Abstract
Abnormalities in mitochondrial complex I, which is responsible for controlling mitochondrial function, have been implicated in a variety of diseases associated with mitochondrial dysfunction, potentially including schizophrenia. The NADH dehydrogenase Fe–S protein 1 (NDUFS1) is the largest subunit of complex I. To explore whether the encoding NDUFS1 gene confers susceptibility to schizophrenia or is associated with the severity of typical symptoms of schizophrenia, we recruited 519 stable schizophrenia patients receiving clozapine treatment and 594 healthy controls for genotyping to investigate the association of four selected tagging single-nucleotide polymorphisms (SNPs) of NDUFS1 and both schizophrenia risk and symptom severity. The severity of psychotic symptoms was evaluated using the Positive and Negative Syndrome Scale and then tested for association with the four SNPs. The SNP rs1044120 showed significant association with schizophrenia (adjusted P=0.032). The frequency of the G allele of rs1044120 was significantly higher in patients than among the healthy controls (adjusted P=0.008). Stratification by sex revealed a significant association between the rs1044120 polymorphism and schizophrenia among males (adjusted P=0.036 and 0.008 in genotypic and allelic comparisons, respectively). We also observed a significant difference in the negative symptom scores among the three genotypes among these males (adjusted P=0.036). Post hoc comparisons showed that rs1044120 G/G carriers had higher negative symptom scores than those with G/T and T/T carriers (raw P=0.035 and 0.005, respectively). Our findings suggest that NDUFS1 may confer susceptibility to schizophrenia in male subjects, acting as a causative factor for the severity of negative symptoms in schizophrenia.
Similar content being viewed by others
Introduction
While schizophrenia affects only ~1% of the global population, the severity of the impairment it can cause ranks it among the leading causes of mental health disease burden worldwide. Since schizophrenia is a complex neuropsychiatric disorder, elucidating its underlying cellular and molecular pathology can be difficult;1 therefore, improving the understanding of its etiology and pathogenesis has become increasingly important to modern psychiatric research.2 Previous epidemiological studies have demonstrated that schizophrenia is a familial disorder with heritability reaching >80%, but the mode of its inheritance and heritability is quite complex and varies among different populations, as do the clinical symptoms and their severity.3, 4 This leaves efforts to better understand schizophrenia fraught with the problem of having to deal with a clinically and genetically heterogeneous disorder.3
Data accumulated to date point to the role of abnormal brain energy metabolism in schizophrenia,5 implicating defects in mitochondrial function as potential factors in both the risk and severity of schizophrenia.6 For example, several clinical studies on patients with schizophrenia have noted abnormalities in the oxidative phosphorylation system enzyme activity as well as mRNA and protein expression in both the periphery and the brain.7 The oxidative phosphorylation system system is controlled by mitochondrial complex I, the first, largest and most intricate multimeric component of the mitochondrial respiratory chain; abnormalities in this complex have been suggested to lead to a variety of diseases associated with mitochondrial dysfunction (for example, Parkinson’s disease, Alzheimer’s disease, type 2 diabetes mellitus).8,9 This complex’s largest subunit, the NADH dehydrogenase Fe–S protein 1 (NDUFS1), is localized in the iron–sulfur center of the hydrophilic arm and is involved in NADH dehydrogenase and oxidoreductase activity, transferring electrons from NADH to the respiratory chain. The human gene encoding the NDUFS1 protein is located in chromosome 2q33-34, a candidate region previously implicated in linkage studies of schizophrenia.10 Unsurprisingly, genetic variations in NDUFS1 have also been reported to be a cause of complex I deficiency and apoptosis, implying further outcomes for respiratory chain disorders.11,12 Building on this premise, it is plausible that NDUFS1 may be a promising candidate gene for schizophrenia susceptibility or symptom severity. To the best of our knowledge, no genetic study has addressed this potential association.
In the present study, we hypothesized that NDUFS1 may be associated with schizophrenia, and investigated whether the presence of common variants within NDUFS1 confers susceptibility to schizophrenia among a Chinese Han population. As a secondary aim, since mitochondrial function and energy metabolism are exhibited in a sex-specific manner,13 we wanted to examine whether there were any observable sex-specific differences in the potential correlation between NDUFS1 and schizophrenia. Third, since schizophrenia is a heterogeneous and polygenic disorder, wherein specific genetic loci may contribute to the pathophysiology and phenotypic variability, as previously seen in schizophrenia cases,14 we wanted to see whether NDUFS1 is associated with any specific symptom clusters.
Materials and methods
Subjects
A total of 519 schizophrenia patients and 594 healthy controls were enrolled in this study. All subjects provided written informed consent prior to performing any of the procedures related to this study. All procedures were reviewed and approved by the Institutional Review Boards of the Shanghai Mental Health Center, and performed in strict accordance with the Declaration of Helsinki, and other relevant national and international regulations.
Detailed information on the patients included in the previous study has already been published.15, 16, 17 In brief, patients were included based on the following criteria: (1) they met the Diagnostic and Statistical Manual of Mental Disorders, Fourth Edition (DSM-IV) criteria for schizophrenia; (2) they had received clozapine treatment; (3) they had maintained a stable condition for more than 6 months before entry into the study; and (4) they had no physical disease or other psychiatric disorder aside from schizophrenia. First-episode patients were excluded because initial diagnoses are often unreliable.2,18 For this study, the diagnosis and review of psychiatric case records were independently checked and verified by two experienced psychiatrists to ensure consistency. The 594 healthy controls were interviewed by a specialized psychiatrist using the Structured Clinical Interview for DSM-IV-TR Axis I Disorders—Patient Edition; subjects with any prior or current psychiatric disorder and/or chronic physical disease were excluded from our analysis. All subjects in both the healthy control group and the schizophrenia group were of Han Chinese origin. Demographic characteristics for the control and schizophrenia groups are summarized in Supplementary Table S1.
Clinical symptom assessment
The severity of psychotic symptoms exhibited by the schizophrenia patients was evaluated using the Positive and Negative Syndrome Scale (PANSS), which was used to evaluate the clinical status of schizophrenia. PANSS evaluations were conducted by three experienced psychiatrists who had simultaneously attended a training session in the use of PANSS. Repeated assessments found that a correlation coefficient >0.8 was maintained for the PANSS total scores.
Single-nucleotide polymorphism (SNP) selection and genotyping
The principal hypothesis underlying this study is that common SNPs in NDUFS1 may confer susceptibility to schizophrenia. In total, four selected tagging SNPs of NDUFS1 were genotyped to test for possible association. To set the inclusion criteria for tagging SNPs, we retrieved CHB data from the HapMap database (http://www.hapmap.org) and defined linkage disequilibrium (LD) blocks using Haploview 4.2 (Broad Institute, Cambridge, MA, USA). Haplotype-tagging SNPs (htSNPs) were selected with an r2 cutoff >0.8 and minor allele frequency (MAF) >0.1, and four htSNPs were captured (Supplementary Table S2). Genomic DNA of all participants was extracted from peripheral blood using a Tiangen DNA Isolation Kit (Tiangen Biotech, Beijing, China). The four htSNPs were amplified independently via polymerase chain reaction (PCR) and genotyped by direct sequencing using an ABI PRISM 3730 Genetic Analyzer (Perkin-Elmer Applied Biosystems, Foster City, CA, USA). Genotyping was carried out according to the methods described in our previous studies.15,16 PCR amplification was performed in a volume of 25 μl containing primer pairs for each SNP. PCR primers were also used for sequencing. Sequencing results were handled by DNAStar (DNASTAR Inc., Madison, WI, USA), and the original sequencing chromatograms of each sample were then manually checked.
Quality control methods were based on our previous work,3,17,19 with 5% of samples randomly selected and independently sequenced to evaluate genotype accuracy. Here, our re-sampling results were 100% concordant with the initial results.
Statistical analysis
Hardy–Weinberg equilibrium testing, and allele and genotype frequency analysis were conducted using SHEsis (http://analysis.bio-x.cn). Pairwise LD of all pairs of htSNPs was performed using HaploView 4.2 (Broad Institute), and the extent of LD was measured by the standardized D′ and r2. Power calculations were carried out with Quanto 1.2.3 (http://hydra.usc.edu/GxE). To adjust for multiple testing, the level of significance was corrected via Bonferroni correction. Statistical significance of differences between clinical features and polymorphisms in patients was assessed using ANCOVA, with variables that may potentially affect the symptoms (for example, age, sex, course of disease, treatment duration and doses) as covariates. Effect sizes were measured with Cohen’s d.
Results
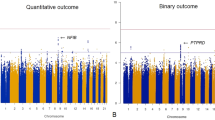
Initial analysis showed a significant difference between the schizophrenia patients and control subjects with respect to age (P<0.01), but not sex (P=0.33). Genotype distributions revealed no deviation from Hardy–Weinberg equilibrium in either healthy cases or controls (allele and genotype frequencies of the four NDUFS1 SNPs are shown in Table 1). In our sampled population, the SNP rs1044120 showed significant association with schizophrenia (P=0.008; P=0.032 following Bonferroni correction). The frequency of G allele of rs1044120 was significantly higher among schizophrenia patients than among the controls (OR=1.36, 95% CI: 1.12–1.65, P=0.002; P=0.008 after Bonferroni correction). After stratifying the samples by sex, we noted significant association between the rs1044120 polymorphism and schizophrenia in male subjects (corrected P=0.036 and 0.008 in genotypic and allelic comparisons, respectively), with no significant difference observed among females. Analysis of pairwise LD was conducted in SNP pairs. Generally, the SNPs were not in significant LD. Further analysis showed a strong LD between rs1044120 and rs6435330 polymorphisms that was only found in females, but not in males (Supplementary Figure S1). In view of this strong LD, we performed a two-SNP haplotype analysis, analyzing only those common haplotypes with at least 3% of frequency in either case or control samples (P-values corresponding to the haplotypes are shown in Supplementary Table S3). No significant difference was found for any haplotype in females.
We next examined the relationship between the rs1044120 polymorphism and psychopathologic symptoms in the male schizophrenia patients by comparing the scores of four factors of the PANSS scale with genotypes of the rs1044120 polymorphism, which showed a significant difference in the negative symptom scores among the three genotypes in males (P=0.009; P=0.036, after Bonferroni correction, shown in Table 2). Post hoc comparisons showed that rs1044120 G/G carriers had higher negative symptom scores than G/T and T/T carriers (P=0.035 (Cohen’s d=0.27) and P=0.005 (Cohen’s d=0.49), respectively; Figure 1).
The power of the four SNPs was >80% (for α=0.05) for schizophrenia samples, with a genotype relative risk of 1.5, under the assumption of a log additive model and disease prevalence of 1%. Analysis of continuous variables and not categorical variables holds the advantage of an increased statistical power of ~50%. Accordingly, the likelihood of a type II error in our sample should be considerably low.
Discussion
To date, the function and dysfunction of mitochondrial complex I has been widely studied in connection with several different neuropsychiatric disorders. For example, deficiencies in the activity of complex were suggested to be a potentially important cause of (or at least contributing factor to) schizophrenia.20,21 Complex I is composed of at least 43 structural subunits encoded in both nuclear and mitochondrial DNA, most of which are encoded by nuclear genes, with only seven being mitochondrially encoded.22 In this study, we attempted to investigate the association between the nuclear genes of complex I and schizophrenia in Han Chinese, and found significant differences in allele and genotype frequencies at the rs1044120 polymorphism between patients diagnosed with schizophrenia and healthy controls. After stratification by sex, this significant association only held in males, but not in females. Further analysis showed a positive association between rs1044120 polymorphism and severity of negative symptoms in male schizophrenia patients. However, we did not find any association between complex I-associated genes and schizophrenia, including genes encoding Leucine-rich repeat kinase 2 (LRRK2),23 NADH dehydrogenase (ubiquinone) Fe–S protein 7 (NDUFS7)9 and NADH dehydrogenase (ubiquinone) flavoprotein 2 (NDUFV2).24
On the whole, the present results suggested that NDUFS1 may potentially contribute to schizophrenia susceptibility. The NDUFS1 gene encodes the largest subunit of complex I, which acts as a critical caspase substrate in the mitochondria and is responsible for the disruption of mitochondrial function during apoptosis.11 Apoptosis generally serves to eliminate excess neurons and maintain synaptic stability in the brain,25 but mounting evidence suggests that altered apoptosis may be involved in the abnormal neurodevelopment and neurodegenerative processes previously associated with schizophrenia.26 Moreover, the rs1044120 polymorphism is located at the 3′-untranslated region (3′-UTR) of NDUFS1, and while the effect of 3′-UTR on gene expression or function is not well known it is possible that variants on the 3′-UTR could potentially produce changes in messenger RNA processing.27 If accurate, then genetic variations in the 3′-UTR of NDUFS1 may result in complex I deficiency and alter apoptosis in the brain, and by extension help to account for the results we encountered in this study. Unfortunately, further research is needed to verify this supposition, since the exact role of rs1044120 polymorphism in the function of the NDUFS1 protein is not yet known, even though we have some preliminary results suggesting an association.
A number of studies have demonstrated that sex-specific patterns are commonly seen in schizophrenia in terms of age at onset, symptomatology, cognitive impairment, treatment response or even the pathophysiology.28, 29, 30, 31 Here, following sex stratification we noted a positive association of the NDUFS1 gene and schizophrenia among male patients. These results suggest that sex-specific regulation may also play an important role in the association between the NDUFS1 and schizophrenia. This would not be unexpected, since previous studies demonstrated that NDUFS1 is correlated with inefficiency of oxidative phosphorylation in a sex-specific manner.32,33 Similarly, Misiak et al.34 indicated that neuronal cell death due to increased mitochondrial impairment is more pronounced in males than in females, and that male neurons exhibited an ~1.5–2-fold higher number of apoptotic and necrotic cells as compared to those found in females. By extension, male neuronal cells should have diminished respiratory chain activity and subsequently lower ATP levels. Again, presuming this extrapolation is correct, our findings suggest that disrupted energy metabolism in the brain induced by NDUFS1 may potentially be more sensitive in males than in females, thereby accounting for the sex-specific association between the NDUFS1 and schizophrenia. Further targeted research may either better verify this possibility or suggest a more viable alternative.
One inherent difficulty in studies on schizophrenia is the clinical heterogeneity of this disorder that can preclude or hinder more detailed inquiries into the disorder’s genetic structure, since schizophrenia manifests in a constellation of symptoms, including positive symptoms, negative symptoms and cognitive dysfunctions. This obstacle could be somewhat offset by better clinical phenotyping based on symptomology that would allow for more accurate delineation of the underlying genetic mechanism at play in the various manifested symptoms of schizophrenia.35 To that end, we further analyzed the association of NDUFS1 with this disorder in male subjects and found that NDUFS1 rs1044120 G/G carriers have higher negative symptom scores than G/T and T/T carriers. This finding may imply an association between NDUFS1 and negative symptoms. Negative symptoms are generally characterized by affective blunting, apathy and social withdrawal.36 Positron emission tomography previously showed that schizophrenia patients with predominantly negative symptoms have greater metabolic abnormalities than subjects with predominantly positive symptoms, particularly in frontal, temporal and cerebella circuitry.37 Another study using 31P-magnetic resonance spectroscopy reported that ATP in the frontal lobe among patients with schizophrenia was correlated with negative symptoms.38 These lines of evidence suggest that aberrant brain metabolism induced by NDUFS1 variation may be a causative factor for the severity of negative symptoms in schizophrenia patients. If further study can verify this possibility, our present findings may serve as foundational evidence in developing more detailed phenotypic symptomology that can assist mental health researchers in working past some of the inherent difficulties in characterizing the underlying genetic or molecular bases of schizophrenia.
Despite the promising implications of our results, there are three methodological limitations that should be noted. First, the present report details an exploratory study done on a subset of the overall Han Chinese population. The size of our sample is modest, especially for the purposes of sex stratification. As such, the findings are largely suggestive of potentially provocative trends, but need validation through studies using a larger sample size. Second, selection bias is inevitable in such studies. In this case, our sample was recruited from the Shanghai region, and may not be representative of the Han Chinese population in general, nor other closely related populations in the region. Third, a consensus five-factor model of PANSS has recently been proposed that may better represent the clinical symptom profiles of patients,39 since it may be more sensitive to capture PANSS structure in schizophrenia patients as compared with the three-subscale PANSS that we used in this study.
In conclusion, in the present study we explored the potential relationship between genetic variations in the NDUFS1 gene and schizophrenia among a Han Chinese population. Our findings suggest that NDUFS1 may confer susceptibility to schizophrenia in male subjects, potentially serving as a causative factor for the severity of negative symptoms exhibited in schizophrenia. Further investigation will be required to uncover the exact role of NDUFS1 in the development and heritability of schizophrenia, as well as the impact of genetic variations to this gene as a causative factor for the severity symptoms manifested by among male patients with schizophrenia.
References
Zhang, C., Zhang, J., Fan, J., Cheng, W., Du, Y., Yu, S. et al. Identification of ANKK1 rs1800497 variant in schizophrenia: new data and meta-analysis. Am. J. Med. Genet. B Neuropsychiatr. Genet. 165, 564–571 (2014).
Tang, W. X., Cai, J., Yi, Z. H., Zhang, Y., Lu, W. H. & Zhang, C. Association study of common variants within the G protein-coupled receptor kinase 6 gene and schizophrenia susceptibility in Han Chinese. Hum. Psychopharm. Clin. 29, 100–103 (2014).
Zhang, C., Cai, J., Zhang, J., Li, Z., Guo, Z., Zhang, X. et al. Genetic modulation of working memory deficits by ankyrin 3 gene in schizophrenia. Prog. Neuropsychopharmacol. Biol. Psychiatry 50, 110–115 (2014).
Goldman, A. L., Pezawas, L., Mattay, V. S., Fischl, B., Verchinski, B. A., Zoltick, B. et al. Heritability of brain morphology related to schizophrenia: a large-scale automated magnetic resonance imaging segmentation study. Biol. Psychiatry 63, 475–483 (2008).
Robicsek, O., Karry, R., Petit, I., Salman-Kesner, N., Muller, F. J., Klein, E. et al. Abnormal neuronal differentiation and mitochondrial dysfunction in hair follicle-derived induced pluripotent stem cells of schizophrenia patients. Mol. Psychiatry 18, 1067–1076 (2013).
Ben-Shachar, D. The interplay between mitochondrial complex I, dopamine and Sp1 in schizophrenia. J. Neural Transm. 116, 1383–1396 (2009).
Ben-Shachar, D. & Karry, R. Neuroanatomical pattern of mitochondrial complex I pathology varies between schizophrenia, bipolar disorder and major depression. PLos ONE 3, e3676 (2008).
Distelmaier, F., Koopman, W. J. H., van den Heuvel, L. P., Rodenburg, R. J., Mayatepek, E., Willems, P. H. G. M. et al. Mitochondrial complex I deficiency: from organelle dysfunction to clinical disease. Brain 132, 833–842 (2009).
Ma, L., Zhang, W., Tang, J. S., Tan, L. W., Yao, Y. G. & Chen, X. G. No association between genetic polymorphisms of the NDUFS7 gene and schizophrenia in Han Chinese. Psychiatr. Genet. 23, 29–32 (2013).
Aberg, K., Axelsson, E., Saetre, P., Jiang, L., Wetterberg, L., Pettersson, U. et al. Support for schizophrenia susceptibility locus on chromosome 2q detected in a Swedish isolate using a dense map of microsatellites and SNPs. Am. J. Med. Genet. B 147B, 1238–1244 (2008).
Ricci, J. E., Munoz-Pinedo, C., Fitzgerald, P., Bailly-Maitre, B., Perkins, G. A., Yadava, N. et al. Disruption of mitochondrial function during apoptosis is mediated by caspase cleavage of the p75 subunit of complex I of the electron transport chain. Cell 117, 773–786 (2004).
Ferreira, M., Torraco, A., Rizza, T., Fattori, F., Meschini, M. C., Castana, C. et al. Progressive cavitating leukoencephalopathy associated with respiratory chain complex I deficiency and a novel mutation in NDUFS1. Neurogenetics 12, 9–17 (2011).
Velarde, M. C. Mitochondrial and sex steroid hormone crosstalk during aging. Longev. Healthspan 3, 2 (2014).
DeRosse, P., Lencz, T., Burdick, K. E., Siris, S. G., Kane, J. M. & Malhotra, A. K. The genetics of symptom-based phenotypes: toward a molecular classification of schizophrenia. Schizophrenia Bull. 34, 1047–1053 (2008).
Zhang, Y., Chen, M., Chen, J., Wu, Z., Yu, S., Fang, Y. et al. Metabolic syndrome in patients taking clozapine: prevalence and influence of catechol-O-methyltransferase genotype. Psychopharmacology (Berl) 231, 2211–2218 (2014).
Zhang, Y., Chen, M., Wu, Z., Chen, J., Yu, S., Fang, Y. et al. Association study of Val66Met polymorphism in brain-derived neurotrophic factor gene with clozapine-induced metabolic syndrome: preliminary results. PLos ONE 8, e72652 (2013).
Cai, J., Zhang, W., Yi, Z., Lu, W., Wu, Z., Chen, J. et al. Influence of polymorphisms in genes SLC1A1, GRIN2B, and GRIK2 on clozapine-induced obsessive-compulsive symptoms. Psychopharmacology (Berl.) 230, 49–55 (2013).
Zhang, C., Li, Z. Z., Shao, Y., Xie, B., Du, Y. S., Fang, Y. R. et al. Association study of tryptophan hydroxylase-2 gene in schizophrenia and its clinical features in Chinese Han population. J. Mol. Neurosci. 43, 406–411 (2011).
Zhang, C., Li, Z. Z., Wu, Z. G., Chen, J., Wang, Z. W., Peng, D. H. et al. A study of N-methyl-D-aspartate receptor gene (GRIN2B) variants as predictors of treatment-resistant major depression. Psychopharmacology 231, 685–693 (2014).
Park, C. & Park, S. K. Molecular links between mitochondrial dysfunctions and schizophrenia. Mol. Cells 33, 105–110 (2012).
Rosenfeld, M., Brenner-Lavie, H., Gal-Ben Ari, S., Kavushansky, A. & Ben-Shachar, D. Perturbation in mitochondrial network dynamics and in complex i dependent cellular respiration in schizophrenia. Biol. Psychiatry 69, 980–988 (2011).
Benit, P., Chretien, D., Kadhom, N., de Lonlay-Debeney, P., Cormier-Daire, V., Cabral, A. et al. Large-scale deletion and point mutations of the nuclear NDUFV1 and NDUFS1 genes in mitochondrial complex I deficiency. Am. J. Hum. Genet. 68, 1344–1352 (2001).
Li, X., Zhang, W., Zhang, C., Gong, W., Tang, J., Yi, Z. et al. No association between genetic variants of the LRRK2 gene and schizophrenia in Han Chinese. Neurosci. Lett. 566, 210–215 (2014).
Zhang, W., Chen, X. G., Gong, W., Tang, J. S., Tan, L. W., Guo, H. et al. Common promoter variants of the NDUFV2 gene do not confer susceptibility to schizophrenia in Han Chinese. Behav. Brain Funct. 6, 75 (2010).
Zhang, C., Wu, Z. G., Hong, W., Wang, Z. W., Peng, D. H., Chen, J. et al. Influence of BCL2 gene in major depression susceptibility and antidepressant treatment outcome. J. Affect. Disorders. 155, 288–294 (2014).
Gasso, P., Mas, S., Molina, O., Lafuente, A., Bernardo, M. & Parellada, E. Increased susceptibility to apoptosis in cultured fibroblasts from antipsychotic-naive first-episode schizophrenia patients. J. Psychiatr. Res. 48, 94–101 (2014).
Conne, B., Stutz, A. & Vassalli, J.D. The 3' untranslated region of messenger RNA: a molecular 'hotspot' for pathology? Nat. Med. 6, 637–641 (2000).
Yi, Z., Zhang, C., Wu, Z., Hong, W., Li, Z., Fang, Y. et al. Lack of effect of brain derived neurotrophic factor (BDNF) Val66Met polymorphism on early onset schizophrenia in Chinese Han population. Brain Res. 1417, 146–150 (2011).
Lu, W. H., Zhang, C., Yi, Z. H., Li, Z. Z., Wu, Z. G. & Fang, Y. R. Association between BDNF Val66Met polymorphism and cognitive performance in antipsychotic-naive patients with schizophrenia. J. Mol. Neurosci. 47, 505–510 (2012).
Wang, L. H., Chen, J. Y., Liou, Y. J., Wang, Y. C., Lai, I. C., Liao, D. L. et al. Association of missense variants of the PRKC, apoptosis, WT1, regulator (PAWR) gene with schizophrenia. Prog. Neuro-Psychopharmacol. 32, 870–875 (2008).
Leung, A. & Chue, P. Sex differences in schizophrenia, a review of the literature. Acta Psychiatr. Scand. 101, 3–38 (2000).
Dato, S., Soerensen, M., Lagani, V., Montesanto, A., Passarino, G., Christensen, K. et al. Contribution of genetic polymorphisms on functional status at very old age: a gene-based analysis of 38 genes (311 SNPs) in the oxidative stress pathway. Exp. Gerontol. 52, 23–29 (2014).
Martin, M. A., Blazquez, A., Gutierrez-Solana, L. G., Fernandez-Moreira, D., Briones, P., Andreu, A. L. et al. Leigh syndrome associated with mitochondrial complex I deficiency due to a novel mutation in the NDUFS1 gene. Arch. Neurol. 62, 659–661 (2005).
Misiak, M., Beyer, C. & Arnold, S. Gender-specific role of mitochondria in the vulnerability of 6-hydroxydopamine-treated mesencephalic neurons. Bba-Bioenergetics 1797, 1178–1188 (2010).
Cherlyn, S. Y., Woon, P. S., Liu, J. J., Ong, W. Y., Tsai, G. C. & Sim, K. Genetic association studies of glutamate, GABA and related genes in schizophrenia and bipolar disorder: a decade of advance. Neurosci. Biobehav. Rev. 34, 958–977 (2010).
Kuperberg, G. & Heckers, S. Schizophrenia and cognitive function. Curr. Opin. Neurobiol. 10, 205–210 (2000).
Potkin, S. G., Alva, G., Fleming, K., Anand, R., Keator, D., Carreon, D. et al. A PET study of the pathophysiology of negative symptoms in schizophrenia. Am. J. Psychiatry 159, 227–237 (2002).
Yacubian, J., de Castro, C. C., Ometto, M., Barbosa, E., de Camargo, C. P., Tavares, H. et al. 31P-spectroscopy of frontal lobe in schizophrenia: alterations in phospholipid and high-energy phosphate metabolism. Schizophr Res. 58, 117–122 (2002).
Wallwork, R. S., Fortgang, R., Hashimoto, R., Weinberger, D. R. & Dickinson, D. Searching for a consensus five-factor model of the Positive and Negative Syndrome Scale for schizophrenia. Schizophr Res. 137, 246–250 (2012).
Acknowledgements
We are deeply grateful to all participants. We thank the two reviewers for their insightful comments. This work was supported by the National Natural Science Foundation of China (81000581 and 81471358), the Shanghai Science & Technology Development Foundation (12140904200), the Medical Health Technique Project of Zhejiang Province (2010KYA047) and the National Key Clinical Disciplines at Shanghai Mental Health Center (OMA-MH, 2011-873).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Zhu, Y., Wang, Z., Ni, J. et al. Genetic variant in NDUFS1 gene is associated with schizophrenia and negative symptoms in Han Chinese. J Hum Genet 60, 11–16 (2015). https://doi.org/10.1038/jhg.2014.94
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2014.94
This article is cited by
-
NDUFV2 pseudogene (NDUFV2P1) contributes to mitochondrial complex I deficits in schizophrenia
Molecular Psychiatry (2020)
-
Effects of omega-3 fatty acids on metabolic syndrome in patients with schizophrenia: a 12-week randomized placebo-controlled trial
Psychopharmacology (2019)
-
Brain-derived neurotrophic factor as a biomarker for cognitive recovery in acute schizophrenia: 12-week results from a prospective longitudinal study
Psychopharmacology (2018)
-
Genetic association analysis of microRNA137 and its target complex 1 with schizophrenia in Han Chinese
Scientific Reports (2017)
-
Do nuclear-encoded core subunits of mitochondrial complex I confer genetic susceptibility to schizophrenia in Han Chinese populations?
Scientific Reports (2015)