Abstract
Immunodeficiency, centromeric instability and facial anomalies (ICF) syndrome is a rare autosomal recessive disorder that shows DNA hypomethylation at pericentromeric satellite-2 and -3 repeats in chromosomes 1, 9 and 16. ICF syndrome is classified into two groups: type 1 (ICF1) patients have mutations in the DNMT3B gene and about half of type 2 (ICF2) patients have mutations in the ZBTB24 gene. Besides satellite-2 and -3 repeats, α-satellite repeats are also hypomethylated in ICF2. In this study, we report three novel ZBTB24 mutations in ICF2. A Japanese patient was homozygous for a missense mutation (C383Y), and a Cape Verdean patient was compound heterozygous for a nonsense mutation (K263X) and a frame-shift mutation (C327W fsX54). In addition, the second Japanese patient was homozygous for a previously reported nonsense mutation (R320X). The C383Y mutation abolished a C2H2 motif in one of the eight zinc-finger domains, and the other three mutations caused a complete or large loss of the zinc-finger domains. Our immunofluorescence analysis revealed that mouse Zbtb24 proteins possessing a mutation corresponding to either C383Y or R320X are mislocalized from pericentrometic heterochromatin, suggesting the importance of the zinc-finger domains in proper intranuclear localization of this protein. We further revealed that the proper localization of wild-type Zbtb24 protein does not require DNA methylation.
Similar content being viewed by others
Introduction
Immunodeficiency, centromeric instability and facial anomalies (ICF) syndrome (MIM 242860) is an extremely rare autosomal recessive disorder accompanied by recurrent infectious diseases. The immunodeficiency phenotype is characterized by reduction in the number of circulating memory B cells and in various types of serum immunoglobulin.1, 2 The centromeric instability is manifested by undercondensation of the pericentromeric heterochromatin of chromosomes 1, 9 and 16, and multiradial chromosome configurations involving these regions.3 The cytological defects are associated with DNA hypomethylation of satellite-2 and -3 repeats at the pericentromeric heterochromatin regions.4, 5, 6, 7, 8, 9, 10, 11 Facial anomalies distinctive for ICF syndrome patients include hypertelorism, low-set ears, epicanthic folds and macroglossia.
About half of ICF syndrome patients are classified as type 1 (ICF1), which shows DNA hypomethylation of only satellite-2 and -3 repeats, and the other half are classified as type 2 (ICF2), which shows hypomethylation of α-satellite repeats, in addition to satellite-2 and -3 repeats at the centromeric regions of all chromosomes.8 ICF1 is caused by mutations in the DNMT3B gene, which encodes one of the two de novo DNA methyltransferases.9, 10, 11 On the other hand, although about half of ICF2 patients possess mutations in the ZBTB24 gene,12, 13 the gene(s) responsible for the other half of ICF2 patients is still unknown. The ZBTB24 protein has a BR-C, ttk and bab (BTB) domain, and an AT-hook domain in its N-terminal region and eight C2H2-type zinc-finger domains in its C-terminal region, but the molecular function of this protein remains elusive. The other members of the BTB–zinc-finger domain family include Kaiso/ZBTB33, which recognizes methylated DNA by its zinc-finger domains14 and the B-cell lymphoma 6 transcription factor (also called ZBTB27), which is required for differentiation of naive B cells into memory B cells in germinal centers.15 Thus, ZBTB24 might function in regulation of DNA methylation and/or transcription.
In this study, we identified three novel ZBTB24 mutations, all of which affected the zinc-finger domains of the protein, in Japanese and Cape Verdean ICF2 patients. Moreover, we show that mutations corresponding to C383Y and R320X of ICF2 patients cause mislocalization of the mouse Zbtb24 protein fused with enhanced green fluorescent protein (EGFP) in the nucleus, thus providing evidence for the importance of the zinc-finger domains in heterochromatin targeting.
Materials and methods
Patient samples
Three ICF2 patients were studied. Two of them were Japanese males (P6 and P7) and the last one was a Cape Verdean female (P8). Research protocols and consent forms were reviewed and approved by the ethical committee of each University/Medical Center/Hospital. Genomic DNA was isolated from peripheral blood leukocytes of the patients by a standard method.
Southern blotting
Genomic DNA of 500 ng was digested with BstBI (New England Biolabs, Beverly, MA, USA) for 12 h at 65 °C (to study satellite-2) or with HhaI (Nippon Gene, Tokyo, Japan) for 12 h at 37 °C (to study α-satellite). The two restriction enzymes are both sensitive to cytosine methylation at their recognition sites. The digested DNA fragments were separated by electrophoresis using 1% agarose gel and transferred to Hybond XL nylon membranes (GE Healthcare, Little Chalfont, UK) in 0.4 N NaOH. The membranes were hybridized with either a 32P-labeled satellite-2 probe (5′-TCGAGTCCATTCGATGAT-3′) (for the BstBI digests) or a 32P-labeled α-satellite probe (5′-ATGTGTGCATTCAACTCACAGAGTTGAAC-3′)16 (for the HhaI digests). Hybridization was carried out for 24 h at 42 °C in 6 × saline sodium citrate, 5 × Denhardt’s solution and 0.1% SDS. After hybridization, the membranes were washed in 6 × saline sodium citrate and 0.1% SDS three times at room temperature, followed by autoradiography.
Mutation search
Mutations were identified in ICF2 patients by sequencing the PCR products together covering all the exons and exon/intron boundaries of the DNMT3B and ZBTB24 genes. The PCR primer sequences are listed in Supplementary Table 1. PCR was performed using KOD–plus (TOYOBO, Osaka, Japan) under the following conditions: 35 cycles of 98 °C for 10 s, 60 °C for 30 s and 68 °C for 30 s. The PCR products were purified using QIAquick Gel Extraction Kit (Qiagen, Hilden, Germany). The sequencing reaction was performed using BigDye Terminator v3.1 Cycle Sequencing kit (Applied Biosystems, Foster City, CA, USA) with either of the two PCR primers and analyzed on 3130xl Genetic Analyzer (Applied Biosystems).
Plasmids and cell lines
Mouse Zbtb24 and Dnmt3b cDNAs containing the entire coding regions were cloned into pEGFP C1 plasmid vector (Clontech, Mountain View, CA, USA). Mutations corresponding to those found in two Japanese ICF2 patients were introduced into the mouse Zbtb24 cDNA using PCR primers carrying the mutations. NIH3T3 mouse fibroblast cells were obtained from American Type Culture Collection (Manassas, VA, USA) and maintained in Dulbecco’s modified Eagle’s medium (Sigma-Aldrich, St Louis, MO, USA) supplemented with 10% fetal bovine serum and penicillin/streptomycin at 37 °C under 5% CO2. Wild-type mouse embryonic stem (ES) cells (J1) and Dnmt1/Dnmt3a/Dnmt3b triple-knockout (TKO) ES cells17 were maintained in Dulbecco’s modified Eagle’s medium supplemented with 15% fetal bovine serum, 1 × MEM nonessential amino-acid solution (Life Technologies, Carlsbad, CA, USA), 0.1 mM β-mercaptoethanol, 500 U ml−1 leukemia inhibiting factor (LIF) and penicillin/streptomycin on irradiated mouse embryonic fibroblasts at 37 °C under 5% CO2. Cells were transfected with the wild-type EGFP-Zbtb24, two mutant EGFP-Zbtb24, C382Y and R319X, and wild-type EGFP-Dnmt3b expression plasmids using FuGENE HD transfection reagent (Promega, Madison, WI, USA) according to the manufacturer’s protocol.
Immunofluorescence analysis
After 48 h of transfection, the cells grown on glass slides were fixed with 4% formaldehyde at room temperature for 30 min, permeabilized by 0.1% Triton X at 4 °C for 3 min and blocked by 1 × Block Ace (DS Pharma Biomedical, Osaka, Japan) at room temperature for 1 h. Fixed cells were incubated with an anti-HP1α mouse monoclonal antibody (MAB3584; Merck Millipore, Billerica, MA, USA; 1:1000 dilution) for 1 h at room temperature. Then the cells were incubated with Alexa Fluor 594 goat anti-mouse IgG (H+L) antibody (A11005; Life Technologies; 1:1000 dilution) for 1 h at room temperature. Nuclei were visualized by SlowFadeR Gold Antifade Reagent with DAPI (Life Technologies). Fluorescent images were taken using LSM510 confocal laser scanning microscopy (Carl Zeiss, Oberkochen, Germany).
Results
Clinical characteristics
P6: A 5-year-old Japanese boy born in 2003 from nonconsanguineous parents. His birth weight was 2880 g, height was 51.0 cm and occipitofrontal circumference was 32.7 cm. He had craniofacial features including macrocephaly, hypertelorism, epicanthal folds, midface flatness, low nasal root, long and flat philtrum, and thick lips (upper lips < lower lips). He had a thin habitus with slender arms, knock-knee and slightly irregular toes. He had hypoplastic primary teeth and bilateral hydronephrosis. His developmental quotient was 80, with delayed motor development and apparently normal mental development. He showed short stature because of hypopituitarism resulting from compression by Rathke’s cleft cyst, which was treated with replacement of human growth hormone and thyroid hormone. He had skin abnormalities including eruption of blisters at the onset of fever and freckle formation by exposure to sunlight with no butterfly erythema. He suffered from refractory diarrhea and recurrent respiratory tract infections. He finally died of fatal viral infection at the age of 7. His serum IgG2, IgA and IgM levels were decreased and the IgG3 level was increased (Table 1). The number of T cells and B cells were in the normal range, but CD27+ memory B cells were hardly detectable (Table 1). The activity of natural killer cell was low. The lymphocyte blastoid transformation test using phytohaemagglutinin was in the normal range. He did not produce antibodies in response to vaccines or pathogens. A cytogenetic analysis revealed that 7% (2/30 cells) of peripheral blood leukocytes had a large deletion of the long arm of chromosome 1, designated as del(1)(q21) (Figure 1a).
Chromosome abnormalities observed in the two Japanese ICF2 patients, P6 and P7. (a) Giemza-stained mitotic chromosomes of peripheral blood leukocytes from P6. About 7% of the cells lacked the long arm of chromosome 1 (arrow). (b) Representative mitotic chromosome spreads of P7. Multiradial configurations with multiple p and q arms derived from chromosomes 1 and 16 were observed (arrows).
P7: A 39-year-old Japanese man born in 1971 from consanguineous parents. His birth weight was 2600 g. He showed dysmorphic facial features including micrognathia and hypertelorism. He showed a delay in language development and had a learning disability. He had recurrent pneumonia and sinusitis and was on intravenous immunoglobulin therapy. At the age of 39, he had ingravescence of tremor, articulation disorder and left hemiplegia. He also showed involuntary movements of the right lower limb and left toes, choreoathetotic movement of the right upper limb, ambulation difficulty, increased tendon reflex, Babinski’s reflex and vesicorectal disorders accompanied by incontinence without sensory impairment. The brain magnetic resonance imaging showed cortical atrophy. His intelligence quotient was 47. He had progressive multifocal leukoencephalopathy due to John Cunningham virus infection and died at the age of 41. His serum levels of IgG, IgG1, IgG2, IgA and IgM were decreased (Table 1). The number of T cells was in the normal range, but that of B cells was decreased (Table 1). The natural killer cell activity and lymphocyte blastoid transformation count were also decreased. A cytogenetic analysis revealed multiradial chromosome configurations with multiple p and q arms from chromosomes 1 and 16 (Figure 1b).
P8: A 16-year-old Cape Verdean girl born in 1996 from nonconsanguineous parents.18 She had characteristic facial features including epicanthic folds. She had recurrent thrush caused by candida infection in the first year of life. Her initial speech was delayed. She had recurrent ear, nose and throat infections and had a secondary hearing impairment. She suffered from recurrent bacterial pulmonary infections that caused bronchiectasis and atelectasis. She had been on intravenous immunoglobulin therapy since she was 10 years old. She had a lobotomy (left lower lobe) at the age of 12 because of the severe bronchiectasis. She had mental retardation (intelligence quotient <70) and did not acquire reading at the age of 16. Her serum IgG and IgA levels before intravenous immunoglobulin treatment were in the normal range, but the IgM level was decreased (Table 1). The number of T cells was in the normal range, but that of CD27+ memory B cells was decreased (Table 1). She had low antibody titers against tetanus (0.32 IU ml−1), no antibody against pneumococcus and no IgM blood group isohemagglutinin. T cells proliferated normally in response to phytohaemagglutinin and tetanic toxin. A cytogenetic analysis of peripheral blood leukocytes showed that 8% of the cells had whole-arm deletions and/or breaks in chromosomes 1 and 16. Multibranched chromosomes were also observed.18
DNA methylation status at satellites
We examined the DNA methylation status of the satellite-2 and α-satellite repeats in the three ICF syndrome patients by Southern blot analysis using methylation-sensitive restriction enzymes. When satellite-2 was examined, the genomic DNAs from the three patients, as well as an ICF1 patient (P3),19 gave multiple low-molecular-weight bands, indicating that this repeat was hypomethylated (Figure 2a). By contrast, the DNAs from the mother of P6 and a normal control individual (N) were resistant to the digestion. When α-satellite was examined, the DNAs from all patients gave low-molecular-weight bands, whereas those from an ICF1 patient (P3) and a normal control (N) were resistant to the digestion (Figure 2b). Therefore, the three patients P6, P7 and P8 showed a DNA methylation pattern diagnostic of ICF2.
DNA methylation status of the satellite-2 and α-satellite repeats in peripheral blood leukocytes from the three ICF2 patients. (a) Southern blot analyses with the satellite-2 probe. Genomic DNAs from P3 (previously diagnosed as ICF1), P6, P7, P8, the mother of P6 (P6M) and a normal control (N) was digested with a methylation-sensitive restriction enzyme BstBI and separated by 1% agarose gel electrophoresis before blotting. (b) Southern blot analyses with the α-satellite probe. Genomic DNAs from P3, P6, P7, P8 and a normal control (N) was digested with a methylation-sensitive restriction enzyme HhaI and separated by 1% agarose gel electrophoresis.
Identification of ZBTB24 mutations
Sequencing of the PCR products from the three patients revealed no mutation in all coding exons of the DNMT3B gene, further supporting that these patients are ICF2. We therefore looked for mutations in the ZBTB24 gene, which was recently reported as a candidate gene for about a half of ICF2 patients. We found a total of four mutations, three of which were novel (Table 2 and Figure 3a). P6 was homozygous for a G-to-A substitution (c.1148G>A) causing an amino-acid substitution at Cys-383 in a zinc-finger domain (C383Y). His parents were heterozygous for this mutation although they were not consanguineous. P7 was homozygous for a C-to-T substitution (c.958C>T) causing a nonsense mutation at codon 320 (R320X). This mutation was previously reported in ICF2.12 His parents were consanguineous and the mother was found to be heterozygous for the mutation. P8 was a compound heterozygote with a nonsense mutation (c.787A>T) at codon 263 (K263X) and a frame-shift mutation (c.980_981delGT) at codon 327, generating a termination codon at 54-codon downstream (C327W fsX54). Besides the missense mutation identified in P6 (C383Y), only one other missense mutation was previously reported (c.1222T>G, C408G) (Table 2 and Figure 3a).12, 20 Interestingly, the affected cysteines are located in the fourth and fifth C2H2 motifs, respectively, and are highly conserved among vertebrates (Figure 3c). This strongly suggests the importance of these motifs for the protein function and/or structure.
Mutation analysis of the ZBTB24 gene in ICF2 patients. (a) A schematic representation of the ZBTB24 protein structure and locations of all reported mutations including those identified in this study (red) (Table 2).12, 13, 20, 21 (b) Sequencing electrophoregrams of the regions containing the mutations. P6F is the father of P6 and P7M is the mother of P7. Other symbols are as in Figure 2. (c) Amino-acid sequence alignments of the fourth and fifth zinc-finger domains of the vertebrate ZBTB24 proteins (from 374G to 431E of the human ZBTB24 protein). The two amino-acid substitutions, C383Y and C408G, altered the highly conserved cysteine in the C2H2 motifs. Asterisks indicate the positions of amino acids conserved across vertebrates. Two cysteins (red) and two histidines (blue) are highlighted.
Subnuclear localization
To gain insights into the biological function of the zinc-finger domains of ZBTB24 and also the impact of ICF2 mutations on this function, we examined the subnuclear localization of the wild-type and mutant proteins. We used mouse NIH3T3 cells for the analysis because they have distinct pericentromeric heterochromatin foci seen as DAPI-dense regions. The transiently overexpressed wild-type mouse Zbtb24 protein fused with EGFP (EGFP-Zbtb24) showed colocalization with HP1α, a heterochromatin protein, in multiple DAPI-dense regions (Figure 4a). The EGFP-Dnmt3b fusion protein also showed heterochromatin localization (Figure 4a). By contrast, EGFP-Zbtb24 proteins with either of the mutations C382Y and R319X, equivalent to those identified in P6 (C383Y) and P7 (R320X), failed to show such localization, suggesting the importance of the zinc-finger domains in heterochromatin targeting. The mutant proteins showed a rather dispersed distribution in the nuclei (Figure 4a). Using DNA methylation-deficient Dnmt TKO mouse ES cells, we further observed that the complete loss of DNA methylation does not affect heterochromatin localization of the EGFP-Zbtb24 (wild-type) protein, suggesting that Zbtb24 does not require DNA methylation for proper localization (Figure 4b).
Subnuclear localization of mouse Zbtb24 fusion proteins. (a) Confocal microscopic images of NIH3T3 nuclei overexpressing EGFP-fused mouse Zbtb24 without (WT) or with ICF2 mutations (C382Y and R319X) (green). EGFP-Dnmt3b was also analyzed (green). Mouse C382Y is equivalent to human C383Y and mouse R319X is equivalent to human C320X. HP1α (red) was used as a heterochromatin marker and DAPI (blue) was used for DNA staining. Scale bars, 2 μm. (b) Confocal microscopic images of WT ES (J1) and Dnmt TKO ES nuclei overexpressing EGFP-fused mouse Zbtb24 (WT). HP1α (red) was used as a heterochromatin marker and DAPI (blue) was used for DNA staining. Scale bars, 2 μm.
Discussion
In this report, we identified four ZBTB24 mutations, of which three were novel, in three ICF2 patients (Table 2 and Figure 3a). Three of the four mutations were nonsense or frame-shift mutations (R320X, K263X and C327W fsX54). These mutations left the BTB domain and the AT-hook motif intact, but made the zinc-finger domains partially or completely lost. Consistent with our finding, the majority of the ZBTB24 mutations, previously reported in ICF2 patients, were also nonsense or frame-shift mutations (Table 2 and Figure 3a).12, 13, 20, 21 The last mutation that we identified altered a highly conserved cysteine in the C2H2 motif of the fourth zinc-finger domain (C383Y). Interestingly, the only one missense mutation previously reported (C408G) affected a highly conserved cysteine in the next zinc-finger domain (Figure 3c).12, 20 These data highlight the importance of the zinc-finger domains of ZBTB24 in the pathogenesis of ICF2. However, the degree of defect in the ZBTB24 protein structure does not appear to be correlated with the severity of immunodeficiency and other clinical manifestations.
Because ICF2 patients have hypomethylated centromeric and pericentromeric repeats, ZBTB24 appears to have a role in the establishment and/or maintenance of DNA methylation at these repeats. Consistent with this function, we observed heterochromatic localization of the wild-type mouse Zbtb24 fusion protein. By contrast, disruption of the zinc-finger domains by either a missense mutation (C382Y) or a truncation mutation (R319X) abolished the heterochromatin targeting. We also found that wild-type mouse Zbtb24 localizes at heterochromatin even in the absence of DNA methylation. These results indicate that the zinc-finger domains of ZBTB24 are important for the proper localization of this protein, and this localization does not involve methylated DNA. It is possible that ZBTB24 recognizes its target region through interactions with some structural components of the chromatin. Although ZBTB24 mutations affect DNA methylation at both centromeric and pericentromeric repeats, DNMT3B mutations affect only pericentromeric repeats. Therefore, ZBTB24 may interact, directly or indirectly, with a DNA methyltransferase other than DNMT3B. Further studies are needed to clarify the detailed molecular function of ZBTB24 and the pathogenesis of ICF2.
References
Ehrlich, M., Sanchez, C., Shao, C., Nishiyama, R., Kehrl, J., Kuick, R. et al. ICF, an immunodeficiency syndrome: DNA methyltransferase 3B involvement, chromosome anomalies, and gene dysregulation. Autoimmunity 41, 253–271 (2008).
Blanco-Betancourt, C. E., Moncla, A., Milili, M., Jiang, Y. L., Viegas-Pequignot, E. M., Roquelaure, B. et al. Defective B-cell-negative selection and terminal differentiation in the ICF syndrome. Blood 103, 2683–2690 (2004).
Maraschio, P., Zuffardi, O., Dalla Fior, T. & Tiepolo, L. Immunodeficiency, centromeric heterochromatin instability of chromosomes 1, 9, and 16, and facial anomalies: the ICF syndrome. J. Med. Genet. 25, 173–180 (1988).
Smeets, D. F., Moog, U., Weemaes, C. M., Vaes-Peeters, G., Merkx, G. F., Niehof, J. P. et al. ICF syndrome: a new case and review of the literature. Hum. Genet. 94, 240–246 (1994).
Jeanpierre, M., Turleau, C., Aurias, A., Prieur, M., Ledeist, F., Fischer, A. et al. An embryonic-like methylation pattern of classical satellite DNA is observed in ICF syndrome. Hum. Mol. Genet. 2, 731–735 (1993).
Miniou, P., Jeanpierre, M., Blanquet, V., Sibella, V., Bonneau, D., Herbelin, C. et al. Abnormal methylation pattern in constitutive and facultative (X inactive chromosome) heterochromatin of ICF patients. Hum. Mol. Genet. 3, 2093–2102 (1994).
Tuck-Muller, C. M., Narayan, A., Tsien, F., Smeets, D. F., Sawyer, J., Fiala, E. S. et al. DNA hypomethylation and unusual chromosome instability in cell lines from ICF syndrome patients. Cytogenet. Cell Genet. 89, 121–128 (2000).
Jiang, Y. L., Rigolet, M., Bourc'his, D., Nigon, F., Bokesoy, I., Fryns, J. P. et al. DNMT3B mutations and DNA methylation defect define two types of ICF syndrome. Hum. Mutat. 25, 56–63 (2005).
Xu, G. L., Bestor, T. H., Bourc'his, D., Hsieh, C. L., Tommerup, N., Bugge, M. et al. Chromosome instability and immunodeficiency syndrome caused by mutations in a DNA methyltransferase gene. Nature 402, 187–191 (1999).
Okano, M., Bell, D. W., Haber, D. A. & Li, E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell 99, 247–257 (1999).
Hansen, R. S., Wijmenga, C., Luo, P., Stanek, A. M., Canfield, T. K., Weemaes, C. M. et al. The DNMT3B DNA methyltransferase gene is mutated in the ICF immunodeficiency syndrome. Proc. Natl Acad. Sci. USA 96, 14412–14417 (1999).
de Greef, J. C., Wang, J., Balog, J., den Dunnen, J. T., Frants, R. R., Straasheijm, K. R. et al. Mutations in ZBTB24 are associated with immunodeficiency, centromeric instability, and facial anomalies syndrome type 2. Am. J. Hum. Genet. 88, 796–804 (2011).
Weemaes, C. M., Tol, M. J., Wang, J., Ostaijen-Ten Dam, M. M., Eggermond, M. C., Thijssen, P. E. et al. Heterogeneous clinical presentation in ICF syndrome: correlation with underlying gene defects. Eur. J. Hum. Genet. (e-pub ahead of print 13 March 2013; doi:10.1038/ejhg2013.40).
Prokhortchouk, A., Hendrich, B., Jorgensen, H., Ruzov, A., Wilm, M., Georgiev, G. et al. The p120 catenin partner Kaiso is a DNA methylation-dependent transcriptional repressor. Genes Dev. 15, 1613–1618 (2001).
Dent, A. L., Shaffer, A. L., Yu, X., Allman, D. & Staudt, L. M. Control of inflammation, cytokine expression, and germinal center formation by BCL-6. Science 276, 589–592 (1997).
Tagarro, I., Ferna'ndez-Peralta, A. M. & Gonza'lez-Aguilera, J. J. Chromosomal localization of human satellites 2 and 3 by a FISH method using oligonucleotides as probes. Hum. Genet. 93, 383–388 (1994).
Tsumura, A., Hayakawa, T., Kumaki, Y., Takebayashi, S., Sakaue, M., Matsuoka, C. et al. Maintenance of self-renewal ability of mouse embryonic stem cells in the absence of DNA methyltransferases Dnmt1, Dnmt3a and Dnmt3b. Genes Cells 11, 805–814 (2006).
Dupont, C., Guimiot, F., Perrin, L., Marey, I., Smiljkovski, D., Le Tessier, D. et al. 3D position of pericentromeric heterochromatin within the nucleus of a patient with ICF syndrome. Clin. Genet. 82, 187–192 (2012).
Shirohzu, H., Kubota, T., Kumazawa, A., Sado, T., Chijiwa, T., Inagaki, K. et al. Three novel DNMT3B mutations in Japanese patients with ICF syndrome. Am. J. Med. Genet. 112, 31–37 (2002).
Cerbone, M., Wang, J., Van der Maarel, S. M., D'Amico, A., D'Agostino, A., Romano, A. et al. Immunodeficiency, centromeric instability, facial anomalies (ICF) syndrome, due to ZBTB24 mutations, presenting with large cerebral cyst. Am. J. Med. Genet. A. 158A, 2043–2046 (2012).
Chouery, E., Abou-Ghoch, J., Corbani, S., El Ali, N., Korban, R., Salem, N. et al. A novel deletion in ZBTB24 in a Lebanese family with immunodeficiency, centromeric instability, and facial anomalies syndrome type 2. Clin. Genet. 82, 489–493 (2012).
Acknowledgements
We would like to thank all patients and their families for participation in this study. We thank Dr Masaki Okano for providing us with the Dnmt TKO mouse ES cells. We are also grateful for the useful advice of Drs Takashi Sado, Yufeng Li, Tomoko Ichiyanagi and Kei Fukuda, and the technical assistance of Mr. Hiroyasu Furuumi. This study was supported by JSPS KAKENHI 23249019 and a research grant from the Uehara Memorial Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Nitta, H., Unoki, M., Ichiyanagi, K. et al. Three novel ZBTB24 mutations identified in Japanese and Cape Verdean type 2 ICF syndrome patients. J Hum Genet 58, 455–460 (2013). https://doi.org/10.1038/jhg.2013.56
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2013.56
Keywords
This article is cited by
-
Evaluation of Clinical and Immunological Alterations Associated with ICF Syndrome
Journal of Clinical Immunology (2024)
-
Enhanced CD19 activity in B cells contributes to immunodeficiency in mice deficient in the ICF syndrome gene Zbtb24
Cellular & Molecular Immunology (2023)
-
Clinical and Immunological Characterization of ICF Syndrome in Japan
Journal of Clinical Immunology (2018)
-
Genetic, Cellular and Clinical Features of ICF Syndrome: a French National Survey
Journal of Clinical Immunology (2016)
-
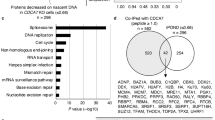
Mutations in CDCA7 and HELLS cause immunodeficiency–centromeric instability–facial anomalies syndrome
Nature Communications (2015)