Abstract
With the advent of large-scale genotyping technologies, enormous quantities of genotype data that were generated have been well exploited through phased haplotypes, and the haplotype-based association study is used as one of the major statistical methods for gene mapping of human complex traits. However, haplotype-based method depends on the information of haplotype frequencies, which results in infeasible computation when haplotypes are not directly observed. This paper provides a genotype-based statistic with multiple tightly linked markers for association analysis using entropy theory. The statistic here does not require haplotype phasing and only requires genotype data. The distribution and the power of the statistic are investigated by simulative study. The results show that the statistic has very reasonable performance. We demonstrated the powerfulness of the statistic by applying our approach to a specific example on hereditary hemochromatosis.
Similar content being viewed by others
Introduction
Linkage disequilibrium (LD), the non-random co-occurrence of alleles from different loci, has a fundamental role in genetic studies as a tool for gene mapping of human complex traits. However, the level of LD is often influenced by a number of factors, including genetic linkage, selection, the rate of recombination and mutation, genetic drift, non-random mating, population structure and other non-biological forces. These bring a great many challenges for LD mapping or association analysis in genetic studies. One of those challenges is to develop novel statistical methods to improve the power of gene mapping. Although many LD methods have been well developed currently for complex disease genes, haplotype-based analysis is one of the major statistical methods because haplotypes of multiple single-nucleotide polymorphisms (SNPs) are considered a more informative format of polymorphisms for genetic analysis than single SNP.1
The classical haplotype-based statistic is to compare haplotype frequencies between affected and unaffected individuals1, 2 or to compare haplotype similarities between affected and unaffected individuals.3, 4 Recently, Zhao et al.5 proposed an entropy-based statistic TPE for a genome-wide association study through a non-linear transformation of haplotype frequencies. Nonetheless, the results of these methods are not uniformly consistent.4, 5 An important reason is that haplotype-based method depends on the information of haplotype frequencies, which results in infeasible computation for estimating haplotype frequencies when haplotypes are not directly observed.
In this paper, we will propose an entropy-based statistic; here we denote it as TGE, as an alternative method of Zhao et al.,5 which only allows for genotype data at linked markers. We will investigate the performance of the statistic TGE by computer simulations and apply it to a real data set on hereditary hemochromatosis (HH).
Materials and methods
Composite LD measure
We consider two marker loci ‘1’ and ‘2’, with co-dominant alleles A and a for loci 1 and B and b for loci 2. The frequencies of alleles A, a, B and b are given by PA, Pa, PB and Pb, and the frequencies of genotype AA, Aa, aa, BB, Bb and bb are given by PAA, PAa, Paa, PBB, PBb and Pbb, respectively. Let PAB, PAb, PaB and Pab be the haplotype frequencies of AB, Ab, aB and ab, respectively. Let P/ be the joint frequency of alleles for loci 1 and 2 in two different gametes, for example, PA/B is the joint frequency of alleles A and B in two different gametes. The composite LD coefficient is defined as ΔAB=PAB+PA/B−2PAPB, and an estimator of ΔAB is defined as  , where
, where  , N is the total number of subjects, n is a count of the number of subjects with the phenotype indicated by its subscript, and P̂A and P̂B are estimates of allele frequencies. It can be seen that these estimators only rely on the information of genotype. We define a two-dimensional random variable X=(X1,X2)T as the state of two alleles A and B, where X1 and X2 denote the number of copies ‘A’ and ‘B’ for marker ‘1’ and ‘2’, respectively. The probability distributions of X1 and X2 are described in Table 1. It can be shown that
, N is the total number of subjects, n is a count of the number of subjects with the phenotype indicated by its subscript, and P̂A and P̂B are estimates of allele frequencies. It can be seen that these estimators only rely on the information of genotype. We define a two-dimensional random variable X=(X1,X2)T as the state of two alleles A and B, where X1 and X2 denote the number of copies ‘A’ and ‘B’ for marker ‘1’ and ‘2’, respectively. The probability distributions of X1 and X2 are described in Table 1. It can be shown that  , and the covariance of X1 and X2 is equal to twice that of the composite LD coefficient ΔAB:cov(X1, X2)=2ΔAB.
, and the covariance of X1 and X2 is equal to twice that of the composite LD coefficient ΔAB:cov(X1, X2)=2ΔAB.
The entropy-based statistic TGE
Suppose that there are k markers, each of which has two alleles A and a. The frequency of allele A is denoted by Pi for ith marker (i=1,…, k). The entropy of the frequencies Pi for the ith marker is defined as Hi=−Pi log Pi.6 Denote the first partial derivatives of the entropy Hi with respect to the frequency Pj as zij: zij=−1−logPi for i=j, zij=0 for i≠j, i, j=1,…, k. To simplify our presentation, a measure with a superscript ‘A’ indicates a measure in affected individuals and a measure with a superscript ‘C’ indicates a measure in unaffected individuals. The frequencies of alleles A in affected individuals for k markers are given by the vector  and the frequencies of alleles A in unaffected individuals are given by the vector
and the frequencies of alleles A in unaffected individuals are given by the vector  . Let
. Let  . Denote Xi as the state of allele A, that is, the number of copies ‘A’ for marker ‘i’, i=1,…, k. Define a k-dimensional random variable X=(X1, X2,…Xk)T. Suppose that there are nA affected individuals and nC unaffected individuals (nA+nC=2N). The total number of the alleles A for marker ‘i’ in nA affected individuals and nC unaffected individuals are denoted as NiA and NiC, respectively.
. Denote Xi as the state of allele A, that is, the number of copies ‘A’ for marker ‘i’, i=1,…, k. Define a k-dimensional random variable X=(X1, X2,…Xk)T. Suppose that there are nA affected individuals and nC unaffected individuals (nA+nC=2N). The total number of the alleles A for marker ‘i’ in nA affected individuals and nC unaffected individuals are denoted as NiA and NiC, respectively.  be the state of allele A for the ith (i=1,·2,…,nA) affected individual and the jth (j=1,·2,…,nC) unaffected individual, respectively. It is easy to see that
be the state of allele A for the ith (i=1,·2,…,nA) affected individual and the jth (j=1,·2,…,nC) unaffected individual, respectively. It is easy to see that

Let  . Note that
. Note that  are the maximum likelihood estimator of the vector PA and PC, respectively, here; let
are the maximum likelihood estimator of the vector PA and PC, respectively, here; let  , and from the asymptotic normality of the maximum likelihood estimator, we have
, and from the asymptotic normality of the maximum likelihood estimator, we have

,
where  for i≠j,7 here, σAii is the variance of Xi for marker ‘i’, and ΔAij and ΔCij are the composite LD coefficient for marker ‘i’ and marker ‘j’, respectively.
for i≠j,7 here, σAii is the variance of Xi for marker ‘i’, and ΔAij and ΔCij are the composite LD coefficient for marker ‘i’ and marker ‘j’, respectively.
 . Let
. Let  , be the estimators of HA, HC, WA and WC, respectively. Then the entropy-based statistic can be defined as
, be the estimators of HA, HC, WA and WC, respectively. Then the entropy-based statistic can be defined as

The statistic TGE is asymptotically a central χ2(k) distribution under the null hypothesis of no association and is asymptotically a non-central χ2(k) distribution8 with the non-centrality parameter  under the alternative hypothesis of association. When the inverse of the matrix
under the alternative hypothesis of association. When the inverse of the matrix  does not exist, the generalized inverse of the matrix will be used.
does not exist, the generalized inverse of the matrix will be used.
Results
Distribution of the statistic TGE and type I error rate
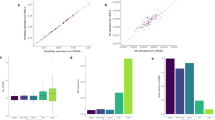
To assess the distribution of the TGE in finite samples, a simulation study is performed to investigate the distribution of the statistic TGE under the null hypothesis of no association. The simulations using the computer program SNaP9 are implemented similar to those in Zhao et al.5 We consider two biallelic marker loci that generate four haplotypes (AB, Ab, aB and ab) with frequencies 0.2952, 0.2562, 0.1957 and 0.2529, respectively. We randomly generate 20 000 individuals in the general population and divide them into equal groups of cases and controls. Then, we randomly sample N individuals (here, nA=nC=N) each from the cases and controls with 10 000 replicates; and thus, it follows 10 000 values of the statistic TGE. Figure 1 plots the histograms of the test statistic TGE when 2N is 200. It can be seen that the distributions of TGE is similar to the theoretical central χ2(k) distributions. For a given significance level α (α=0.05), the type I error rate is estimated as the proportion of rejecting the null hypothesis in 10 000 replicates performed when the null hypothesis holds. The estimated type I error rates for sample sizes (2N) from 200 to 1000 individuals are exhibited in Table 2. We can see that the type I error rates are around the nominal levels α=0.05. All these indicate the validity of the statistic TGE.
The power of the test statistic TGE
To evaluate the power of the statistic TGE and compare the power of the TGE with that of the statistic TPE, we performed simulations under the alternative hypothesis of association. Consider a biallelic disease locus with alleles D and d and two biallelic marker loci that generate four haplotypes (AB, Ab, aB and ab) with frequencies 0.2952, 0.2562, 0.1957 and 0.2529, respectively. The disease allele frequency is taken as PD=0.3. The penetrances of genotypes DD, Dd and dd are denoted by f11, f12 and f22, respectively. The overall LD10 between the four haplotypes and the allele D at the disease locus are chosen as 0.0728, −0.0448, 0.0192 and −0.0472, respectively. We consider four genetic models: additive model, dominant model, recessive model and multiplicative model. Under a specific genetic model, for a particular sample size, the power is estimated as the proportion of rejecting the null hypothesis in 10 000 replicates performed when the alternative hypothesis holds. Figure 2 shows the power curves against sample size at the 0.05 significance level for TGE and TPE. It can be seen that when the sample size is larger than 100, the statistic TGE has consistently higher power than the statistic TPE under the dominant model. The power of the TGE is almost the same as that of the TPE under the multiplicative model and the additive model, except when the sample size is smaller than 200 for the additive model. But we can see that the statistic TPE dominates the statistic TGE under the recessive model when the sample size is smaller than 400.
Power curves against sample size at the 0.05 significance level for the statistic TGE (o) and TPE (*) under the recessive model (a, f11=1, f12=0.1, f22=0.1), the additive model (b, f11=1, f12=0.5, f22=0), the dominant model (c, f11=1, f12=1, f22=0) and the multiplicative model (d, f11=0.81, f12=0.045, f22=0.0025).
Applications to HH
HH is inherited as a recessive disease resulting in excessive iron absorption from diet and leads to chronic disease and early death. It is one of the most common inherited diseases among people of European descent, with an estimated prevalence of 1/200 to 1/400, with an even higher prevalence likely in the Irish population.11 About 1 in 8 to 1 in 10 Australians of Northern European ancestry are genetic carriers for HH.12 In the course of cloning the hemochromatosis gene, genotypes in 101 HH cases and 64 controls were genotyped at 43 microsatellite repeat markers that span the 6.5-Mb HH gene region13 (http://link.springer.de/link/service/journals/00439/index.htm). We analyzed the data using the statistic TGE. To simplify our computation, four markers are used in our analysis: 2229, 2241, 2242 and 2236. We take the ancestral allele given in Thomas et al.13 at each marker as allele A and all the other alleles as allele a. In Table 3, we present the values of the statistic TGE and the corresponding P-values for testing the association of two, three and four SNP markers with HH. It is evident that the statistic TGE obtained a very small P-value.
Discussion
With the availability of high-density maps of SNP markers, population-based LD mapping or association study provides an unprecedented opportunity for identifying genetic variants that influence human complex trait. Haplotype-based analysis is used as one of the major statistical methods for mapping gene of human complex trait. When only genotype data at multiple loci are collected from a sample of unrelated individuals, haplotype-based methods need estimating haplotype phases and frequencies. Although current computational and laboratory methods promise improved determination of haplotype phase, the haplotype-based method is not yet cost-effective.14, 15 Here, we proposed an entropy-based statistic using genotype data for association analysis. The validity and the power of the statistic are demonstrated by simulation analysis and case study. In contrast with haplotype-based association study, our method does not need the haplotype information, and the power of our method is sometimes higher than that of haplotype-based method (for example, the method of Zhao et al.5) when the sample size is not very small. Our paper here mainly investigated the effect of mode of inheritance on the power and the effect of LD between SNPs on the power is not examined. More work awaits to be done with the effect of LD between SNPs in the future.
References
Akey, J., Jin, L. & Xiong, M. Haplotypes vs single marker linkage disequilibrium tests: what do we gain? Eur. J. Hum. Genet. 9, 291–300 (2001).
Chapman, N. H. & Wijsman, E. M. Genome screens using linkage disequilibrium tests: optimal marker characteristics and feasibility. Am. J. Hum. Genet. 63, 1872–1885 (1998).
Bourgain, C., Genin, E., Margaritte-Jeannin, P. & Clerget-Darpoux, F. Maximum identity length contrast: a powerful method for susceptibility gene detection in isolated populations. Genet. Epidemiol. Suppl. 21, S560–S564 (2001).
Tzeng, J. Y., Devlin, B., Wasserman, L. & Roeder, K. On the identification of disease mutations by the analysis of haplotype similarity and goodness of fit. Am. J. Hum. Genet. 72, 891–902 (2003).
Zhao, J. Y., Boerwinkle, E. & Xiong, M. An entropy-based statistic for genomewide association studies. Am. J. Hum. Genet. 77, 27–40 (2005).
Shannon, C. E. A mathematical theory of communication. Bell. Systems Tech. J. 27, 379–423 (1948).
Rao, C. R. Linear Statistical Inference and its Applications, John Wiley: New York, (1973).
Lehmann, E. L. Theory of Point Estimation, John Wiley & Sons: New York, (1983).
Nothnagel, M. Simulation of LD block-structured SNP haplotype data and its use for the analysis of case-control data by supervised learning methods. Am. J. Hum. Genet. 71 (Suppl), A2363 (2002).
Xiong, M., Zhao, J. Y. & Boerwinkle, E. Haplotype block linkage disequilibrium mapping. Front. Biosci. 8, 85–93 (2003).
Feder, J. N., Gnirke, A., Thomas, W., Tsuchihashi, Z., Ruddy, D. A., Basava, A. A. et al. A novel MHC class I-like gene is mutated in patients with hereditary haemochromatosis. Nat. Genet. 13, 399–408 (1996).
Edward, C. Q., Griffen, L. M., Goldgar, D., Drummond, C., Solnick, M. H. & Kushner, J. P. Prevalence of haemochromatosis among 11,065 presumbly healthy donors. N Engl. J. Med. 318, 1355–1362 (1988).
Thomas, W., Fullan, A., Loeb, D. B., McClelland, E. E., Bacon, B. R. & Wolff, R. K. A haplotype and linkage disequilibrium analysis of the hereditary haemochromatosis gene region. Hum. Genet. 102, 517–525 (1998).
Browning, S. R. & Browning, B. L. Haplotype phasing: existing methods and new developments. Nat. Rev. Genet. 12, 703–714 (2011).
Ruiz-Marín, M., Matilla-García, M., Cordoba, J. A., Susillo-González, J. L., Romo-Astorga, A., González-Pérez, A. et al. An entropy test for single-locus genetic association analysis. BMC Genet. 23, 11–19 (2010).
Acknowledgements
This study was supported by the Foundation of Hunan Educational Committee (11B095) and National Social Science Fund Youth Project (11CTJ003).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Li, YM., Xiang, Y. Genotype-based association analysis via entropy. J Hum Genet 57, 734–737 (2012). https://doi.org/10.1038/jhg.2012.102
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2012.102
Keywords
This article is cited by
-
An adaptive strategy for association analysis of common or rare variants using entropy theory
Journal of Human Genetics (2017)