1998; 17: 3112–3123.
Stein T, Crighton D, Warnock LJ, Milner J, White RJ . Several regions of p53 are involved in repression of RNA polymerase III transcription. Oncogene 2002; 21: 5540–5547.
Chang TC, Yu D, Lee YS, Wentzel EA, Arking DE, West KM et al. Widespread microRNA repression by Myc contributes to tumorigenesis. Nat Genet 2008; 40: 43–50.
Ofir M, Hacohen D, Ginsberg D . MiR-15 and miR-16 are direct transcriptional targets of E2F1 that limit E2F-induced proliferation by targeting cyclin E. Mol Cancer Res 2011; 9: 440–447.
Kaplan D, Smith D, Meyerson H, Pecora N, Lewandowska K . CD5 expression by B lymphocytes and its regulation upon Epstein-Barr virus transformation. Proc Natl Acad Sci USA 2001; 98: 13850–13853.
Acknowledgements
We thank Dante Tatone and Lia De Amicis for excellent technical and management assistance. Thanks to the genetic laboratories of Professor L Stuppia and Professor G. Calabrese (University of Chieti) that performed the cell lines authentication by STR and karyotype analysis. Thanks to Reza Ghasemi for help with the writing of the manuscript. AV is supported by a Marie Curie Career Integration Grants. This study was supported by Associazione Italiana Ricerca sul Cancro, AIRC Start Up grant 2010 (10054) and Marie Curie Career Integration Grant (GA-2011-303735) to RV.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on the Leukemia website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/
About this article
Cite this article
Veronese, A., Pepe, F., Chiacchia, J. et al. Allele-specific loss and transcription of the miR-15a/16-1 cluster in chronic lymphocytic leukemia. Leukemia 29, 86–95 (2015). https://doi.org/10.1038/leu.2014.139
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2014.139
This article is cited by
-
MicroRNA-15a-5p acts as a tumor suppressor in histiocytosis by mediating CXCL10-ERK-LIN28a-let-7 axis
Leukemia (2022)
-
Chronic lymphocytic leukaemia
Nature Reviews Disease Primers (2017)
-
Effects of miRNA-15 and miRNA-16 expression replacement in chronic lymphocytic leukemia: implication for therapy
Leukemia (2017)
-
Evolution of multiple cell clones over a 29-year period of a CLL patient
Nature Communications (2016)
-
Role of microRNA in chronic lymphocytic leukemia onset and progression
Journal of Hematology & Oncology (2015)