Abstract
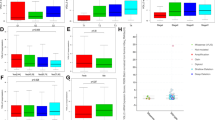
Mucosa-associated lymphoid tissue (MALT) lymphoma is characterized by t(11;18)(q21;q21)/API2-MALT1, t(1;14)(p22;q32)/BCL10-IGH and t(14;18)(q32;q21)/IGH-MALT1, which commonly activate the nuclear factor (NF)-κB pathway. Gastric MALT lymphomas harboring such translocations usually do not respond to Helicobacter pylori eradication, while most of those without translocation can be cured by antibiotics. To understand the molecular mechanism of these different MALT lymphoma subgroups, we performed gene expression profiling analysis of 21 MALT lymphomas (13 translocation-positive, 8 translocation-negative). Gene set enrichment analysis (GSEA) of the NF-κB target genes and 4394 additional gene sets covering various cellular pathways, biological processes and molecular functions have shown that translocation-positive MALT lymphomas are characterized by an enhanced expression of NF-κB target genes, particularly toll like receptor (TLR)6, chemokine, CC motif, receptor (CCR)2, cluster of differentiation (CD)69 and B-cell CLL/lymphoma (BCL)2, while translocation-negative cases were featured by active inflammatory and immune responses, such as interleukin-8, CD86, CD28 and inducible T-cell costimulator (ICOS). Separate analyses of the genes differentially expressed between translocation-positive and -negative cases and measurement of gene ontology term in these differentially expressed genes by hypergeometric test reinforced the above findings by GSEA. Finally, expression of TLR6, in the presence of TLR2, enhanced both API2-MALT1 and BCL10-mediated NF-κB activation in vitro. Our findings provide novel insights into the molecular mechanism of MALT lymphomas with and without translocation, potentially explaining their different clinical behaviors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Isaacson PG, Du MQ . MALT lymphoma: from morphology to molecules. Nat Rev Cancer 2004; 4: 644–653.
Ye H, Liu H, Attygalle A, Wotherspoon AC, Nicholson AG, Charlotte F et al. Variable frequencies of t(11;18)(q21;q21) in MALT lymphomas of different sites: significant association with CagA strains of H pylori in gastric MALT lymphoma. Blood 2003; 102: 1012–1018.
Ye H, Gong L, Liu H, Hamoudi RA, Shirali S, Ho L et al. MALT lymphoma with t(14;18)(q32;q21)/IGH-MALT1 is characterized by strong cytoplasmic MALT1 and BCL10 expression. J Pathol 2005; 205: 293–301.
Streubel B, Simonitsch-Klupp I, Mullauer L, Lamprecht A, Huber D, Siebert R et al. Variable frequencies of MALT lymphoma-associated genetic aberrations in MALT lymphomas of different sites. Leukemia 2004; 18: 1722–1726.
Goatly A, Bacon CM, Nakamura S, Ye H, Kim I, Brown PJ et al. FOXP1 abnormalities in lymphoma: translocation breakpoint mapping reveals insights into deregulated transcriptional control. Mod Pathol 2008; 21: 902–911.
Lucas PC, Yonezumi M, Inohara N, McAllister-Lucas LM, Abazeed ME, Chen FF et al. Bcl10 and MALT1, independent targets of chromosomal translocation in MALT lymphoma, cooperate in a novel NF-kappa B signaling pathway. J Biol Chem 2001; 276: 19012–19019.
Zhou H, Wertz I, O’Rourke K, Ultsch M, Seshagiri S, Eby M et al. Bcl10 activates the NF-kappaB pathway through ubiquitination of NEMO. Nature 2004; 427: 167–171.
Lucas PC, Kuffa P, Gu S, Kohrt D, Kim DS, Siu K et al. A dual role for the API2 moiety in API2-MALT1-dependent NF-kappaB activation: heterotypic oligomerization and TRAF2 recruitment. Oncogene 2007; 26: 5643–5654.
Baens M, Fevery S, Sagaert X, Noels H, Hagens S, Broeckx V et al. Selective expansion of marginal zone B cells in Emicro-API2-MALT1 mice is linked to enhanced IkappaB kinase gamma polyubiquitination. Cancer Res 2006; 66: 5270–5277.
Li Z, Wang H, Xue L, Shin DM, Roopenian D, Xu W et al. Emu-BCL10 mice exhibit constitutive activation of both canonical and noncanonical NF-kappaB pathways generating marginal zone (MZ) B-cell expansion as a precursor to splenic MZ lymphoma. Blood 2009; 114: 4158–4168.
Zhou Y, Ye H, Martin-Subero JI, Hamoudi R, Lu YJ, Wang R et al. Distinct comparative genomic hybridisation profiles in gastric mucosa-associated lymphoid tissue lymphomas with and without t(11;18)(q21;q21). Br J Haematol 2006; 133: 35–42.
Sagaert X, Theys T, Wolf-Peeters C, Marynen P, Baens M . Splenic marginal zone lymphoma-like features in API2-MALT1 transgenic mice that are exposed to antigenic stimulation. Haematologica 2006; 91: 1693–1696.
Ho L, Davis RE, Conne B, Chappuis R, Berczy M, Mhawech P et al. MALT1 and the API2-MALT1 fusion act between CD40 and IKK and confer NF-kappa B-dependent proliferative advantage and resistance against FAS-induced cell death in B cells. Blood 2005; 105: 2891–2899.
Liu H, Ye H, Ruskone-Fourmestraux A, de Jong D, Pileri S, Thiede C et al. T(11;18) is a marker for all stage gastric MALT lymphomas that will not respond to H pylori eradication. Gastroenterology 2002; 122: 1286–1294.
Ye H, Gong L, Liu H, Ruskone-Fourmestraux A, de Jong D, Pileri S et al. Strong BCL10 nuclear expression identifies gastric MALT lymphomas that do not respond to H pylori eradication. Gut 2006; 55: 137–138.
Okabe M, Inagaki H, Ohshima K, Yoshino T, Li C, Eimoto T et al. API2-MALT1 fusion defines a distinctive clinicopathologic subtype in pulmonary extranodal marginal zone B-cell lymphoma of mucosa-associated lymphoid tissue. Am J Pathol 2003; 162: 1113–1122.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA 2005; 102: 15545–15550.
Saxena V, Orgill D, Kohane I . Absolute enrichment: gene set enrichment analysis for homeostatic systems. Nucleic Acids Res 2006; 34: e151.
Ortutay C, Vihinen M . Immunome: a reference set of genes and proteins for systems biology of the human immune system. Cell Immunol 2006; 244: 87–89.
Pepper SD, Saunders EK, Edwards LE, Wilson CL, Miller CJ . The utility of MAS5 expression summary and detection call algorithms. BMC Bioinformatics 2007; 8: 273.
Gomariz RP, Arranz A, Juarranz Y, Gutierrez-Canas I, Garcia-Gomez M, Leceta J et al. Regulation of TLR expression, a new perspective for the role of VIP in immunity. Peptides 2007; 28: 1825–1832.
Chng WJ, Remstein ED, Fonseca R, Bergsagel PL, Vrana JA, Kurtin PJ et al. Gene expression profiling of pulmonary mucosa-associated lymphoid tissue lymphoma identifies new biologic insights with potential diagnostic and therapeutic applications. Blood 2009; 113: 635–645.
Rubtsov AV, Swanson CL, Troy S, Strauch P, Pelanda R, Torres RM . TLR agonists promote marginal zone B cell activation and facilitate T-dependent IgM responses. J Immunol 2008; 180: 3882–3888.
Akira S, Takeda K . Toll-like receptor signalling. Nat Rev Immunol 2004; 4: 499–511.
Ohmae T, Hirata Y, Maeda S, Shibata W, Yanai A, Ogura K et al. Helicobacter pylori activates NF-kappaB via the alternative pathway in B lymphocytes. J Immunol 2005; 175: 7162–7169.
Yokota S, Ohnishi T, Muroi M, Tanamoto K, Fujii N, Amano K . Highly-purified Helicobacter pylori LPS preparations induce weak inflammatory reactions and utilize Toll-like receptor 2 complex but not Toll-like receptor 4 complex. FEMS Immunol Med Microbiol 2007; 51: 140–148.
Sancho D, Gomez M, Sanchez-Madrid F . CD69 is an immunoregulatory molecule induced following activation. Trends Immunol 2005; 26: 136–140.
Erlanson M, Gronlund E, Lofvenberg E, Roos G, Lindh J . Expression of activation markers CD23 and CD69 in B-cell non-Hodgkin's lymphoma. Eur J Haematol 1998; 60: 125–132.
de Jong D, Koster A, Hagenbeek A, Raemaekers J, Veldhuizen D, Heisterkamp S et al. Impact of the tumor microenvironment on prognosis in follicular lymphoma is dependent on specific treatment protocols. Haematologica 2009; 94: 70–77.
Hopken UE, Wengner AM, Loddenkemper C, Stein H, Heimesaat MM, Rehm A et al. CCR7 deficiency causes ectopic lymphoid neogenesis and disturbed mucosal tissue integrity. Blood 2007; 109: 886–895.
Nie Y, Waite J, Brewer F, Sunshine MJ, Littman DR, Zou YR . The role of CXCR4 in maintaining peripheral B cell compartments and humoral immunity. J Exp Med 2004; 200: 1145–1156.
Springael JY, Urizar E, Parmentier M . Dimerization of chemokine receptors and its functional consequences. Cytokine Growth Factor Rev 2005; 16: 611–623.
Lopez-Giral S, Quintana NE, Cabrerizo M, Alfonso-Perez M, Sala-Valdes M, De Soria VG et al. Chemokine receptors that mediate B cell homing to secondary lymphoid tissues are highly expressed in B cell chronic lymphocytic leukemia and non-Hodgkin lymphomas with widespread nodular dissemination. J Leukoc Biol 2004; 76: 462–471.
Hopken UE, Foss HD, Meyer D, Hinz M, Leder K, Stein H et al. Up-regulation of the chemokine receptor CCR7 in classical but not in lymphocyte-predominant Hodgkin disease correlates with distinct dissemination of neoplastic cells in lymphoid organs. Blood 2002; 99: 1109–1116.
Trentin L, Cabrelle A, Facco M, Carollo D, Miorin M, Tosoni A et al. Homeostatic chemokines drive migration of malignant B cells in patients with non-Hodgkin lymphomas. Blood 2004; 104: 502–508.
Novak A, Akasaka T, Manske M, Gupta M, Witzig T, Dyer MJS et al. Elevated expression of GPR34 and its association with a novel translocation t(X;14)(p11;q32) involving IGHS and GPR34 in MALT lymphoma. Blood 2008; 112: 785, Abstract 2251.
Du MQ, Atherton JC . Molecular subtyping of gastric MALT lymphomas: implications for prognosis and management. Gut 2006; 55: 886–893.
McNamara D, El Omar E . Helicobacter pylori infection and the pathogenesis of gastric cancer: a paradigm for host-bacterial interactions. Dig Liver Dis 2008; 40: 504–509.
de Jong D, Vyth-Dreese F, Dellemijn T, Verra N, Ruskone-Fourmestraux A, Lavergne-Slove A et al. Histological and immunological parameters to predict treatment outcome of Helicobacter pylori eradication in low-grade gastric MALT lymphoma. J Pathol 2001; 193: 318–324.
Acknowledgements
We would like to thank Professor Ahmet Dogan and Dr Ellen Remstein, Department of Pathology, Mayo Clinic for sharing their pulmonary MALT lymphoma gene expression microarray data; Dr Ian McFarlane, the Microarray CoreLab, National Institute of Health Research, Cambridge Comprehensive Biomedical Research Centre for his technical assistance, Dr Koichi Kuwano, Department of Infectious Medicine, Kurume University School of Medicine, Japan for providing TLR1, TLR2 and TLR6 expression constructs and both previous and present members of Du lab for helpful discussion and assistance. The research in Du lab was supported by research grants from Leukemia Research, UK and the National Institute for Health Research Cambridge Biomedical Research Center. BS was supported by Grant FWF (P19346-B12). LdL is a senior research associate of the FRS-FNRS.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Authors contribution
RAH designed the experiment, collected and analyzed the data. AA, HY and LG contributed to the design and experimental data collection and analysis; ARF, BS, AC, MR, IW, CDWP, KAM, LdL and PGI provided lymphoma cases; MQD designed, analyzed the data and wrote the paper.
Supplementary Information accompanies the paper on the Leukemia website
Supplementary information
Rights and permissions
About this article
Cite this article
Hamoudi, R., Appert, A., Ye, H. et al. Differential expression of NF-κB target genes in MALT lymphoma with and without chromosome translocation: insights into molecular mechanism. Leukemia 24, 1487–1497 (2010). https://doi.org/10.1038/leu.2010.118
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/leu.2010.118
Keywords
This article is cited by
-
Molecular pathogenicity of 1-nonadecene and l-lactic acid, unique metabolites in radicular cysts and periapical granulomas
Scientific Reports (2023)
-
Constitutive expression of c-REL in uveal melanoma patients: correlation with clinicopathological parameters and patient outcome
Clinical and Translational Oncology (2020)
-
CARD–BCL-10–MALT1 signalling in protective and pathological immunity
Nature Reviews Immunology (2019)
-
Using discretization for extending the set of predictive features
EURASIP Journal on Advances in Signal Processing (2018)
-
Comparative gene-expression profiling of the large cell variant of gastrointestinal marginal-zone B-cell lymphoma
Scientific Reports (2017)