Abstract
To elucidate the mechanism underlying transport and processing defects from the viewpoint of enzyme folding, we constructed three-dimensional models of human acid α-glucosidase encompassing 27 relevant amino acid substitutions by means of homology modeling. Then, we determined in each separate case the number of affected atoms, the root-mean-square distance value and the solvent-accessible surface area value. The analysis revealed that the amino acid substitutions causing a processing or transport defect responsible for Pompe disease were widely spread over all of the five domains comprising the acid α-glucosidase. They were distributed from the core to the surface of the enzyme molecule, and the predicted structural changes varied from large to very small. Among the structural changes, we paid particular attention to G377R and G483R. These two substitutions are predicted to cause electrostatic changes in neighboring small regions on the molecular surface. The quality control system of the endoplasmic reticulum apparently detects these very small structural changes and degrades the mutant enzyme precursor (G377R), but also the cellular sorting system might be misled by these minor changes whereby the precursor is secreted instead of being transported to lysosomes (G483R).
Similar content being viewed by others
Introduction
Acid α-glucosidase (GAA; EC 3. 2.1.20/3) is a lysosomal exoglycosidase that catalyzes the hydrolysis of the α-1,4- and α-1,6-glucosidic bonds of glycogen, yielding glucose. The enzyme is encoded by the GAA gene localized to 17q25, and is synthesized as a protein of 952 amino acids including an amino-terminal signal peptide for transport into the lumen of the endoplasmic reticulum (ER). On entry into the ER, the enzyme is N-glycosylated, resulting in a precursor form with an apparent molecular mass of 110 kDa. Subsequently, the enzyme is transported to the Golgi apparatus, in which further modification of the sugar chains occurs. Proteolytic processing in the trans-Golgi apparatus and in the lysosomes, in which the enzyme exerts its function, results in a 95-kDa intermediate form and finally the mature forms of 76 and 70 kDa. The small peptides that are cleaved-off remain linked to the protein core through disulfide linkages. Some cell types, such as fibroblasts, secrete a small amount of the newly formed 110-kDa acid α-glucosidase precursor.1, 2, 3, 4
A genetic deficiency of acid α-glucosidase causes the accumulation of glycogen in the lysosomes, and thereby leads to Pompe disease (glycogen storage disease type II, acid maltase deficiency; OMIM 232300). The clinical spectrum of this disease ranges from an early-onset rapidly progressive infantile form characterized by cardiac failure, severe hypotonia and sometimes hepatomegaly, to a late-onset slowly progressive juvenile/adult forms characterized by skeletal muscle weakness.1 The overall incidence of this disease has been estimated to be from 1 in 14 000 to 1 in 137 000.1 In a recent study involving newborn screening in Taiwan, four patients with Pompe disease were discovered among 132 538 apparently unaffected newborns.5
So far, close to 300 genetic mutations responsible for Pompe disease have been reported.1, 6, 7, 8 Among them there is a group of mutations that affects the post-translational processing and transport of the enzyme protein. As it is well known that the detrimental effect of most mutations in lysosomal diseases is on protein folding, giving rise to processing and transport defects,9 it is necessary to elucidate the structural cause of processing and transport defects. This will facilitate in understanding the molecular basis of Pompe disease. However, information regarding this is very scarce.
Earlier, we built a structural model of the catalytic domain and the surrounding region of human acid α-glucosidase by means of homology modeling using the structural information on α-glucosidase MalA from Sulfolobus solfataricus (PDB: 2G3N)10 as a template, and examined the structural changes caused by pathogenic and non-pathogenic amino acid substitutions.11 However, the investigations were restricted to a limited region of the enzyme protein because the amino acid sequence identity between the human acid α-glucosidase and the template was low.
In this study, we constructed an improved three-dimensional model of human acid α-glucosidase involving the whole structure using the recently reported crystal structure of the N-terminal subunit of human intestinal maltase-glucoamylase12 as a template. We then used this new model to investigate genetically determined protein structural changes giving rise to defects in the processing and transport of acid α-glucosidase in Pompe disease.
Materials and methods
Amino acid substitutions causing a processing/transport defect in Pompe disease
As mentioned above, close to 300 genetic mutations associated with Pompe disease have been reported (http://www.pompecenter.nl/). Here, we analyzed the structural implications of 27 specific amino acid substitutions, which were earlier shown to cause a processing/transport defect judging from the results of biochemical investigations. The amino acid substitutions, biochemical data and references to relevant literature are listed in Table 1. The Pompe disease mutation database at http://www.pompecenter.nl/ provides an additional background information, which can easily be accessed by electronic links to all publications dealing with each of the 27 amino acid substitutions described in the present publication, including case reports if available.
Structural modeling of the wild-type and mutant human acid α-glucosidase
A structural model of wild-type human acid α-glucosidase was built using molecular modeling software, Molecular Operating Environment, MOE (CCG-Chemical Computing Group Inc., Montreal, Quebec, Canada), by means of homology modeling. The crystal structure of N-terminal subunit of human intestinal maltase-glucoamylase (PDB: 2QLY) was used as a template, and energy minimization was carried out. The root-mean-square gradient value was set at 0.05 kcal/molÅ. The structural models of mutant species incorporating the amino acid substitutions were constructed using molecular modeling software, TINKER, developed by Ponder et al.13, 14, 15, 16, 17
Calculation of the numbers of atoms influenced by amino acid substitutions
To determine the number of atoms affected, each mutant model was superimposed on the wild-type structure on the basis of the Cα atoms by the least-square-mean fitting method.18, 19, 20, 21, 22 We defined that the structure was influenced by an amino acid substitution when the position of an atom in a mutant differed from that in the wild-type structure by more than the cut-off distance (0.15 Å) on the basis of the total root-mean-square distance (RMSD), as described earlier.18, 19, 20, 21, 22 The numbers of influenced atoms in the main chain and the side chain were calculated.
Determination of RMSD values of all atoms in mutant human acid α-glucosidase structures
The RMSD values of all atoms in the mutant enzyme structures were determined according to Weiner's method to predict the severity of the structural changes, as described earlier.20, 22, 23
Determination of the solvent-accessible surface area (ASA) values of amino acid residues causing a processing/transport defect in Pompe disease
The ASA value of each residue in the wild-type human acid α-glucosidase was calculated using ACCESS,24 and the position of a substituted amino acid residue causing a processing/transport defect was determined, as described earlier.19, 20, 21, 22
Coloring of the atoms influenced by representative amino acid substitutions
Coloring of the influenced atoms in the three-dimensional structure of human acid α-glucosidase was carried out for representative amino acid substitutions including G377R and G483R, which were predicted to cause structural changes in a small region on the surface of the molecule. The analysis is on the basis of the geographic distances between the wild-type and substituted amino acid residues in the three-dimensional model, according the method described earlier.19, 20, 21, 22, 25
Structural modeling of the surface region affected by representative amino acid substitutions
The surface structure of wild-type human acid α-glucosidase and the mutant structures with the G377R and G483R substitutions were built using Jmol, version 11 (an open-surface Java viewer for chemical structures in 3D), and the molecular surfaces were presented by coloring.
Results
Structural model of human acid α-glucosidase and locations of amino acid substitutions associated with a processing/transport defect in Pompe disease
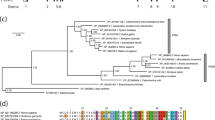
To investigate structural changes that give rise to processing/transport defects in Pompe disease, we constructed a three-dimensional model of human acid α-glucosidase encompassing the whole structure by means of homology modeling. We used the existing crystallographic data for the N-terminal subunit of human intestinal maltase-glucoamylase as a template. Figure 1 shows the sequence alignment of these enzymes, human acid α-glucosidase (LYAG) and human intestinal maltase-glucoamylase (PDB: 2QLY), completed with α-glucosidase MalA from S. solfataricus (PDB: 2G3N) that was earlier used for three-dimensional modeling.11 The amino acid identity between human acid α-glucosidase and the N-terminal subunit of human intestinal maltase-glucoamylase was 41%, this being sufficiently high to model the entire structure, whereas the homology with S. solfatarics MalA was only 28 %, this being too low to construct the entire model. The residues responsible for the catalytic activity (D518 and D616) and sequence motifs of family 31 glycosyl hydrolases were well conserved.26
Sequence alignment of human acid α-glucosidase (LYAG), the N-terminal subunit of human intestinal maltase-glucoamylase (2QLY) and Sulfolobus solfataricus MalA (2G3N). Catalytic residues are indicated by gray shadowing and conserved sequence regions of family 31 glycosyl hydrolases are surrounded by boxes. The secondary structure scheme for the catalytic subunit of the N-terminal subunit of human intestinal maltase-glucoamylase (2QLY) is shown with reference to those reported by Sim et al.12 The seven N-glycosylation sites of human acid α-glucosidase (LYAG) are indicated by arrowheads below the sequences. Cleavage sites of LYAG (CS 1, 2 and 3) are indicated by solid arrows above the sequences according to Moreland et al.3 Disulfide bridges in the trefoil type-P domain of LYAG link C82 to C109; C92 to C108 and C103 to C127.3, 36
The overall structure of the template, human intestinal maltase-glucoamylase and the modeled structure of human acid α-glucosidase are superimposed in Figure 2a. According to the constructed model, human acid α-glucosidase is composed of five domains; that is, a trefoil type-P domain (residues 89–135), an N-terminal β-sandwich domain (residues 136–346), a catalytic (β/α)8 barrel domain (residues 347–723) with two inserted loops, which include insert 1 (residues 444–491) and insert 2 (residues 522–567) protruding out between β3 and α3, and between β4 and α4, respectively, a proximal C-terminal domain (residues 724–818) and a distal C-terminal domain (residues 819–952) (Figure 2b). We determined the locations of the putative acid/base catalyst D616 and the nucleophile D518, and the amino acid residues giving rise to processing/transport defects when substituted by other residues through mutation (Table 1). The locations are shown in Figure 3, representing the predicted structure of the wild-type human acid α-glucosidase. The amino acid residues involved in substitutions were distributed over all of the five domains comprising the enzyme molecule, although they were relatively far away from the catalytic residues.
Superimposed Cα traces of human acid α-glucosidase and template N-terminal subunit of human intestinal maltase-glucoamylase, and the domains of human acid α-glucosidase. (a) Human acid α-glucosidase and human intestinal maltase-glucoamylase are depicted in blue and green, respectively. Yellow lines indicate disulfide bonds. N-Glycosylation sites (N140, N233, N390, N470, N652, N882 and N925) are indicated by black arrows. Major cleavage sites (CS 1, 2 and 3) are indicated by red arrows. Front view (left) and side view (right). (b) The individual domains of human GAA are colored as follows: trefoil type-P domain (cyan), N-terminal domain (yellow), catalytic (β/α)8 barrel domain (red), proximal C-terminal domain (blue) and distal C-terminal domain (green). The structure is presented as a ribbon model. N-Glycosylation sites and major cleavage sites are indicated by black and red arrows, respectively. The color code is according to Sim et al.,12 for 2QLY. Front view (left) and side view (right).
Locations of the amino acid residues that cause processing/transport defects in human acid α-glucosidase when substituted. The backbones are displayed as a ribbon model. The positions of both the catalytic residues (red) and the residues subject to substitution (orange) are displayed as space filling models. Front view (a) and side view (b).
Structural changes in human acid α-glucosidase caused by amino acid substitutions associated with a processing/transport defect in Pompe disease
We used the constructed three-dimensional model of each of the mutant proteins to calculate the number of atoms influenced by the amino acid substitution. The results are summarized in Table 1. The average numbers were 66 in the main chain and 79 in the side chain, with very high standard deviation (s.d.) values of 125 and 150 for the two types of chains, respectively. Regarding the main chain, the number of affected atoms was <10 for 10 of the 27 substitutions (37% of cases), ⩾10 and <50 for 7 other substitutions (26% of cases) and ⩾50 in the remaining 10 cases (37%). As to the side chain, the number of affected atoms was <10 for 7 of the 27 substitutions (26% of cases), ⩾10 and <50 for 9 other substitutions (33% of cases) and ⩾50 in the remaining 11 cases (41%).
The RMSD values for the amino acid substitutions associated with a processing/transport defect were also determined and are included in Table 1. The average RMSD value of the substitutions was 0.043 Å, with again a very high s.d. value of 0.042 Å. In 10 of the 27 cases (37%), the RMSD value was <0.02 Å, in 7 other cases (26%) it was ⩾0.02 Å and <0.04 Å and in the remaining 10 cases (37%) the RMSD value was ⩾0.04 Å. The variation in the RMSD value paralleled the number of atoms affected in the main chain.
To determine the locations of the amino acid residues involved in substitutions that cause a processing/transport defect, the ASA values of the residues in the wild-type structure were calculated (Table 1). The average ASA value was 5.7 Å2 (s.d. 9.5 Å2). In 13 of the 27 cases (48 %), the ASA value was <1.0 Å2, in 11 other cases (41 %) it was ⩾1.0 Å2 and <10 Å2 and in the 3 remaining cases (11%) it was ⩾10 Å2.
We paid particular attention to G377R and G483R. According to Sim et al.,12 G377 is predicted to be located in the loop region between β1 and α1, and G483 in insert 1 of the human acid α-glucosidase structure. For both residues, the ASA values were high (⩾25 Å2), but the numbers of affected atoms in the main chain were very small (⩽5), as were the RMSD values (<0.02 Å). These figures suggest that each of the two amino acid substitutions causes a small structural change on the surface of acid α-glucosidase. Coloring of the atoms affected by the mutations supported this interpretation, and it revealed that they affected neighboring small regions on the molecular surface (Figure 4).
Color imaging of the atoms affected by representative amino acid substitutions. Front view (left) and side view (right) of G377R (a) and G483R (b). Each atom is colored according to the distance between the atom in the mutant and the corresponding atom in the wild-type structure. The color code is as follows: gray <0.15 Å; cyan ⩾0.15 Å and <0.30 Å and green ⩾0.30 Å. The whole structure is depicted as a ball and stick model, and the affected atoms as a space-filling model.
Finally, we investigated what changes G377R and G483R might cause on the molecular surface. As the uncharged amino acid glycine is replaced by a positively charged arginine, both these substitutions are expected to affect neighboring residues and to locally affect the electrostatic surface of the enzyme, as shown in Figure 5. On comparing the wild-type and mutant structures, G377R was predicted to affect R375, W376, Y378, S379, Q386, N417 and L678, whereas G483R was predicted to influence R411, S449 and S484.
Surface presentations of the wild-type human acid α-glucosidase and two pathogenic mutants. The molecular surface of the wild type (a), the G377R (b) and the G483R (c) mutants are shown in front view. Both the whole structure (left) as well as the region affected by the amino acid substitutions (right) are depicted. The molecular surface of G377 and G483 in the wild type (a) and that of residues affected by each substitution in the mutants (b and c) are colored according to the character: Yellow (neutral), green (aromatic) and blue (basic).
Discussion
Processing and transport defects of lysosomal enzymes commonly occur as a result of missense mutations in the respective genes and cause lysosomal diseases.8, 9 To elucidate the basis of such defects in Pompe disease from a structural viewpoint, we constructed three-dimensional models that allowed us to compare the structures of the wild-type and mutant forms of human acid α-glucosidase.
In this study, we used the crystal structure of the N-terminal subunit of human intestinal maltase-glucoamylase, which is highly homologous to human acid α-glucosidase, as a template, and we managed to construct a tentative model encompassing the entire structure of the enzyme. The analysis revealed that the overall structure of human acid α-glucosidase probably resembles that of the template quite closely. Likewise, the enzyme protein has five domains. The trefoil type-P domain is characteristically found in all subunits of maltase-glucoamylase and sucrose-isomaltase, but its function is as yet unknown. The catalytic domain has a (β/α)8 barrel structure (Figures 2b and 3) and is shared by α-amylases and several other enzymes with rather different function.27 Precise analysis of the structural differences between the catalytic domain of human acid α-glucosidase and maltase-glucoamylase may reveal the basis of their different substrate specificities. It is noted that the cleavage sites in human acid α-glucosidase in which proteolytic processing occurs are located at positions where the primary sequences of both enzymes deviate one and another (Figures 1 and 2).3, 4 The projected locations of the disulfide bonds suggest how the various peptides that arise from proteolytic processing are held together. Of further note, the seven N-linked glycosylation sites predicted by the primary structure of human acid α-glucosidase and, indeed, occupied by carbohydrate side chains are all located at the outer surface of the molecule (Figure 2).28, 29
Then, we mapped the locations of amino acid residues involved in substitutions that were earlier shown to hamper the post-translational processing and transport of human acid α-glucosidase (Table 1). In these cases, the 110-kD precursor enzyme was synthesized, but defective processing and transport is indicated by the far less than normal amounts of the acid α-glucosidase species of 95 kDa and 76/70 kDa detectable by polyacrylamide gel electrophoresis in the presence of SDS-polyacrylamide gel electrophoresis (PAGE).8 The processing and transport defects can also be deduced from the lack of secretion of the 110-kDa α-glucosidase precursor, and from the higher degree of sialylation of the carbohydrate side-chains that can occur when a mutant protein is retarded in the trans-Golgi cisternae. This leads to an increase in the apparent molecular mass on SDS-PAGE.4 In the constructed three-dimensional model of human acid α-glucosidase, the amino acid substitutions giving rise to processing/transport defects seemed to be distributed in the all five domains and from the core to the surface of the molecule. The structural changes caused by each individual substitution varied from small to large, as judged with several criteria including the numbers of affected atoms in the main chain and side chain, and the RMSD values.
Among the substitutions, we investigated G377R and G483R in more detail, as they were predicted to cause only minor structural changes in a small region on the molecular surface. The replacement of an uncharged residue by a positively charged residue is apparently sufficient to change the conformation of the enzyme precursor such that it leads to a folding or transport defect. It is noteworthy that G377R is not secreted and probably does not pass the ER quality control system, whereas G483R, having excessive sialic acids, must have passed through trans-Golgi apparatus, being secreted in a normal amount and converted, to some extent, into the 95-kDa processing intermediate (Table 1). Thus, we tentatively conclude that the region in question is sensitive to even small structural changes, with far reaching consequences for proper folding and transport of human acid α-glucosidase.
In conclusion, through molecular modeling involving the recently published crystal structure of the homologous intestinal enzyme maltase-glucoamylase, we have provided one of the very first insights into the tentative three-dimensional structure of human acid α-glucosidase. The model awaits confirmation and probably minor corrections, but all attempts to crystallize the enzyme have failed so far. A simpler three-dimensional model not covering the entire length of the protein has already shown its value for predicting the location of amino acid residues that are involved in catalytic function.11
We have used this new model, encompassing the entire structure of acid α-glucosidase, to gain an impression of the structural changes brought about by amino acid substitutions that affect the post-translational processing and transport of the enzyme en route from the ER to lysosomes and causing Pompe disease.
Among the substitutions, we focused on G377R and G483R resulting in electrostatic changes that particularly influence small neighboring regions on the molecular surface. Even these minor changes are apparently detected by the ER's quality control system or make the enzyme vulnerable to degradation in other pre-lysosomal or lysosomal compartments. Structural studies will not only help us to further elucidate the molecular basis of Pompe disease, but also to clarify the mechanism underlying protein quality control.
References
Hirschhorn, R. & Reuser, A. J. J. in The Metabolic and Molecular Bases of Inherited Disease (eds Scriver, C.R., Beaudet, A.L., Sly, W.S.& Valle, D.) 3389–3420 (McGraw-Hill, New York, 2001).
Hasilik, A. & Neufeld, E. F. Biosynthesis of lysosomal enzyme in fibroblasts. Phosphorylation of mannose residues. J. Biol. Chem. 255, 4946–4950 (1980).
Moreland, R. J., Jin, X., Zhang, X. K., Decker, R. W., Lee, K. L., Cauthron, R. D. et al. Lysosomal acid alpha-glucosidase consists of four different peptides processed from a single chain precursor. J. Biol. Chem. 280, 6780–6791 (2005).
Wisslaar, H. A., Kroos, M. A., Hermans, M. M. P., van Beeuman, J. & Reuser, A. J. J. Structural and functional changes of lysosomal acid α-glucosidase during intracellular transport and maturation. J. Biol. Chem. 268, 2223–2231 (1993).
Chien, Y. H., Chiang, S. C., Zhang, X. K., Keutzer, J., Lee, N. C., Huang, A. C. et al. Early detection of Pompe disease by newborn screening is feasible: results from the Taiwan screening program. Pediatrics 122, e39–e45 (2008).
Hermans, M. M., Van Leenen, D., Kroos, M. A., Beesley, C. E., Van Der Ploeg, A. T., Sakuraba, H. et al. Twenty-two novel mutations in the lysosomal alpha-glucosidase gene (GAA) underscore the genotype- phenotype correlation in glycogen-storage disease type II. Hum. Mutat. 23, 47–56 (2004).
Montalvo, A. L., Bembi, B., Donnarumma, M., Filocamo, M., Parenti, G., Rossi, M. et al. Mutation profile of the GAA gene in 40 Italian patients with late onset glycogen storage disease type II. Hum. Mutat. 27, 999–1006 (2006).
Kroos, M., Pomponio, R. J., van Vliet, L., Palmer, R. E., Phipps, M., Van der Helm, R. et al. GAA Database Consortium, Update of the Pompe disease mutation database with 107 sequence variant and a format for severity rating. Hum. Mutat. 29, e13–e26 (2008).
Mahuran, D. J. Biochemical consequences of mutations causing the GM2 gangliosidoses. Biochem. Biophys. Acta. 1455, 105–138 (1999).
Ernst, H. A., Leggio, L. L., Willemoës, M., Leonard, G., Blum, P. & Larson, S. Structure of the Sulfolobus solfatarics alpha-glucosidase: implications for domain conservation and substrate recognition in GH31. J. Mol. Biol. 358, 1106–1124 (2006).
Tajima, Y., Matsuzawa, F., Aikawa, S., Okumiya, T., Yoshimizu, M., Tsukimura, T. et al. Structural and biochemical studies on Pompe disease and a ‘pseudodeficiency of acid alpha-glucosidase’. J. Hum. Genet 52, 898–906 (2007).
Sim, L., Quezade-Calvillo, R., Sterchi, E. E., Nichols, B. L. & Rose, D. R. Human intestinal maltase-glucoamylase: crystal structure of the N-terminal catalytic subunit and basis of inhibition and substrate specificity. J. Mol. Biol. 375, 782–792 (2008).
Kundrot, C. E., Ponder, J. W. & Richards, F. M. Algorithms for calculation excluded volume and its derivative as a function of molecular conformation and their use in energy minimization. J. Comput. Chem. 12, 402–819 (1991).
Dudek, M. J. & Ponder, J. M. Accurate modeling of the intramolecular electrostatic energy of proteins. J. Comput. Chem. 16, 791–816 (1995).
Kong, M. J. & Ponder, J. W. Reaction field methods for off-center multipoles. J. Chem. Phys. 107, 481–492 (1997).
Pappu, R. V., Hart, R. W. & Ponder, J. W. Analysis and application of potential energy smoothing for global optimization. J. Phys. Chem. B. 102, 9725–9742 (1998).
Ren, P. & Ponder, J. W. Polarizable atomic multipole water model for molecular mechanics simulation. J. Phy. Chem. B. 107, 5933–5947 (2003).
Matsuzawa, F., Aikawa, S., Doi, H., Okumiya, T. & Sakuraba, H. Fabry disease: correlation between structural changes in alpha-galactosidase, and clinical and biochemical phenotype. Hum. Genet. 117, 317–328 (2005).
Saito, S., Ohno, K., Sugawara, K. & Sakuraba, H. Structural and clinical implications of amino acid substitutions in N-acetylgalactosamine-4-sulfatase: insight into mucopolysaccharidosis type VI. Mol. Genet. Metab. 93, 419–425 (2008).
Sugawara, K., Saito, S., Ohno, K., Okuyama, T. & Sakuraba, H. Structural study on mutant alpha-L-iduronidase: insight into mucopolysaccharidosis type I. J. Hum. Genet. 53, 467–474 (2008).
Ohno, K., Saito, S., Sugawara, K. & Sakuraba, H. Structural consequences of amino acid substitutions causing Tay-Sachs disease. Mol. Genet. Metab. 94, 462–468 (2008).
Sugawara, K., Ohno, K., Saito, S. & Sakuraba, H. Structural characterization of mutant alpha-galactosidases causing Fabry disease. J. Hum. Genet. 53, 812–824 (2008).
Weiner, S. J., Kallman, P. A., Case, D. A., Singh, U. C., Ghio, C., Alagona, G. et al. A new force field for molecular mechanical simulation of nucleic acid and proteins. J. Am. Chem. Soc. 106, 765–784 (1984).
McDonald, I. K. & Thornton, J. M. Satisfying hydrogen bonding potential in proteins. J. Mol. Biol. 238, 777–793 (1994).
Yoshimizu, M., Tajima, Y., Matsuzawa, F., Aikawa, S., Iwamoto, K., Kobayashi, T. et al. Binding parameters and thermodynamics of the interaction of imino sugars with a recombinant human acid alpha-glucosidase (alglucosidase alfa): insight into the complex formation mechanism. Clin. Chim. Acta. 391, 68–73 (2008).
Davies, G. & Henrissat, B. Structures and mechanism of glycosyl hydrolases. Structure 3, 853–859 (1995).
Farber, G. K. & Petsko, G. A. The evolution of α/β barrel enzyme. TIBS 15, 228–234 (1990).
Hoefsloot, L. H., Hoogeveen-Westerveld, M., Kroos, M. A., Van Beeumen, J., Oostra, B. A. & Reuser, A. J. J. Primary structure and processing of lysosomal α-glucosidase; homology with the intestinal sucrase-isomaltase complex. EMBO J 7, 1697–1704 (1998).
Hermans, M. M., Wisselar, H. A., Kroos, M. A., Oostra, B. A. & Reuser, A. J. J. Human lysosomal α-glucosidase: functional characterization of the glycosylation sites. Biochem. J 289, 681–686 (1993).
Fernandez-Hojas, R., Huie, M. L., Navarro, C., Dominguez, C., Roig, M., Lopez-Coronas, D. et al. Identification of six novel mutations in the acid alpha-glucosidase gene in three Spanish patients with infantile onset glycogen storage disease type II (Pompe disease). Neuromusc. Disord 12, 159–166 (2002).
Van den Hout, J. M., Kamphoven, J. H., Winkel, L. P., Arts, W. F., De Klerk, J. B., Loonen, M. C. et al. Long-term intravenous treatment of Pompe disease with recombinant human alpha-glucosidase from milk. Pediatrics 113, e448–e457 (2004).
Gort, L., Coll, M. J. & Chabás, A. Glycogen storage disease type II in Spanish patients: high frequency of c.1076-1G>C mutation. Mol. Genet. Metab. 92, 183–187 (2007).
Müller-Felber, W., Horvath, R., Gempel, K., Podskarbi, T., Shin, Y., Pongratz, D. et al. Late onset Pompe disease: clinical and neurophysiological spectrum of 38 patients including long-term follow-up in 18 patients. Neuromusc. Disord. 17, 698–706 (2007).
Hermans, M. M., de Graaff, E., Kroos, M. A., Wisselaar, H. A., Willemsen, R., Oostra, B. A. et al. The conservative substitution Asp-645 → Glu in lysosomal alpha-glucosidase affects transport and phosphorylation of the enzyme in an adult patient with glycogen-storage disease type II. Biochem. J. 289, 687–693 (1993).
Kroos, M. A., Kirschner, J., Gellerich, F. N., Hermans, M. M., Van Der Ploeg, A. T., Reuser, A. J. et al. A case of childhood Pompe disease demonstrating phenotypic variability of p.Asp645Asn. Neuromusc. Disord. 14, 371–374 (2004).
Reuser, A. J. J. & Kroos, M. A. in Lysosomal Storage Disorders (eds Barranger, J.A. & Cabrera-Salazar, M.A.) 473–498 (Springer Science and Business Media, LLC, 2007).
Acknowledgements
We thank Dr J Ponder (the Department of Biochemistry and Molecular Biophysics, Washington University) for providing us with the TINKER software. This work was partly supported by grants from the Japan Society for the Promotion of Science; the High-Tech Research Center Project of the Ministry of Education, Science, Sports and Culture of Japan; the Ministry of Health and Welfare of Japan; the Japan Science and Technology Agency and CREST.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sugawara, K., Saito, S., Sekijima, M. et al. Structural modeling of mutant α-glucosidases resulting in a processing/transport defect in Pompe disease. J Hum Genet 54, 324–330 (2009). https://doi.org/10.1038/jhg.2009.32
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/jhg.2009.32
Keywords
This article is cited by
-
Mutations in GAA Gene in Tunisian Families with Infantile Onset Pompe Disease: Novel Mutation and Structural Modeling Investigations
Journal of Molecular Neuroscience (2020)
-
Genotype-phenotype correlation in Pompe disease, a step forward
Orphanet Journal of Rare Diseases (2014)
-
Biochemical and structural study on a S529V mutant acid α-glucosidase responsive to pharmacological chaperones
Journal of Human Genetics (2011)