Abstract
Fabry disease (FD) is a rare X-linked recessive glycosphingolipid-storage disorder caused by deficient activity of the lysosomal enzyme alpha-galactosidase A. Intravenous enzyme replacement therapy (ERT) has been used to supplement deficient enzyme activity in patients with FD. Despite its clinical effect and manifestations, clear criteria for the clinical effectiveness and cost-effectiveness of ERT have not been well established. In this study, we investigated the pharmacodynamic actions and short-term effects of ERT in patients with FD through direct molecular profiling from blood samples of patients before and after ERT. Based on this comparison, we observed that immune/inflammation-related pathways and growth factor-related pathways such as innate/adaptive immune pathway, lymphocyte proliferation and leukocyte proliferation were actively regulated under ERT. We also found that TINAGL1, DAAM2, CDK5R1 and MYO5B known to be related with clinical symptoms of FD showed increased levels after ERT, leading to the amelioration of clinical manifestations. Especially the catabolic process-related genes, including USP15 and ERUN1, showed direct increasing after ERT in vivo in male patients. These results suggest that male patients with FD respond more actively to ERT than do female patients with FD. Pathway analysis revealed that oxidative phosphorylation pathway-related genes are downregulated under ERT. ERT has a role to protect the proteins from oxidative damage and such deactivation of oxidative phosphorylation is one of direct pharmacodynamic actions of ERT. These results extended our understanding of the pathophysiology of ERT. To our knowledge, this is the first study to observe the molecular basis for the mechanism of ERT in vivo through the comprehensive comparison of transcriptome study with next-generation sequencing data.
Similar content being viewed by others
Introduction
Fabry disease (FD) is an X-linked recessive lysosomal storage disorder caused by deficient activity of alpha-galactosidase A (GAL-A, E.C. 3.2.1.22), which results in the intralysosomal accumulation of glycosphingolipids in the body.1 The initial symptoms, including acroparesthesia and pain with fever in childhood, make the timely diagnosis of FD difficult. Intravenous enzyme replacement therapy (ERT) has been used in patients with FD to supplement the deficient enzyme, but clear criteria for the clinical effectiveness and cost-effectiveness of ERT in symptomatic patients have not yet been established, despite the high annual costs associated with therapy for lysosomal diseases.2, 3
Although there have been numerous studies to investigate the efficacy of ERT on multiple organ systems,4, 5, 6, 7, 8, 9, 10, 11, 12, 13 most of the studies were focused on the long-term clinical effects of ERT based on quantifiable clinical measurements such as measurements of pain, renal, cardiac and cerebral outcomes and health-related quality of life.14, 15, 16, 17, 18 Although ERT ameliorates many of the manifestations of FD and leads to dramatically improved quality of life for many patients, little is known about the pharmacodynamics of action or the pathway-level mechanisms during the course of ERT in patients. Recently, some studies reported the presence of antibodies against ERT in 70–100% of patients,19, 20, 21, 22 but there are few studies regarding the effectiveness or clinical impact of antibodies for the therapy.19, 20, 21, 23
Until now, no direct comparison of in vivo molecular profiles before and after ERT has been performed, and the in vivo properties of ERT remain unclear. Therefore, we need to understand the molecular basis for the mechanism of action of ERT. Here we first investigated the short-term effects of ERT through a direct comparison of molecular perturbation patterns among Korean patients with FD. To our knowledge, this is the first study reporting the molecular basis for the mechanism of ERT at the molecular pathway level using a transcriptome study. The advent of rapid sequencing technologies has allowed for detailed profiling of gene-expression levels with reduced costs, and sequence-based expression analysis is now increasingly being adopted for clinical use.24, 25 We applied RNA-seq to identify the differentially expressed genes (DEGs) and the molecular pathways directly responding to ERT. We found that immune/inflammation-related genes are significantly upregulated after ERT, while the genes associated with oxidative phosphorylation are downregulated under ERT. The consideration of gender revealed that male patients with FD were more actively responsive than female patients to ERT.
Materials and methods
Ethics approval
The study was approved by the institutional review board of the Severance Hospital (No. 4-2014-0787) of Yonsei University. Written informed consent was obtained from the patients (>20 years) or the parents of child patients before the study began.
Human patient samples
Prospective data were collected from six patients with FD (two male patients with classical phenotypes and four symptomatic female carriers) who had been receiving ERT. The patients were aged between 17 and 35 years. The patients were receiving ERT with Agalsidase beta (Fabrazyme, Genzyme Corporation, Cambridge, MA, USA), which was intravenously infused at a dose of 1 mg kg−1 per 2 weeks. Blood samples (3 ml) were obtained immediately before and after ERT for the transcriptome study to observe the molecular-level changes. RNAs were extracted from the blood samples. For a fair comparison, the patients were guided not to drink or eat any kinds of food or water after ERT to prevent the perturbation of molecular signals owing to the ingestion process.
Illumina strand-specific RNA sequencing
After extracting the total RNA from the samples, mRNA was isolated using oligo-dT magnetic beads from 1 μg total RNA. Isolated RNA was fragmented into pieces about 200 nucleotides long and purified for cDNA synthesis. Then sequencing adaptors were ligated to the fragments, and size selection of the adaptor-ligated cDNA was performed using Agencourt AMPure XP (Beckman Coulter, Brea, CA, USA) Beads. These fragments were purified and PCR-amplified to construct the sequencing library. The primary sequencing reads were subjected to quality control. Sequencing was performed on an Illumina MiSeq instrument (Illumina, Inc., San Diego, CA, USA).
Data analysis and statistical modeling
All primary analysis of sequence data was performed with Bowtie226 and TopHat27 packages. Bowtie2 is an ultrafast and memory-efficient tool for aligning sequencing reads to long reference sequences. TopHat is a specially designed tool for fast splice-junction mapping of RNA-seq reads. It also analyzes the mapping results to identify splice junctions between exons. In our study, raw FASTQ files were generated using a MiSeq machine from Illumina, and sequence reads were mapped to the UCSC hg19 reference genome using Botwie2 and TopHat. Once the sequence reads were successfully mapped to the reference genome, the htSeq program, which contains powerful options for analyzing high-throughput sequencing data, was used to count the reads mapped to each gene. It counts reads that overlap with any exon for the given gene, counting the untranslated region as part of the first exon. Once the count of the reads for each gene was finished, the tags with very low counts were filtered out in order to prevent the intervention of weakly expressed tags. We kept the tags expressed in at least two of the samples. Then a normalization process was applied to account for the differences in the number of reads produced in different sequencing runs as well as technical biases caused by the library preparation protocols, sequencing platforms and nucleotide compositions.28 The edgeR29 package was used for differential gene-expression analysis. To address the overdispersion problem, which is caused by the assumption of a Poisson distribution where the mean is equal to the variance, edgeR assumes a negative binomial distribution, where the relation between the variance υ and mean μ is defined as υs=μ+αμ2. Using the negative bionomical assumption, we estimated the coefficient of variance for each gene. Then a generalized linear model was applied for the statistical comparison of gene expression before and after ERT. First, we tested the ERT effect in a paired design under the additive model by estimating the baseline differences among patients, which would be subtracted out. DEGs under ERT were identified by comparing the expression profiles before and after ERT. Then gender was considered in order to identify DEGs under ERT, which would be a critical factor owing to the characteristics of FD. Because FD is X-linked, males and females can both have symptoms, but males are usually more severely affected than females. Hence, we expect that there might be differences in the responsiveness and effectiveness of ERT depending on the degree of clinical manifestations. Indeed, we observed distinct differences in gene-expression patterns under ERT between female and male patients.
Results
Analysis of DEGs associated with ERT
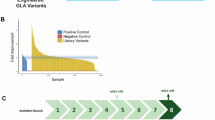
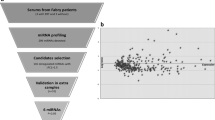
First, we identified DEGs responding to ERT. There were about 30 DEGs, with significantly (P-value<0.05) perturbed expression under ERT. Because ERT for patients with FD only involves the replacement of the deficient enzyme, we would not expect dramatic changes, including serious side effects in the body owing to the intravenous infusion of recombinant enzyme. However, there would be clear molecular responsiveness associated with ERT. Indeed, the DEGs responding to ERT were enriched for immune-related pathways such as lymphocyte proliferation and leukocyte proliferation. The infusion of recombinant enzyme would trigger immune reactions, leading to subtle molecular-level changes in the body. However, there is no confirmed association between the immune responses associated with antibodies and the clinical effectiveness. Such initial immune response is natural and causes only minor effects; there is a small fraction of genes responding to ERT. To further elucidate the biological differences between normal physiological and pharmacological conditions, the profiles of pre-ERT and post-ERT conditions were compared (Figure 1a). The RNA-seq data demonstrated minor differences after ERT, which is also consistent with the results in Figure 1d. Most of the female patients revealed very minor gene-expression differences before and after ERT, while most of the gene-expression levels stayed the same (Figure 1a). However, changes of gene expression in the male patients were much clearer; the regression line for the expression levels before and after ERT shows that many of the genes were downregulated right after ERT (for example, GDY and SHM). Figure 1b represents the scatter plot of gene-expression levels showing significant differences before and after ERT in the male patients. Interestingly, although the gene-expression levels generally tended to decrease slightly (Figure 1a, bottom right), highly ranked, significantly changed genes (2-fold (in red) and <0.5-fold (in blue)) showed upregulated patterns after ERT (Figure 1b). Comparison of the gene-expression levels with hierarchical clustering showed that male patients were clearly distinguished from female patients based on the fold changes in gene expression (Figure 1c), which is consistent with Figure 1e.
Comparison of biological differences before and after ERT (two male patients and four female patients, two samples (precondition/postcondition) per patient). Each sample was obtained before and after ERT from each patient. (a) Scatter plots between normal physiological (pre) conditions and pharmacological (post) conditions. HKS, KYM, KWK and PYM represent female patients, and GDY and SHM represent male patients. (b) Scatter plots of gene-expression levels (FPKM) showing significant differences between precondition and postcondition in a male patient. The genes having >2-fold (in red) and <0.5-fold (in blue) differences are plotted. (c) Hierarchical clustering of six Fabry patients based on log fold change ratio. Male patients were clearly distinguished from female patients based on their expression perturbation patterns. (d) Twelve samples including pre-ERT and post-ERT conditions. (e) The same samples annotated only by gender. The samples from male patients reveal more clear differences between pre-ERT and post-ERT conditions (light green vs deep green). A full color version of this figure is available at the Journal of Human Genetics journal online.
Gender-specific responsiveness to ERT
We performed a principal component analysis of all individual samples to see the overall patterns of responding genes based on the RNA-seq expression profiles (Figures 1d and e). For fair comparison, we excluded the genes in sex chromosome. Figure 1d shows no clear distinction between pre-ERT and post-ERT conditions at the molecular level. However, it is interesting to note that male patients revealed much more active gene-expression changes associated with ERT than did female patients (Figure 1e). These data suggest that gender would be a critical factor to identify the molecular signals associated with ERT. Thus we identified DEGs responding to ERT under the consideration of gender. As we see in Figure 1, there were more DEGs in the male patients than in the female patients (167 DEGs in male patients, 37 DEGs in female patients; P-value<0.05), which indicates the occurrence of an active response to ERT in the male patients at the molecular level. Because Fabry disorder is X-linked, male patients usually experience much more progressive and severer multi-organ damages, while there are considerable variations in the clinical manifestations in female carriers. Therefore, the active responses in the male patients might be related to severe clinical manifestations or damages. Pathway-enrichment analysis was performed to identify overrepresented or under-represented gene ontology terms in the course of ERT using the DAVID program,30 which provides comprehensive functional annotations to understand biological mechanisms associated with DEGs. From the above results, we found that ‘proteolysis involved in cellular protein catabolic process (P-value<1.4 × E-02)’, ‘cellular protein catabolic process (P-value<1.4 × E-02)’, ‘enzyme-linked receptor protein signaling pathway (P-value<4.3 × E-04)’ and ‘regulation of glucokinase activity (P-value<2.2 × E-02)’ were identified as the most highly enriched pathways.
To better understand the pattern of enriched pathways responding to ERT, the expression profiles of genes associated with protein catabolic processes were plotted in Figure 2. As shown in Figure 2a, all genes associated with two protein catabolism-related pathways (‘proteolysis involved in cellular protein catabolic process’ and ‘cellular protein catabolic process’) were highly upregulated right after ERT in the male patients. However, in the female patients (Figure 2b), the genes associated with those pathways were more likely randomly perturbed or downregulated right after ERT. Thus we could observe that male patients responded more actively to ERT by stimulating the protein catabolic pathway-related genes. Meanwhile, the ‘regulation of cell cycle’ pathway-related genes were enriched (P-value<3.0 × E-02) in the female patients, although the significance was not critical.
Expression patterns of genes associated with the top two highly enriched pathways: ‘proteolysis involved in cellular protein catabolic process’ and ‘cellular protein catabolic process’. (a) Expression profiles of genes belonging to the above two pathways in male patients. (b) Expression profiles of the same genes in female patients. A full color version of this figure is available at the Journal of Human Genetics journal online.
Pathway analysis with GSEA (Gene Set Enrichment Analysis)
In order to understand the molecular basis associated with ERT in FD, we performed GSEA analysis. GSEA31 is a computational method that determines the statistical significance of predefined genes (for example, biological pathways) related with two biological states. Figure 3 represents the upregulated and downregulated pathways responding to ERT. First of all, immune-related pathways (for example, innate/adaptive pathway, interleukin 2 (IL2)-related pathway, T-cell receptor pathway and so on) were identified as highly upregulated pathways. Such enrichment of immune reaction was very natural owing to injection of enzyme. Growth factor-related pathways, especially associated with a cell proliferation, such as EGF_pathway and PDFG_pathway were also activated after ERT, and inflammatory-related pathways were also detected as activated pathways. Recent study32 showed that globotriaosylceramide is correlated with inflammation in Fabry patients treated with ERT. Furthermore, interestingly, we identified an oxidative phosphorylation pathway as a downregulated one after ERT. As oxidative damage is usually increased in patients with FD and ERT has a role to protect from the protein oxidative damage,32 such a deactivation of oxidative phosphorylation was very critical process indeed for treatment.
Network analyses of DEGs in male patients
We investigated the functional similarity of the DEGs through a network analysis, which might reveal hidden regulatory relationships. In this study, we used HumanNet,33 which is a probabilistic functional gene network constructed by a modified Bayesian integration of various types of genomic data from multiple organisms. It covers about 87% of the protein-coding genes in humans. Figure 4 represents the functional connections of the DEGs in the male patients. As we can see in Figure 4, most of the significant DEGs are directly or indirectly connected with the overall upregulation patterns, which reveal the functional connections among the DEGs. Such functional modules associated with DEGs demonstrate that those DEGs are not just randomly expressed but are more likely perturbed under functionally common biological mechanisms. For example, even though two significantly upregulated genes such as CDK5R1 and TINAG1 (Module 2) are not directly connected to each other, they share many common genes in the functional network, which leads to the overall upregulation patterns, including the immediate immune reaction under ERT. The module composed of ERLIN1 (Module 3) and its neighborhoods was functionally enriched for ‘AMP biosynthetic process’ (P-value<2.4 × E-3) and ‘AMP metabolic process’ (P-value<4.1 × E-3). Figure 4 show the module annotations (Modules 1, 2 and 3) and corresponding enriched gene ontology terms associated with these modules. Interestingly, each of those modules represents ERT response-related pathways. The functionally connected genes in Module 1 are mostly related to antibody response-related pathways, which supports our observation of the occurrence of antibodies in the male patients. Similarly, Modules 2 and 3 show the enrichment of cell cycle regulation- and immune-related pathways. In the next section, we further investigate the significant DEGs in detail and show that they are closely related to known clinical manifestations of FD.
Functionally connected DEGs in HumanNet in male patients (P-value<0.01): Overview of functional networks around the differentially expressed genes. The connected genes in HumanNet are marked as modules (Module1: red circle, Module 2: blue circle, Module 3: functionally connected gene set in the center). The enriched biological pathways for each module are also listed. Note that node size represents the significance of differential expression (∝ reverse of P-value), and the bigger the node size, the more differentially expressed the gene. The border color represents the upregulated/downregulated pattern. The light green color demonstrates the genes with an upregulated pattern after ERT, and the black color represents the genes with a downregulated pattern after ERT. The deeper the color, the larger the fold change log ratio. The nodes with red color represent differentially expressed genes under ERT (seed genes) while the nodes with cyan color represent the genes that are directly connected with the seed genes even though they were not differentially expressed in this experiment. A full color version of this figure is available at the Journal of Human Genetics journal online.
Analysis of DEGs in male patients with FD
The up-regulated genes, such as IL18R1 and IL1R1 on chromosome 2, form a gene cluster, and they are known to be associated with various clinical features of FD, including pyogenic arthritis, pyoderma gangrenosum and acne (Table 1). ER lipid raft associated 1 (ERLIN1) is also identified as one of the upregulated DEGs after ERT in the male patients. It is a component of the ENLIN1/ERLIN2 complex, which mediates the endoplasmic reticulum-associated degradation of inositol 1, 4, 5-trisphosphate receptors. In FD, the degradation of mutant lysosomal alpha-galactosidase was demonstrated to occur within the endoplasmic reticulum-associated degradation as a result of the misfolding of mutant proteins.34 Such upregulation of ERLIN1 in male patients with FD would be closely related to the degradation process of alpha-galactosidase A. In addition, DAAM2, which helps to alleviate the retinitis-related symptoms in male patients, also showed very high fold changes in its perturbation patterns after ERT. Analysis of the DEGs revealed that most of the genes are functionally related to clinical symptoms of FD in male patients, and such molecular-level changes due to ERT are well explained by the active relief of symptoms in male patients. Such clinical effectiveness of ERT in male patients was also shown in several studies based on quantifiable clinical parameters or manifestations such as measurements of pain, renal, cardiac and cerebral outcomes and health-related quality of life.14, 15, 16, 17, 18, 35
Discussion
Male patients had more active responses to ERT, which were especially related to protein catabolism-related pathways. We further investigated the biological mechanism of the enrichment of protein catabolism-related pathways in male patients with FD. Recent studies reported the occurrence and effects of antibodies.19, 20, 21, 22 The incidence of antibodies toward the infused enzyme is commonly observed in male patients with FD and is related to negative impacts on Gb3 reduction.20 According to recent studies,4, 19 antibody emergence occurs in more than half of male patients with FD regardless of enzyme types, such as Agalsidase alpha or beta, but not in any female patients. Because female patients usually have residual enzyme activities owing to X-chromosome inactivation, the infused enzyme is not recognized as immunogenic. Therefore, male patients have a greater tendency to be exposed to the antibodies than female patients. We observed that male patients were more actively responsive to ERT than female patients and protein catabolic process-related pathways were significantly upregulated in male patients. This suggests that the activation of protein catabolic processes in the male patients is closely related to the incidence of antibodies after ERT. We will further investigate the direct connection between them in the future. In summary, our study shows the molecular-level perturbation caused by ERT in patients with FD. To our knowledge, this is the first study to observe the short-term effectiveness of ERT through the comprehensive comparison of transcriptome data with next-generation sequencing data. In this study, we could only collect six samples from patients with FD owing to the rareness of the disease; a limitation of the study was the relatively small sample size, thus we are planning to collect more samples in future to further explain more detailed pharmacodynamics of ERT. We note that FD usually affect endothelial cells, cardiomyocytes, podocytes and and so on; however, these cells are very hard to be efficiently isolated in large numbers with high purity. Thus this study was carried out by comparing the molecular profiles before and after ERT in blood cells. We hope that these limitations can be addressed in future research studies.
In conclusion, we investigated the mechanism of action of ERT based on molecular-level perturbations in Korean patients with FD through a transcriptome study. Our molecular pathway-level study reveals that male patients are more actively responsive to ERT than female patients in the short term. Protein catabolism-related pathways and inflammation/immune-related pathways are especially upregulated right after the course of ERT in male patients, which explains the emergence of antibodies owing to the infused enzymes while oxidative phosphorylation pathway is identified as a downregulated one owing to the role of ERT to protect the protein oxidative damage. Our study provides valuable information for understanding the direct molecular-level response to ERT and further clarifying the optimal care for patients with FD.
References
Brady, R. O., Gal, A. E., Bradley, R. M., Martensson, E., Warshaw, A. L. & Laster, L. Enzymatic defect in Fabry's disease. Ceramidetrihexosidase deficiency. N. Engl. J. Med. 276, 1163–1167 (1967).
Pollack, A . in The New York Times. Genzyme Drug Shortage Leaves Users Feeling Betrayed (2010). http://www.nytimes.com/2010/04/16/business/16genzyme.html. Accessed on 13 March 2015.
Bouwman, M. G., de Ru, M. H., Linthorst, G.E., Hollak, C.E., Wijburg, F.A. & van Zwieten, M. C. Fabry patients' experiences with the timing of diagnosis relevant for the discussion on newborn screening. Mol. Genet. Metab. 109, 201–207 (2013).
Hughes, D. A., Elliott, P. M., Shah, J., Zuckerman, J., Coghlan, G., Brookes, J. et al. Effects of enzyme replacement therapy on the cardiomyopathy of Anderson-Fabry disease: a randomised, double-blind, placebo-controlled clinical trial of agalsidase alfa. Heart 94, 153–158 (2008).
Germain, D. P., Waldek, S., Banikazemi, M., Bushinsky, D. A., Charrow, J., Desnick, R. J. et al. Sustained, long-term renal stabilization after 54 months of agalsidase beta therapy in patients with Fabry disease. J. Am. Soc. Nephrol. 18, 1547–1557 (2007).
Schiffmann, R., Kopp, J. B., Austin, H. A. 3rd, Sabnis, S., Moore, D. F., Weibel, T. et al. Enzyme replacement therapy in Fabry disease: a randomized controlled trial. JAMA 285, 2743–2749 (2001).
Eng, C. M., Banikazemi, M., Gordon, R. E., Goldman, M., Phelps, R., Kim, L. et al. A phase 1/2 clinical trial of enzyme replacement in fabry disease: pharmacokinetic, substrate clearance, and safety studies. Am. J. Hum. Genet. 68, 711–722 (2001).
Schiffmann, R., Ries, M., Timmons, M., Flaherty, J. T. & Brady, R. O. Long-term therapy with agalsidase alfa for Fabry disease: safety and effects on renal function in a home infusion setting. Nephrol. Dial. Transplant. 21, 345–354 (2006).
Hoffmann, B., Beck, M., Sunder-Plassmann, G., Borsini, W., Ricci, R., Mehta, A. et al. Nature and prevalence of pain in Fabry disease and its response to enzyme replacement therapy—a retrospective analysis from the Fabry Outcome Survey. Clin. J. Pain 23, 535–542 (2007).
Byeon, S. K., Lee, J. Y., Lee, J. S. & Moon, M. H. Lipidomic profiling of plasma and urine from patients with Gaucher disease during enzyme replacement therapy by nanoflow liquid chromatography-tandem mass spectrometry. J. Chromatogr. A 1381, 132–139 (2015).
Schiffmann, R., Pastores, G. M., Lien, Y. H., Castaneda, V., Chang, P., Martin, R. et al. Agalsidase alfa in pediatric patients with Fabry disease: a 6.5-year open-label follow-up study. Orphanet J. Rare Dis. 9, 169 (2014).
Ferreira, S., Ortiz, A., Germain, D. P., Viana-Baptista, M., Caldeira-Gomes, A., Camprecios, M. et al. The alpha-galactosidase A p.Arg118Cys variant does not cause a Fabry disease phenotype: data from individual patients and family studies. Mol. Genet. Metab. 114, 248–258 (2015).
Furujo, M., Kubo, T., Kobayashi, M. & Ohashi, T. Enzyme replacement therapy in two Japanese siblings with Fabry disease, and its effectiveness on angiokeratoma and neuropathic pain. Mol. Genet. Metab. 110, 405–410 (2013).
Lidove, O., West, M. L., Pintos-Morell, G., Reisin, R., Nicholls, K., Figuera, L. E. et al. Effects of enzyme replacement therapy in Fabry disease—a comprehensive review of the medical literature. Genet. Med. 12, 668–679 (2010).
Rombach, S. M., Smid, B. E., Bouwman, M. G., Linthorst, G. E., Dijkgraaf, M. G. & Hollak, C. E. Long term enzyme replacement therapy for Fabry disease: effectiveness on kidney, heart and brain. Orphanet J. Rare Dis. 8, 47 (2013).
Rombach, S. M., Hollak, C. E., Linthorst, G. E. & Dijkgraaf, M. G. Cost-effectiveness of enzyme replacement therapy for Fabry disease. Orphanet J. Rare Dis. 8, 29 (2013).
Mehta, A., Beck, M., Elliott, P., Giugliani, R., Linhart, A., Sunder-Plassmann, G. et al. Enzyme replacement therapy with agalsidase alfa in patients with Fabry's disease: an analysis of registry data. Lancet 374, 1986–1996 (2009).
Vedder, A. C., Breunig, F., Donker-Koopman, W. E., Mills, K., Young, E., Winchester, B. et al. Treatment of Fabry disease with different dosing regimens of agalsidase: effects on antibody formation and GL-3. Mol. Genet. Metab. 94, 319–325 (2008).
Rombach, S. M., Aerts, J. M., Poorthuis, B. J., Groener, J. E., Donker-Koopman, W., Hendriks, E. et al. Long-term effect of antibodies against infused alpha-galactosidase A in Fabry disease on plasma and urinary (lyso)Gb3 reduction and treatment outcome. PLoS ONE 7, e47805 (2012).
Linthorst, G. E., Hollak, C. E., Donker-Koopman, W. E., Strijland, A. & Aerts, J. M. Enzyme therapy for Fabry disease: neutralizing antibodies toward agalsidase alpha and beta. Kidney Int. 66, 1589–1595 (2004).
Benichou, B., Goyal, S., Sung, C., Norfleet, A. M. & O'Brien, F. A retrospective analysis of the potential impact of IgG antibodies to agalsidase beta on efficacy during enzyme replacement therapy for Fabry disease. Mol. Genet. Metab. 96, 4–12 (2009).
Hollak, C. E. & Linthorst, G. E. Immune response to enzyme replacement therapy in Fabry disease: impact on clinical outcome? Mol. Genet. Metab. 96, 1–3 (2009).
Kakkis, E. D., O'Donovan, M., Cox, G., Hayes, M., Goodsaid, F., Tandon, P. K. et al. Recommendations for the development of rare disease drugs using the accelerated approval pathway and for qualifying biomarkers as primary endpoints. Orphanet J. Rare Dis. 10, 16 (2015).
Berger, M. F., Levin, J. Z., Vijayendran, K., Sivachenko, A., Adiconis, X., Maguire, J. et al. Integrative analysis of the melanoma transcriptome. Genome Res. 20, 413–427 (2010).
Ko, Y., Ament, S. A., Eddy, J. A., Caballero, J., Earls, J. C., Hood, L. et al. Cell type-specific genes show striking and distinct patterns of spatial expression in the mouse brain. Proc. Natl Acad. Sci. USA 110, 3095–3100 (2013).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Trapnell, C., Pachter, L. & Salzberg, S. L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Dillies, M.A., Rau, A., Aubert, J., Hennequet-Antier, C., Jeanmougin, M., Servant, N. et al. A comprehensive evaluation of normalization methods for Illumina high-throughput RNA sequencing data analysis. Brief. Bioinform. 14, 671–683 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Huang, D. W., Sherman, B. T., Tan, Q., Collins, J. R., Alvord, W. G., Roayaei, J. et al. The DAVID Gene Functional Classification Tool: a novel biological module-centric algorithm to functionally analyze large gene lists. Genome Biol. 8, R183 (2007).
Subramanian, A., Tamayo, P., Mootha, V. K., Mukherjee, S., Ebert, B. L., Gillette, M. A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Biancini, G. B., Vanzin, C. S., Rodrigues, D. B., Deon, M., Ribas, G. S., Barschak, A. G. et al. Globotriaosylceramide is correlated with oxidative stress and inflammation in Fabry patients treated with enzyme replacement therapy. Biochim. Biophys. Acta 1822, 226–232 (2012).
Lee, I., Blom, U. M., Wang, P. I., Shim, J. E. & Marcotte, E. M. Prioritizing candidate disease genes by network-based boosting of genome-wide association data. Genome Res. 21, 1109–1121 (2011).
Fan, J. Q. & Ishii, S. Active-site-specific chaperone therapy for Fabry disease. Yin and Yang of enzyme inhibitors. FEBS J. 274, 4962–4971 (2007).
MacDermot, K. D., Holmes, A. & Miners, A. H. Anderson-Fabry disease: clinical manifestations and impact of disease in a cohort of 60 obligate carrier females. J. Med. Genet. 38, 769–775 (2001).
Acknowledgements
This study was supported by a grant (HI12C-002) of the Korea Healthcare Technology R&D Project. This work was also supported by the Supercomputing Center/Korea Institute of Science and Technology Information with supercomputing resources, including technical support (KSC-2014-C1-017).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Ko, Y., Lee, C., Moon, M. et al. Unravelling the mechanism of action of enzyme replacement therapy in Fabry disease. J Hum Genet 61, 143–149 (2016). https://doi.org/10.1038/jhg.2015.123
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2015.123
This article is cited by
-
Inflammation across the spectrum of hypertrophic cardiac phenotypes
Heart Failure Reviews (2023)