Abstract
Rhizomelic chondrodysplasia punctata (RCDP) is an autosomal recessive disorder due to the deficiency in ether lipid synthesis. RCDP type 1, the most prominent type, is caused by the dysfunction of the receptor of peroxisome targeting signal type 2, Pex7 (peroxisomal biogenesis factor 7), and the rest of the patients, RCDP types 2 and 3, have defects in peroxisomal enzymes catalyzing the initial two steps of alkyl-phospholipid synthesis, glyceronephosphate O-acyltransferase and alkylglycerone phosphate synthase (Agps). We herein investigated defects of two patients with RCDP type 3. Patient 1 had a novel missense mutation, T1533G, resulting in the I511M substitution in Agps. The plasmalogen level was mildly reduced, whereas the protein level and peroxisomal localization of Agps-I511M in fibroblasts were normal as in the control fibroblasts. Structure prediction analysis suggested that the mutated residue was located in the helix α15 on the surface of V-shaped active site tunnel in Agps, likely accounting for the mild defects of plasmalogen synthesis. These results strongly suggest that an individual with mildly affected level of plasmalogen synthesis develops RCDP. In fibroblasts from patient 2, the expression of AGPS mRNA and Agps protein was severely affected, thereby giving rise to the strong reduction of plasmalogen synthesis.
Similar content being viewed by others
Introduction
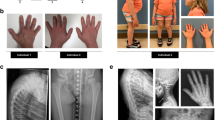
Rhizomelic chondrodysplasia punctata (RCDP) is a rare, peroxisome-related disorder deficient in the synthesis of ether phospholipids such as plasmalogens. Patients with RCDP have clinical features of shortened proximal long bones, rhizomelia, bilateral congenital cataracts, dysmorphic facial features and severe growth and developmental delays. Radiographic examination reveals punctate epiphyseal calcifications, metaphyseal dysplasia and vertebral coronal clefts.1
More than 90% of RCDP patients have defects in peroxisomal biogenesis factor 7 gene (PEX7), referred as RCDP type 1 (RCDP1). Pex7 encoded by PEX7 is the receptor of peroxisome targeting signal type 2-containing enzymes such as 3-ketoacyl-CoA thiolase catalyzing the final step of peroxisomal β-oxidation of fatty acids. Less than 10% of RCDP patients are deficient in two genes encoding two peroxisomal matrix enzymes required for the synthesis of plasmalogens and other alkyl-phospholipids. These are glyceronephosphate O-acyltransferase (Gnpat) and alkylglycerone phosphate synthase (Agps), of which defects are responsible for RCDP types 2 and 3 (RCDP2 and RCDP3), respectively.2
Plasmalogens are a major subclass of choline and ethanolamine glycerophospholipids containing a vinyl-ether-linked long-chain fatty alcohol at the sn-1 position of the glycerol backbone. The initial two steps in plasmalogen synthesis occur inside peroxisomes and the later steps proceed in the endoplasmic reticulum.3 First, Gnpat catalyzes the esterification of dihydroxyacetone phosphate (DHAP) to generate acyl-DHAP. Subsequently, Agps replaces the acyl chain of acyl-DHAP with a long-chain fatty alcohol, generating alkyl-DHAP. The Agps reaction follows a ‘ping-pong’ mechanism, where the fatty acid product is released from acyl-DHAP before the binding of the fatty alcohol.4 The crystal structure of Agps5, 6 shows a hydrophobic V-shaped tunnel matching the 16- to 18-carbon ligand, a gating helix that functions in substrate binding and product release and a catalytic center that forms a flavin-linked intermediate with DHAP.
Although the functional roles of plasmalogens remain largely unknown, several potential functions have been proposed. Their specialized functions include a role in the protection against oxidants,7, 8 transport of LDL-derived cholesterol from cell surface and endocytic compartments to endoplasmic reticulum,9 inflammatory responses10 and composition of lipid raft membranes.11
Several mutations in the AGPS gene have been identified so far in RCDP3 patients.12, 13, 14 In this study, we identified a novel missense mutation in AGPS gene in an Iranian female child, patient 1, who was clinically diagnosed as RCDP. In addition, we also reported a Lebanese RCDP patient, patient 2, with no detectable level of AGPS mRNA.
Materials and methods
Cell culture
Skin fibroblast cell lines were cultured in Dulbecco’s modified Eagle’s medium (high glucose) (Invitrogen, Carlsbad, CA, USA) supplemented with 10% heat-inactivated fetal bovine serum (Sigma, St Louis, MO, USA) in 5% CO2–95% air.15
Morphological analysis
Cells were fixed with 4% paraformaldehyde and permeabilized with 0.1% Triton X-100 at room temperature for 30 min.16 Peroxisomes were visualized by immunostaining with antibodies to Pex1417 and Agps.18 Antigen and antibody complex was detected with Alexa Fluor 488- or 568-labeled secondary antibodies (Invitrogen) as described.16 Cells were observed by an LSM 510 confocal microscope (Carl Zeiss, Oberkochen, Germany). Images were acquired and analyzed with an LSM image browser (Carl Zeiss).
Mutation analysis
AGPS cDNA was generated by reverse transcription-polymerase chain reaction (RT–PCR) method using total RNA extracted from fibroblasts with Trizol (Invitrogen). RT reaction was performed with SuperScript II reverse transcriptase (Life Technologies, Carlsbad, CA, USA) and 18-mer oligo-dT primer (Amersham Biosciences, Piscataway, NJ, USA). PCR was carried out with a pair of primers, hsAGPS5′ (5′-GGGTAAGCTTGGTTCCGGGCGGCAGCACAAGGCGGTAGCC-3′) and hsAGPS3′ (5′-GCGTGCGGCCGCTGACATTTTTTTGTAATGGTACTAATGGAT-3′). The PCR product was digested with HindIII and NotI and cloned into the HindIII and NotI sites of pcDNA3.1Zeo (Invitrogen). The nucleotide sequence was determined by the dideoxynucleotide chain termination method using a Dye-terminator DNA sequence kit (Applied Biosystems, Foster City, CA, USA). Sequences were aligned by using GENETYX-MAC program (Software Development Co., Ltd., Tokyo, Japan). Genomes were extracted from GM16678 with Trizol and regions containing primer annealing sites used for RT–PCR were amplified using sets of primers AGPS Met genome Fw (5′-CCCAGGGTTCTAGGACACCTCGAGG-3′) and AGPS Met genome Rv (5′-CGCTTCTTTGGGATGGTGCCCGACTC-3′), and AGPS Stop Fw (5′-GGGCTTAATGAGCTTGATAGTCTGG-3′) and AGPS Stop Rv (5′-AGGCCAAGCACCTTACAGACGTGAAGGC-3′), and directly sequenced using AGPS Met genome Fw and AGPS Stop Fw, respectively.
Lipid analysis
Cells were metabolically labeled for the indicated time with 14C-ethanolamine (Amersham Biosciences)19 and subjected to the analysis of plasmalogens as described.18
Liquid chromatography-electronspray ionization-tandem mass spectrometry analysis
Total lipids were extracted by Bligh and Dyer method.20 Cells were dissolved in methanol/chloroform/water (v/v/v: 2:1:0.8) and 50 pmol of 1-heptadecanoyl-sn-glycero-3-phosphocholine, 1, 2-didodecanoyl-sn-glycero-3-phosphocholine and 1, 2-didodecanoyl-sn-glycero-3-phosphoethanolamine (DDPE) were added as internal standards. After the incubation for 5 min at room temperature, 1 ml each of water and chloroform was added and the extracts were then centrifuged to collect a lower organic phase. To re-extract the lipids from water phase, 1 ml chloroform was added. The combined organic phase was evaporated under the nitrogen stream and suspended in methanol.
Liquid chromatography-electronspray ionization-tandem mass spectrometry was performed using a 4000 Q-TRAP quadrupole linear ion trap hybrid mass spectrometer (AB Sciex, Framingham, MA, USA) with ACQUITY UPLC System (Waters, Milford, MA, USA). Samples were injected to an ACQUIRY UPLC BEH C18 column (1.7 μm, 1.0 × 150 mm2; Waters) and then directly subjected to ESI-MS/MS analysis. A 10-μl aliquot of each sample was directly introduced by autosampler injector and the samples were separated by step gradient elution with mobile phase A (acetonitrile:methanol:water at 2:2:1 (v/v/v), 0.1% formic acid and 0.028% ammonium) and mobile phase B (isopropanol, 0.1% formic acid and 0.028% ammonium) at the ratios: 100:0 (for 0–5 min), 95:5 (5–20 min), 70:30 (20–21 min), 50:50 (21–45 min), 50:50 (45–100 min) and 100:0 (100–120 min). The flow rate was 70 μl min−1 at 30 °C. Source temperature was at 200 °C, declustering potential was at 60 and collision cell exit potential was at 15. Plasmenylethanolamine (PlsEtns) with alkenyl p16:0, p18:0 and p18:1 at sn-1 position were detected by precursor ion scan of m/z 364, m/z 392 and m/z 390, respectively, at positive ion mode. The data were analyzed and quantified using the Analyst software (AB Sciex).
Antibodies
We used rabbit antisera against Agps18 and Pex14.17 A mouse monoclonal antibody against α-tubulin was purchased from Abcam (Cambridge, UK).
Protein modeling
Automated homology modeling was performed in SWISS-MODEL by aligning Homo sapiens Agps to the crystal structure of Cavia porcellus (PDB 4BBY). The root-mean-square deviation of the Cα atomic coordinates of the superimposed model structure on the reference molecule was 0.07 Å, indicating a good fit. We used PyMOL for the preparation of figure and calculation of root-mean-square deviation. Note that the direction of the side chain at M511 can flexibly change.
Results
Clinical report
Patient 1 was an Iranian female child who was born to a cousin couple, birth weight 2.6 kg. There was no similar case in the family. The predominant features were intrauterine and postnatal growth retardation, motor delay, bilateral congenital cataracts, bilateral congenital diaphragmatic hernia, rhizomelia, joint stiffness and contracture and stippled epiphyses. Patient 1 is alive and she was 5 years 3 months old in December 2013. She could sit without aid, but could only stand with aid. She could walk independently but there was some limitation of large joint movement. She was operated for cataract, and she did not have problem either with vision or hearing. She could eat and swallow foods without any difficulty. She could talk a few words and make simple sentences. Her weight was 12.5 kg, height was 89 cm and head circumference was 46 cm, all below 3% for age.
Coriell Institute reported that patient 2 was a Lebanese female child. The cardinal features were cataracts, rhizomelic shortening of the limbs and stippled epiphyses. These two patients were clinically diagnosed as RCDP by their symptoms. Fibroblasts derived from the patient 1 were established and named R1 and fibroblasts named GM16678 from patient 2 were purchased from Coriell Institute for Medical Research (Camden, NJ, USA).
Agps deficiency in two RCDP patients
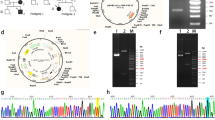
To determine the primary defect of the two RCDP patients, expression levels of PEX7, GNPAT and AGPS mRNA were assessed by RT–PCR using total RNA from R1 and GM16678 and specific primers for respective genes. A single PCR product of about 2.0 kbp apparently corresponding to the entire open reading frame of AGPS was similarly amplified from both control and R1 fibroblasts (Figure 1A, lanes 1 and 2). By contrast, no band was detected in the RT–PCR product from GM16678 (Figure 1A, lane 4). Analysis of the genomic sequence did not show any mutation in the primer annealing sites used for RT–PCR in GM16678. These results suggest that AGPS transcription was defective in GM16678. RT–PCR products of PEX7 and GNPAT were amplified with normal size and level in the fibroblasts from a control, R1 and GM16678 (data not shown). Subsequent sequencing of six independent AGPS cDNA clones from R1 indicated a point mutation of a thymine at nucleotide position 1533 to a guanine, resulting in an amino-acid change from Ile to Met at 511 in Agps, named Agps-I511M. No mutation was identified in cDNAs of PEX 7 and GNPAT from R1 from two and six independent clones, respectively (data not shown). Therefore, two RCDP patients analyzed here are defective in Agps and genetically classified as RCDP3.
Defects of alkylglycerone phosphate synthase (Agps) in two rhizomelic chondrodysplasia punctata (RCDP) patients. (A) Expression levels of AGPS in R1 and GM16678. AGPS mRNA in fibroblasts from a control (control, lanes 1 and 3), R1 (lane 2) and GM16678 (lane 4) was verified by reverse transcription-polymerase chain reaction (RT–PCR) with a pair of primers specific for AGPS (upper panels) or actin (lower panels). (B) Mutation analysis of AGPS in R1. Partial nucleotide sequences of AGPS cDNA isolated from fibroblasts from R1 (R1, left panel) and a control (Control, right panel) and the deduced amino acids are shown. A thymine 1533 in fibroblasts from a control (shaded, right panel) was substituted by a guanine in R1 (shaded, left panel), resulting in the Ile511 codon (ATT) in fibroblasts from a control mutated to a Met511 codon (ATG) in R1. (C) Expression of Agps protein in R1 and GM16678. Fibroblasts from a control (lane 1), R1 (lane 2), and GM16678 (lane 3) were lysed and analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and immunoblotting with antibodies against Agps (upper panel) and α-tubulin as a loading control (lower panel). (D) Intracellular localization of Agps in R1. Fibroblasts from a control (a–c), R1 (d–f) and GM16678 (g–i) were immunostained with antibodies against Agps (a, d and g) and peroxisomal biogenesis factor 14 (Pex14) (b, e and h). Merged views are also shown. Bar, 10 μm.
Protein analysis
Next, we examined the expression level of Agps in R1 and GM16678. Whole-cell lysates were analyzed by immunoblotting with anti-Agps antibody.18 Agps was hardly detectable in GM16678 (Figure 1C, lane 3), consistent with the undetectable level of AGPS mRNA (Figure 1A, lane 4). By contrast, the expression level of Agps in R1 was similar to that in control fibroblasts (Figure 1C, lanes 1 and 2), suggesting that the I511M mutation did not alter the stability of Agps. Furthermore, Agps-I511M was coincided with Pex14, indicative of peroxisomal localization (Figure 1Dd–f), as assessed with wild-type Agps in control fibroblasts (Figure 1Da–c). No fluorescence signal of Agps was detected in GM16678-derived fibroblasts (Figure 1Dg–i). These results suggested that I511M mutation in Agps did not affect the efficiency of its peroxisomal import.
Agps protein modeling
To appreciate the potential effects of Agps point mutation on the activity, we predicted the three-dimensional structure of the mutated H. sapiens Agps using SWISS-MODEL, based on the C. porcellus Agps crystal structure.6 In the amino-acid sequence, H. sapiens Agps shares 92% identity with the C. porcellus (guinea-pig) Agps. The model was visualized with PyMOL to predict the structural changes (Figure 2). The location of a substrate, long-chain fatty alcohol, in H. sapiens Agps was estimated by alignment to Dictyostelium discoideum Agps bound to long-chain fatty alcohol,5 based on their secondary structure elements using secondary structure matching implemented in COOT.21 Similar to the structure of C. porcellus Agps, H. sapiens Agps has a V-shaped tunnel that runs across the cap–flavin adenine dinucleotide domain interface from the gating helix. The I511 was located in the helix α15 along with the V-shaped tunnel and its side chain was predicted to be adjacent to the substrate long-chain fatty alcohol (Figure 2b). On the other hand, a longer bulky side chain of the mutated M511 could penetrate into the V-shaped tunnel (Figure 2a). Thus, it is more likely that the I511M substitution lowers the substrate accessibility to the narrow hydrophobic tunnel by steric hindrance.
Protein modeling of alkylglycerone phosphate synthase (Agps)-I511M mutant. Three-dimensional structures of the wild-type and mutated form of Homo sapiens Agps are modeled based on the crystal structure of Cavia porcellus Agps.6 Close-up views of the hydrophobic active-site tunnel containing the fatty alcohol (C16) substrate (fatty alcohol (FA)) (green chain, open arrowhead) and flavin adenine dinucleotide (FAD) cofactor (green isoalloxazine ring, solid arrowhead) in Agps-I511M (a) and wild-type Agps (b) were presented.
Plasmalogen level in RCDP3 patients
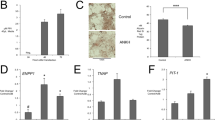
We determined plasmalogen levels in R1 and GM16678. Phosphatidylethanolamine (PE) and PlsEtn are detectable in two distinct bands on one-dimensional thin layer chromatography plate after the trichloroacetic acid treatment, because trichloroacetic acid hydrolyzes alkenyl ether bond of PlsEtn, resulting in the conversion of PlsEtn to 2-acylglycerophosphoethanolamine (lyso-PE).18 Upon trichloroacetic acid treatment, lyso-PE was evident in the case of control fibroblasts, whereas a severely reduced level of lyso-PE was detected in GM16678 cells. On the other hand, lyso-PE was mildly reduced in R1 as compared with that in control fibroblasts (Figure 3a). Furthermore, pulse-labeling experiment with 14C-ethanolamine revealed that newly synthesized PlsEtn was reduced severely in GM16678, but that only mildly reduced in R1 (Figures 3b and c).
Levels of plasmenylethanolamine in R1 and GM16678. (a) Total lipids of R1 and GM16678. Lipids extracted from fibroblasts from a control, R1 and GM16678 were resolved by thin layer chromatography and detected by exposure to iodine vapor. PC, phosphatidylcholine; PI, phosphatidylinositol; PS, phosphatidylserine; SM, sphingomyelin. Note that a plasmalogen, plasmenylethanolamine (PlsEtn), was detected as 2-acylglycerophosphoethanolamine (lyso-PE).18 (b) PlsEtn synthesis in R1 and GM16678. Cells from a normal control, R1 and GM16678 were metabolically labeled with 14C-ethanolamine for the indicated time. Plasmalogen synthesis was analyzed as described in (a) and detected by autoradiography. (c) Lyso-PE was quantified using image analysis software (Multi Gauge; Fuji Photo Film, Tokyo, Japan). Relative amount of newly synthesized plasmalogens in fibroblasts from a control (solid circle), R1 (solid square) and GM16678 (solid triangle) are shown, where the levels in control fibroblasts were designated as 1. Data represent the means±s.d. of four independent experiments. (d) Liquid chromatography-electronspray ionization-tandem mass spectrometry (LC-ESI-MS/MS) analysis of plasmalogens in R1 and GM16678. Lipids were extracted as in (a) without trichloroacetic acid (TCA) treatment and then analyzed by LC-ESI-MS/MS. PlsEtns with alkenyl p16:0, p18:0 and p18:1 at sn-1 position were detected by precursor ion scan of m/z 364, m/z 392 and m/z 390, respectively, at the positive ion mode. Concentrations of phospholipids were calculated using protein concentrations and internal standards. Data represent the means±s.d. of three independent experiments (*P<0.001; t-test versus control).
The sn-1 position of PlsEtn is mostly linked to C16:0, C18:0 and C18:1 fatty alcohols. I511M mutation is positioned at the substrate-binding site tunnel in the Agps molecule. Thus, we suspected that any of three long-chain fatty alcohol substrates might not fit to the tunnel of Agps-I511M. To test this hypothesis, we verified the composition of alkenyl chains in PlsEtns in R1 by a liquid chromatography-electronspray inonization-tandem mass spectrometry analyzer. However, all the alkenyl chain species of PlsEtns were equally reduced in R1 (Figure 3d), implying that the side chain at I511M mutation on the surface of V-shaped tunnel of Agps less likely affects the tunnel in the binding affinity toward any particular type of fatty alcohols.
Discussion
A wide spectrum of mutations, including missense and splicing site mutations, has been reported in AGPS gene in RCDP3 patients.12, 13, 14 We show here two new probands with RCDP3.
We identified a novel one point mutation of AGPS in patient 1. The predicted structure of H. sapiens Agps indicated that the mutated residue I511M identified in R1 is located in the helix α15 in the V-shaped tunnel (Figure 2a). The mutation did not affect the protein stability, peroxisomal localization and specificity of fatty alcohols required for the formation of ether bond (Figure 3d). However, the amounts of PlsEtn species were mildly decreased (Figures 3a–c). The mutation likely altered the packing fraction of the space of V-shaped tunnel and mildly lowered the accessibility of long-chain fatty alcohols or acyl-DHAP to the tunnel.
It is of interest to note that the T568M mutation was identified in an RCDP3 patient.12 T568 is also located within the active site tunnel, close to both the fatty alcohol and flavin adenine dinucleotide cofactor. Despite no predicted change in polar interactions, the substituted methionine residue could cause steric hindrance of the V-shaped tunnel, thereby giving rise to rather severe reduction of plasmalogen synthesis despite normal expression of Agps protein.12
The clinical severity of patients with Agps-T568M and Agps-I511M is high and at the same level, but the PlsEtn synthesis in cells expressing Agps-I511M was only mildly affected. Studies from several RCDP patients proposed that the severity in RCDP patients is predicted by the capacity of plasmalogen synthesis.12 Given these data, we suspected that plasmalogen level in the patient with Agps-I511M is just above the threshold for severe RCDP symptom. Other mutations identified in RCDP3 patients mostly destabilize Agps by loosing structural integrity, thereby giving rise to the strong reduction of the Agps expression.12
Patient 2 was completely compromised in the expression of AGPS mRNA and severely affected in the plasmalogen synthesis. G-banded chromosome analysis showed no large deletion of the long arm of chromosome 2 where AGPS is located (data not shown), suggesting that alternation at a transcriptional level or rapid degradation of AGPS mRNA causes severe reduction of Agps expression in GM16678. In the two patients, posttranslational modification of proteins with glycosylphosphatidylinositol (GPI) may also be affected, because the majority of protein-linked GPI contains 1-alkyl-2-acyl phosphatidylinositol22 and the biosynthesis of alkyl-containing GPI requires peroxisomal alkyl-phospholipid biosynthetic pathway.23 Recently, Kanzawa et al.24 reported that lipid remodeling of GPI-anchored proteins was impaired in fibroblasts from patients with RCDP2 and RCDP3, where only diacyl form of GPI was generated. In addition to the deficiency of plasmalogen synthesis, unusual GPI-anchored proteins may contribute to the complex phenotypes of the RCDP3 patients.
Change history
25 July 2014
This article has been corrected since Advance Online Publication, and an erratum is also printed in this issue.
References
Braverman, N. E. & Moser, A. B. Functions of plasmalogen lipids in health and disease. Biochim. Biophys. Acta 1822, 1442–1452 (2012).
Nimmo, G., Monsonego, S., Descartes, M., Franklin, J., Steinberg, S. & Braverman, N. Rhizomelic chondrodysplasia punctata type 2 resulting from paternal isodisomy of chromosome 1. Am. J. Med. Genet. 152, 1812–1817 (2010).
Nagan, N. & Zoeller, R. A. Plasmalogens: biosynthesis and functions. Prog. Lipid Res. 40, 199–229 (2001).
Brown, A. J. & Snyder, F. Alkyldihydroxyacetone-P synthase. Solubilization, partial purification, new assay method, and evidence for a ping-pong mechanism. J. Biol. Chem. 257, 8835–8839 (1982).
Razeto, A., Mattiroli, F., Carpanelli, E., Aliverti, A., Pandini, V., Coda, A. et al. The crucial step in ether phospholipid biosynthesis: structural basis of noncanonical reaction associated with a peroxisomal disorder. Structure 15, 683–692 (2007).
Nenci, S., Piano, V., Rosati, S., Aliverti, A., Pandini, V., Fraaije, M. W. et al. Precursor of ether phospholipids is synthesized by a flavoenzyme through covalent catalysis. Proc. Natl Acad. Sci. USA 109, 18791–18796 (2012).
Zoeller, R. A., Morand, O. H. & Raetz, C. R. H. A possible role for plasmalogens in protecting animal cells against photosensitized killing. J. Biol. Chem. 263, 11590–11596 (1988).
Tsukamoto, T., Miura, S. & Fujiki, Y. Restoration by a 35K membrane protein of peroxisome assembly in a peroxisome-deficient mammalian cell mutant. Nature 350, 77–81 (1991).
Munn, N. J., Arnio, E., Liu, D., Zoeller, R. A. & Liscum, L. Deficiency in ethanolamine plasmalogen leads to altered cholesterol transport. J. Lipid Res. 44, 182–192 (2003).
Brites, P., Waterham, H. R. & Wanders, R. J. A. Functions and biosynthesis of plasmalogens in health and disease. Biochim. Biophys. Acta 1636, 219–231 (2004).
Rodemer, C., Thai, T. P., Brugger, B., Kaercher, T., Werner, H., Nave, K. A. et al. Inactivation of ether lipid biosynthesis causes male infertility, defects in eye development and optic nerve hypoplasia in mice. Hum. Mol. Genet. 12, 1881–1895 (2003).
Itzkovitz, B., Jiralerspong, S., Nimmo, G., Loscalzo, M., Horovitz, D. D. G., Snowden, A. et al. Functional characterization of novel mutations in GNPAT and AGPS, causing rhizomelic chondrodysplasia punctate (RCDP) types 2 and 3. Hum. Mutat. 33, 189–197 (2012).
Thai, T. P., Rodemer, C., Jauch, A., Hunziker, A., Moser, A., Gorgas, K. et al. Impaired membrane traffic in defective ether lipid biosynthesis. Hum. Mol. Genet. 10, 127–136 (2001).
de Vet, E.C.J.M., Ijlst, L., Oostheim, W., Dekker, C., Moser, H. W., van Den Bosch, H. et al. Ether lipid biosynthesis: alkyl-dihydroxyacetonephosphate synthase protein deficiency leads to reduced dihydroxyacetonephosphate acyltransferase activities. J. Lipid Res. 40, 1998–2003 (1999).
Fukuda, S., Shimozawa, N., Suzuki, Y., Zhang, Z., Tomatsu, S., Tsukamoto, T. et al. Human peroxisome assembly factor-2 (PAF-2): a gene responsible for group C peroxisome biogenesis disorder in humans. Am. J. Hum. Genet. 59, 1210–1220 (1996).
Okumoto, K., Shimozawa, N., Kawai, A., Tamura, S., Tsukamoto, T., Osumi, T. et al. PEX12, the pathogenic gene of group III Zellweger syndrome: cDNA cloning by functional complementation on a CHO cell mutant, patient analysis, and characterization of Pex12p. Mol. Cell. Biol. 18, 4324–4336 (1998).
Shimizu, N., Itoh, R., Hirono, Y., Otera, H., Ghaedi, K., Tateishi, K. et al. The peroxin Pex14p: cDNA cloning by functional complementation on a Chinese hamster ovary cell mutant, characterization, and functional analysis. J. Biol. Chem. 274, 12593–12604 (1999).
Honsho, M., Yagita, Y., Kinoshita, N. & Fujiki, Y. Isolation and characterization of mutant animal cell line defective in alkyl-dihydroxyacetonephosphate synthase: localization and transport of plasmalogens to post-Golgi compartments. Biochim. Biophys. Acta 1783, 1857–1865 (2008).
Nagan, N., Hajra, A. K., Larkins, L. K., Lazarow, P., Purdue, P. E., Rizzo, W. B. et al. Isolation of a Chinese hamster fibroblast variant defective in dihydroxyacetonephosphate acyltransferase activity and plasmalogen biosynthesis: use of a novel two-step selection protocol. Biochem. J. 332, 273–279 (1998).
Bligh, E. G. & Dyer, W. J. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37, 911–917 (1959).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D 60, 2126–2132 (2004).
Chatterjee, S. & Mayor, S. The GPI-anchor and protein sorting. Cell. Mol. Life Sci. 58, 1969–1987 (2001).
Kanzawa, N., Maeda, Y., Ogiso, H., Murakami, Y., Taguchi, R. & Kinoshita, T. Peroxisome dependency of alkyl-containing GPI-anchor biosynthesis in the endoplasmic reticulum. Proc. Natl Acad. Sci. USA 106, 17711–17716 (2009).
Kanzawa, N., Shimozawa, N., Wanders, R. J. A., Ikeda, K., Murakami, Y., Waterham, H. R. et al. Defective lipid remodeling of GPI anchors in peroxisomal disorders, Zellweger syndrome, and rhizomelic chondrodysplasia punctata. J. Lipid Res. 53, 653–663 (2012).
Acknowledgements
We thank K Shimizu for the figure illustrations, Y Nanri for technical assistance and the other members of our laboratory for discussions. This work was supported in part by a CREST grant (to YF) from the Science and Technology Agency of Japan; Grants-in-Aid for Scientific Research (numbers 19058011, 20370039, 24247038 and 25116717 to YF; 23570236 to MH); the Global COE Program and Grants for Excellent Graduate Schools from the Ministry of Education, Culture, Sports, Science and Technology of Japan; and grants from the Japan Foundation for Applied Enzymology and Takeda Science Foundation. MN was a Research Fellow of the Japan Society for the Promotion of Science.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Noguchi, M., Honsho, M., Abe, Y. et al. Mild reduction of plasmalogens causes rhizomelic chondrodysplasia punctata: functional characterization of a novel mutation. J Hum Genet 59, 387–392 (2014). https://doi.org/10.1038/jhg.2014.39
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2014.39