Abstract
To date, four families with spinocerebellar ataxia type 5 (SCA5) with four distinct mutations in the spectrin, beta, nonerythrocytic 2 gene (SPTBN2) have been reported worldwide. In the present study, we identified the first Japanese family with SCA5, and analyzed this family clinically and genetically. The clinical features of the five patients in this family revealed late-onset autosomal-dominant pure cerebellar ataxia. We collected DNA samples from the majority of the family members across two generations, and exome sequencing combined with Sanger sequencing revealed a novel heterozygous three-nucleotide in-frame deletion mutation (c.2608_2610delGAG) in exon 14 of the SPTBN2 gene. This mutation cosegregated with the disease in the family and resulted in a glutamic acid deletion (p.E870del) in the sixth spectrin repeat, which is highly conserved in the SPTBN2 gene. This is the first three-nucleotide in-frame deletion mutation in this region of the beta-3 spectrin protein highly likely to be pathogenic based on exome and bioinformatic data.
Similar content being viewed by others
Introduction
Spinocerebellar ataxia type 5 (SCA5) is a rare autosomal-dominant pure spinocerebellar ataxia (SCA). SCA5 patients predominantly exhibit limb and gait ataxia (>90%); however, truncal ataxia, sensory deficits, abnormal eye movements, dysarthria and exaggerated deep tendon reflexes are also prevalent (25–90%).1 Individuals generally struggle with fine motor coordination but rarely require a wheelchair until late in the disease course.1
SCA5 is caused by mutations in the spectrin, beta, nonerythrocytic 2 gene (SPTBN2). This gene is primarily expressed in the brain. It encodes the beta-III spectrin protein consisting of 2390 amino acids comprising two calponin homology domains, 17 triple-helical spectrin repeats and a pleckstrin homology domain, and as a cytoskeletal protein is highly expressed in Purkinje cells.2
To date, five mutations, that is, three missense (p. L253P, p. T472M and R480W) and two deletion (p.E532_M544del and p.L629_R634delinsW) ones, in the SPTBN2 gene have been reported in SCA5 families originating from the United States, France, Germany and Norway,3, 4, 5, 6 and an SCA5 patient.7 In the present study, we report the first Japanese SCA5 family with a novel heterozygous three-nucleotide in-frame deletion mutation in the SPTBN2 gene.
Materials and methods
Patients
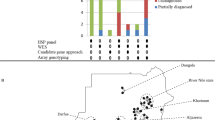
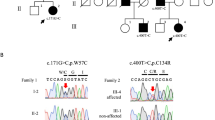
The pedigree is shown in Figure 1a. Eight living family members were interviewed and examined by experienced neurologists, and the diagnosis of familial SCA was made on the basis of the patients’ clinical presentations as well as the results of physiological, radiological and biochemical examinations. Unfortunately, we were not able to obtain the consent of other family members to be examined in this study. Brain magnetic resonance imaging (MRI) in four affected members revealed cerebellar atrophy. Molecular analysis of the proband excluded SCA1, SCA2, MJD, SCA6, SCA7, SCA8, SCA12, SCA17, SCA31, SCA36 and DRPLA. The present clinical and genetic study was approved by the institutional review board of the Yamanashi University, and informed consent was obtained from all participating individuals.
Pedigree of the SCA5 family. (a) Squares and circles indicate males and females, respectively. Filled symbols indicate affected individuals, whereas open symbols indicate unaffected individuals. Slashes indicate the people who have died already. Asterisks indicate the individuals who have been examined clinically and molecularly. (b) Representative sequencing of the SPTBN2 gene in part of the family (II-6, II-8, II-10, III-10, III-12, III-14, III-15 and III-16). The mutation cosegregated with the disease.
Whole-exome sequencing
We carried out whole-exome sequencing of genomic DNA from the proband and the father. Genomic DNA was extracted from blood leukocytes from eight individuals. We used a NEBNest DNA Library Prep Master Mix set (New England Biolabs, Ipswich, MA, USA, E6040) to manufacture the DNA library. Exome capture was performed with a SureSelect Human All Exon V4 kit (Agilent, Santa Clara, CA, USA). Paired-end sequencing was carried out using an Illumina HiSeq 2000, which generated 100-bp reads. Reference sequence data were aligned using a Burrows-Wheeler Aligner.8 Then single-nucleotide variants were analyzed using a genome analysis tool kit. The additional databases utilized included the Single-Nucleotide Polymorphism Database9 and the 1000 Genomes Project.10 As the clinical features of the affected individuals in this family are typical of an autosomal-dominant SCA, we initially filtered for heterozygous variations. We then filtered for those that were shared between the proband and the father. Protein sequence alignment was performed using the PROVEAN11 and Mutation Taster.12
Sanger sequencing
Exon 14 of the SPTBN2 gene was amplified using the following PCR primer sequences: SPTBN2 ex14-F: 5′-CTGCTCAACAAGCACACAGC-3′ and SPTBN2 ex14-R: 5′-GTTGGCAATGACTGAATCTGG-3′. Sanger sequencing was performed according to an established standard protocol for biotechnology. To determine whether or not the mutation identified in the present study is found in the general population, DNA samples from unrelated control subjects (n=90) were screened.
Results
Clinical features
The proband was a 57-year-old male who presented with poor coordination that he first noted at the age of 51. At the age of 52, he noted mild unsteadiness of gait. Neurological examination at the age of 57 revealed mild dysarthria, dysphagia and incoordination of the lower limbs. He had trouble accomplishing the heel-to-knee test and difficulty in standing with feet together; however, he was able to walk without assistance. Knee tendon reflexes were slightly exaggerated bilaterally; however, Babinski’s sign was negative. Eye movements were intact, and no nystagmus was observed. Neither mental retardation nor sensory abnormality was observed.
The clinical features of the five patients in this family are summarized in the Table 1. The ages at onset in the five patients ranged from 50s to 60s. There was no obvious anticipation of age at the onset between the two generations. Limb and trunk ataxia, and cerebellar dysarthria, were the common features in all the affected individuals. Cognitive deterioration, epilepsy, saccadic eye movements, visual disturbance, ophthalmoplegia and peripheral nerve involvement were absent in all the patients. The disease progressed very slowly and the clinical severity correlated with the duration of the disease. Two of the patients (II-6 and II-8) became wheelchair-bound ∼20 to 30 years after the disease onset. Brain MRI of our patients revealed moderate to severe atrophy in the cerebellum (Figure 2). Single-photon emission tomography (SPECT) in the proband (III-15) showed blood flow reduction in the cerebellum (Figure 3).
Mutation analysis
We identified 10 692 and 10 641 variations in the proband and the father, respectively, of which 231 and 241, respectively, were novel heterozygous variants. Of these, 183 variants were shared between the proband and the father (Supplementary Data). Once anonymous open reading frames were excluded and genes associated with SCAs were prioritized, we found one likely candidate three-nucleotide in-frame deletion mutation in the SPTBN2 gene, a gene known to cause SCA5.
On Sanger sequencing, we validated the most possible shared three-nucleotide in-frame deletion mutation in the SPTBN2 gene as the cause of the disease. This mutation cosegregated faithfully with the disease phenotype in this family (Figure 1b). All the five patients had a heterozygous c.2608_2610delGAG (p.E870del) mutation in the SPTBN2 gene, which is located at an evolutionarily conserved amino acid (Figure 4). We performed bioinformatic analyses using the PROVEAN and Mutation Taster (see ‘Materials and methods’ section), which predicted that the detected variant was deleterious and disease-causing, respectively. The variant was not detected in our 180 control chromosomes and was not observed in the databases of normal sequence variations (1000 Genome Project and Single-Nucleotide Polymorphism Database). Thus, the mutation identified was likely to be pathogenic.
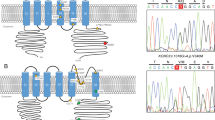
Mutations of the SPTBN2 gene. (a) Illustration of exons (filled rectangles) in the SPTBN2 gene. (b) A schematic representation of the functional domains of β-3 spectrin, modified from the reported representation,2 is shown along with the positions of all found mutations, exhibiting dominant (upper) or recessive (lower) transmission, so far. CH, calponin homology domain; ABD, actin-binding domain; ANK, ankyrin-binding domain; PH, pleckstrin homology domain. (c) Evolutionary conservation of the Japanese SCA5 mutation (p.E870del) among species.
Discussion
We presented here the first Japanese case of SCA5 with a highly likely pathogenic mutation in the SPTBN2 gene. This SCA5 family is the fifth one worldwide. The two deletions (p.E532_M544del and p.L629_R634delinsW) in the SPTBN2 gene reported earlier in American and French SCA5 families comprise distinct and nonoverlapping in-frame mutations in the third of the 17 spectrin repeats.3, 4 Both of these deletions are predicted to disrupt the triple-helical structure of the spectrin repeat and the conformation of the spectrin tetramer, which likely disrupts the protein structure involving the characteristic leucine and tryptophan residues.13 The third SCA5 family reported from Germany has a missense mutation (p. L253P) in the SPTBN2 gene that interferes with the actin-binding site in the second calponin homology domain.5 The fourth family, of Norwegian descent, has a missense mutation (c.1415C>T) that results in a p. T472M substitution in the second spectrin repeat of the beta-III spectrin protein.6
On the other hand, the p.E870del mutation detected in our family is the first one located in the sixth spectrin repeat likely associated with SCA5, resulting in the deletion of one glutamic acid deletion out of a total of 2390 amino acids. Bioinformatic analysis suggests that an amino-acid sequence changed, which might have affected protein features. Similarly, a three-nucleotide in-frame deletion mutation (c.679_681delTTC, p.F227del) in the KCND3 gene has been reported in another type of autosomal-dominant cerebellar ataxia (SCA22).14 This mutation leads to the retention of voltage-gated potassium channel Kv4.3 subunits in the cytoplasm, which is consistent with the lack of A-type K+ channel conductance on whole-cell patch-clamp recording.14 The function of beta-III spectrin, however, might be much more diverse than that of KCND3 and rather different from that of KCND3.
These mutations in the SPTBN2 spectrin repeats may affect the structure, binding properties and overall function of the protein, and also lead to SCA5. To date, no curative treatment for autosomal-dominant spinocerebellar ataxias has been found. Therefore, in addition to its value in genetic counseling, genetic knowledge of the relevant loci might help to identify the possible targets of pharmaceutical interventions.
Clinical presentations similar to those in our family have been reported in other SCA5 families.3, 4, 5, 6 The age at the onset is typically from the third to fifth decade, with highly variable symptoms that worsen over time. The major characteristics of SCA5 are slow progression with patients remaining ambulatory even after decades of suffering from the disease and having an approximately normal longevity.15 The MRI findings in four patients were pronounced cerebellar atrophy, but the brainstem remained intact, in accordance with the major clinical presentation. Our study suggests that SCA5 should be considered in a pure form of autosomal-dominant cerebellar ataxia when SCA6 or SCA31 is excluded, particularly in Japan.
A single infantile-onset SCA5 patient with a heterozygous missense mutation (c.1438C>T, p.R480W) in the SPTBN2 gene has been reported.7 The parents of the patient were healthy and nonconsanguineous. This is a much more severe phenotype with ataxia and global development delay.7 Meanwhile, a homozygous mutation in the SPTBN2 gene has been reported in a consanguineous family with childhood developmental ataxia and cognitive impairment.16, 17 Thus, recessive mutations in the SPTBN2 gene might cause a more severe disorder than SCA5, that is, resembling one infantile-onset SCA5.7 This suggests that beta-III spectrin has an important role in both development and cognition in addition to its function in the cerebellum. Cognitive impairment is an integral part of a recessive ataxic syndrome, and the condition is called spectrin-associated autosomal-recessive cerebellar ataxia type 1 (SPARCA1).13 The identification of SPARCA1 and normal heterozygous carriers of the stop codon in SPTBN2 provides insights into the mechanism underlying the molecular dominance in SCA5 and demonstrates that the cell-specific repertoire of spectrin subunits underlies a novel group of disorders, namely, the neuronal spectrinopathies, which include SCA5, SPARCA1 and a form of West syndrome.16
In summary, there are increasing numbers of SPTBN2 mutations being reported, and we added a novel SPTBN2 mutation to the ‘spectrinopathies’, which will be useful for elucidating the molecular mechanism underlying the ‘spectrinopathies’.
Accession codes
References
Dick, K. A., Ikeda, Y., Day, J. W. & Ranum, L. P. Spinocerebellar ataxia type 5. Handb. Clin. Neurol. 103, 451–459 (2012).
Ikeda, Y., Dick, K. A., Weatherspoon, M. R., Gincel, D., Armbrust, K. R., Dalton, J. C. et al. Spectrin mutations cause spinocerebellar ataxia type 5. Nat. Genet. 38, 184–190 (2006).
Ranum, L. P., Schut, L. J., Lundgren, J. K., Orr, H. T. & Livingston, D. M. Spinocerebellar ataxia type 5 in a family descended from the grandparents of President Lincoln maps to chromosome 11. Nat. Genet. 8, 280–284 (1994).
Stevanin, G., Herman, A., Brice, A. & Dürr, A. Clinical and MRI findings in spinocerebellar ataxia type 5. Neurology 53, 1355–1357 (1999).
Bürk, K., Zühlke, C., König, I. R., Ziegler, A., Schwinger, E., Globas, C. et al. Spinocerebellar ataxia type 5: clinical and molecular genetic features of a German kindred. Neurology 62, 327–329 (2004).
Cho, E. & Fogel, B. L. A family with spinocerebellar ataxia type 5 found to have a novel missense mutation within a SPTBN2 spectrin repeat. Cerebellum 12, 162–164 (2013).
Jacob, F.-D., Ho, E. S., Martinez-Ojeda, M., Darrans, B. T. & Khwaja, O. S. Case of infantile onset spinocerebellar ataxia type 5. J. Child Neurol. 28, 1292–1295 (2012).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinfomatics 38, 1767–1771 (2010).
Sherry, S. T., Ward, M. H., Kholodov, M., Baker, J., Phan, L., Smigielski, E. M. et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res. 29, 308–311 (2001).
1000 Genomes Project Consortium. A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010).
Choi, Y. A fast computation of pairwise sequence alignment scores between a protein and a set of single-locus variants of another protein In Proceedings of the ACM Conference on Bioinformatics, Computational Biology and Biomedicine (BCB ’12) 414–417 (ACM: New York, NY, USA, 2012).
Schwarz, J. M., Rödelsperger, C., Schuelke, M. & Seelow, D. MutationTaster evaluates disease-causing potential of sequence alterations. Nat. Methods 7, 575–576 (2010).
Ylanne, J., Scheffzek, K., Young, P. & Saraste, M. Crystal structure of the alpha-actinin rod reveals an extensive torsional twist. Structure 9, 597–604 (2001).
Lee, Y.-C., Durr, A., Majczenko, K., Huang, Y.-H., Liu, Y.-C., Lien, C.-C. et al. Mutations in KCND3 cause spinocerebellar ataxia type 22. Ann. Neurol. 72, 859–869 (2012).
Bauer, P., Schöls, L. & Riess, O. Spectrin mutations in spinocerebellar ataxia (SCA). Bioessays 28, 785–787 (2006).
Lise, S., Clarkson, Y., Perkins, E., Kwasniewska, A., Sadighi Akha, E., Schnekenberg, R. P. et al. Recessive mutations in SPTBN2 implicate b-III spectrin in both cognitive and motor development. PLoS Genet. 8, e1003074 (2012).
Elsayed, S. M., Heller, R., Thoenes, M., Zaki, M. S., Swan, D., Elsobky, E. et al. Autosomal dominant SCA5 and autosomal recessive infantile SCA are allelic conditions resulting from SPTBN2 mutations. Eur. J. Hum. Genet. 22, 286–288 (2014).
Acknowledgements
We thank the family for participating in this study. We also thank Drs Hayashi and Akiyama, Akiyama Neurosurgery Clinic, for referring the patients to us, and Drs Honda and Shimazaki, Department of Neurology, Jichi Medical University, and Dr Kinya Ishikawa, Department of Neurology and Neurological Science, Graduate School, Tokyo Medical and Dental University, for the molecular testing of the proband for SCA1, SCA2, MJD, SCA6, SCA7, SCA8, SCA12, SCA17, SCA31 and DRPLA. This work was supported by a grant from the Research Committee for Ataxic Diseases (YT) of the Ministry of Health, Labor and Welfare, Japan.
Accession codes The nucleotide sequence data reported are available in the GenBank database under the accession number KJ567653.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, Y., Koh, K., Miwa, M. et al. A Japanese SCA5 family with a novel three-nucleotide in-frame deletion mutation in the SPTBN2 gene: a clinical and genetic study. J Hum Genet 59, 569–573 (2014). https://doi.org/10.1038/jhg.2014.74
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2014.74
This article is cited by
-
Two novel missense variants in SPTBN2 likely associated with spinocerebellar ataxia type 5
Neurological Sciences (2021)
-
Between SCA5 and SCAR14: delineation of the SPTBN2 p.R480W-associated phenotype
European Journal of Human Genetics (2018)
-
A Novel Homozygous Mutation in SPTBN2 Leads to Spinocerebellar Ataxia in a Consanguineous Family: Report of a New Infantile-Onset Case and Brief Review of the Literature
The Cerebellum (2018)
-
A human β-III-spectrin spinocerebellar ataxia type 5 mutation causes high-affinity F-actin binding
Scientific Reports (2016)