Abstract
Multiple single nucleotide polymorphisms (SNPs) within ADAM33 have been reported to be associated with asthma and bronchial hyper-responsiveness in Caucasian populations. We examined whether these SNPs contribute to a predisposition to asthma, especially aspirin-intolerant asthma (AIA), in the Japanese population. Ten polymorphic sites (ST+4, ST+7, T1, T2, T+1, V-3, V-2, V-1, V4, V5) were genotyped in 102 AIA patients, 282 aspirin-tolerant asthma (ATA) patients and 120 control (CTR) subjects by direct sequencing. Haplotype frequencies were estimated by the expectation-maximization method. Differences in allele and haplotype frequencies among phenotypes were analyzed by the chi-square and permutation tests. ST+7, V-1 and V5 sites in the AIA group were significantly different from those in the ATA group (P=0.034–0.004) and from those in the CTR group (P=0.019–0.002). Haplotypes at three sites (ST+7, V-1, and V5) were significantly different in frequency between the AIA and ATA (P=0.008) or CTR (P=0.001) groups. Sequence variations in ADAM33 are likely to correlate with susceptibility to AIA in the Japanese population.
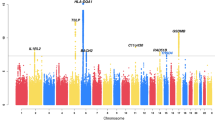
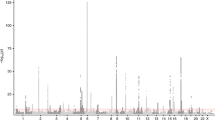
Similar content being viewed by others
Introduction
Bronchial asthma (BA) is a common respiratory disease characterized by chronic airway inflammation leading to bronchial hyper-responsiveness (BHR) and reversible airway obstruction. Although aspects of BA susceptibility can be explained by environmental factors such as allergen exposure and inflammation, genetic factors may also play a key role in its etiology. Multiple sequence variations within the A-disintegrin and metalloprotease (ADAM) 33 gene were reported to be associated with the asthma phenotype and BHR in Caucasian populations (Van Eerdewegh et al. 2002). The ADAM family comprises membrane-anchored metalloproteases with multiple biological functions. For example, some ADAMs have important roles in the shedding of cell surface proteins, such as cytokines and cytokine receptors (Black et al. 1997; Garton et al. 2001; Asakura et al. 2002). Although the functional role of ADAM33 remains poorly understood, ADAM33 probably confers protease activity due to its conserved zinc-binding motif in the metalloprotease domain (Yoshinaka et al. 2002). ADAM33 is abundantly expressed in smooth muscle cells of airway tissue, which might indicate that it plays an important role in airway remodeling and BHR. Aspirin-intolerant asthma (AIA) is a clinically distinct subtype of BA characterized by severe broncho-constriction after oral administration of aspirin or other non-steroidal anti-inflammatory drugs (NSAIDs). The prevalence of AIA, based on respiratory symptoms following NSAIDs use, is reported to be 0.6–2.5% of the general population and 4.3–11% of asthmatic patients (Hedman et al. 1999; Szczeklik et al. 2000; Kasper et al. 2003; Vally et al. 2002). Since most of the agents that evoke AIA inhibit cyclo-oxygenase, a key enzyme in prostaglandin biosynthesis, genes involved in the arachidonic acid cascade may be responsible for the development of AIA. To date, leukotriene C4 synthetase has been reported to be associated with AIA (Sanak et al. 1997; Kawagishi et al. 2002). However, conflicting results have also been reported (Van Sambeek et al. 2000). Recently, by screening of 63 candidate genes of the arachidonic acid cascade in the Japanese population, we demonstrated that promoter polymorphism of the prostaglandin E2 receptor subtype 2 gene (EP2) is associated with AIA (Jinnai et al. 2004). Nevertheless, a single genetic factor cannot explain the genetic background of AIA, and, therefore, other factors conferring susceptibility to AIA remain to be identified.
It is unclear whether the synthesis or the functional activity of ADAM33 is enhanced or decreased by theses polymorphisms. If biological activity is enhanced, shedding of cytokine receptors, which is the putative result of proteolytic activity of ADAM33, might be excessive. In contrast, if the biological activity of ADAM33 is decreased, the presence of certain receptors might be increased. For example, the number of cells with CysLTR1 in the nasal mucosa was higher in AIA patients than in ATA patients (Sousa et al. 2002). If ADAM33 regulates the expression of CysLTR1, a decrease in its shedding activity would cause excess CysLTR1, leading to the development of AIA. Although this is largely speculative, there seems to be a link between ADAM33 and AIA. Recently, aspirin-exacerbated respiratory disease (AERD) has been associated with increased asthma severity (Mascia et al. 2005). Because in previous studies ADAM33 seemed to predispose to airway remodeling or increase in BA severity, it may be inferred that ADAM33 is indirectly linked with AIA.
Although several types of asthma or other lung abnormalities, such as childhood asthma (Van Eerdewegh et al. 2002; Lind et al. 2003; Raby et al. 2004; Noguchi et al. 2006), adult asthma (Hirota et al. 2006), reduced lung function (Jongepier et al. 2004; Van Diemen et al. 2005) and chronic obstructive pulmonary disease (COPD) (Van Diemen et al. 2005), have been reported to be associated with polymorphisms of ADAM33, there have been no studies on its association with aspirin sensitivity. The purpose of this investigation was to test the hypothesis that sequence variations in ADAM33 could contribute to the development of aspirin- and NSAIDs-intolerant asthma. We performed a genotype–phenotype association study, using polymorphisms of ADAM33, that was originally reported to be associated with asthma.
Materials and methods
Population subjects
This study was performed with the approval of the Ethics Committees at the School of Medicine, Niigata University and the Institute of Medical Science at the University of Tokyo. A total of 384 Japanese asthmatic patients was recruited at the Niigata University Hospital in Japan. Characteristics of the study population are shown in Table 1. BA was diagnosed according to the criteria of the Japanese Society of Allergology (1995). It was based on episodes of wheezing and dyspnea, especially at night and in the early morning, and reversible airway obstruction shown by >20% variation in daily values of volume in the first second of forced expiration (FEV1) and peak expiratory flow (PEF) after inhalation or administration of bronchodilator. Atopic-type asthma was identified in patient with elevated levels of serum total non-specific IgE. One hundred and twenty unrelated Japanese non-asthmatic individuals, over 60 years of age and with no history of asthma, comprised the control (CTR). Latent asthma is unlikely in this age group.
Asthma severity was also classified according to the criteria of the Japanese Society of Allergology (1995). This classification is based on a combination of the severity and frequency of asthma exacerbation and the therapeutic regimen required to control daily symptoms. A person with sporadic wheezing and mild episodes of asthma treated by bronchodilators alone is assigned to the mild asthma group. An individual with frequent moderate or severe asthma that restrict his/her activities of daily life is assigned to the severe asthma group. In persons receiving corticosteroids, total corticosteroid dosage, including inhalations, determines severity, independent of symptoms. An individual receiving more than 10 mg of prednisolone daily is defined as having severe asthma. The moderate severity group is defined by characteristics intermediate between the previous two classifications.
The diagnosis of AIA was made on the basis of a history of more than one self-reported episode of bronchial response to aspirin or NSAID ingestion. ATA was defined as BA with no history of NSAID-induced asthma. A provocation test was not applied, mainly because of the risk of significant occult disease, although the likelihood of severe reaction was considered very low (Jenkins et al. 2004).
Determination of the ADAM33 genotype
Genomic DNA was obtained from blood specimens from subjects who gave their written informed consent for DNA analysis. Asthmatic subjects were classified into AIA (n=102) and ATA (n=282), or into severe (n=72), moderate (n=257), or mild (n=55) asthma. Ten polymorphisms in the 3′ portion of the gene, ST+4, ST+7, T1, T2, T+1, V-3, V-2, V-1, V4, and V5, were selected for the association study, because most SNPs that were associated with BA and BHR in the original report (Van Eerdewegh et al. 2002) were in this region. Polymorphisms were genotyped using BigDye Terminator cycle sequencing on an ABI PRISM 3700 DNA analyzer, (Applied Biosystems, Tokyo, Japan). We imported the chromatograms into Sequencher software, version 3.1 (Hitachi Software, Yokohama, Japan) to determine genotypes.
Statistical analysis
For each polymorphism, Hardy–Weinberg equilibrium was tested with the χ2 test. We also performed χ2 tests to compare differences in allele frequency distribution between each phenotype. Haplotype frequencies for multiple sites in phase-unknown samples were estimated, using the expectation–maximization method, and the permutation test was performed to test the deviation of the haplotype frequency with SNPAlyze v5.0 software (DYNACOM, Mobara, Japan), as described elsewhere (Zhao et al. 2000; Jinnai et al. 2004).
Lewontin’s pair-wise linkage disequilibrium (LD) coefficient, D′, and a standardized LD coefficient, r 2, were calculated as described elsewhere (Lewontin 1964; Hill and Robertson 1968).
Results
Allele frequency distribution of ADAM33 polymorphisms
Frequencies of the minor alleles of ADAM33 in the AIA, ATA, and CTR groups are shown in Table 2. The genotype distributions fulfilled the Hardy–Weinberg equilibrium in all polymorphisms and groups. At the ST+7, V-1, and V5 sites, the minor allele frequency distributions of the AIA group were significantly different from those of the ATA group (χ2=5.49–9.35, P=0.034–0.004) and the differences between the AIA group and the CTR group were more prominent (χ2=4.49–8.25, P=0.019–0.002). In contrast, no statistically significant differences in allele frequency distribution between the ATA and CTR groups were observed.
The asthmatic patients were divided into severe and moderate–mild groups, based on clinical status as previously described, and the difference in allelic distributions between the severe and CTR groups and between the severe and moderate–mild groups were evaluated (Table 3). There was a significant difference in allele frequencies between the severe and moderate–mild groups at the ST+7 site (χ2=3.98, P=0.046). At the ST+7, V-1 and V5 sites the allele frequency distribution of the severe group was significantly different from that of the CTR group (χ2=4.01–6.06, P=0.045–0.014).
Linkage disequilibrium in ADAM33
For the estimation of the haplotype block structure of ADAM33, the linkage disequilibrium (LD) coefficients D′ and r 2 were calculated for every pair-wise combination of polymorphisms. Estimates of LD by the D′ and r 2 statistics were obtained from the observed genotype data for all possible pairs of the ten polymorphic sites in all subjects. The pair-wise LD among the ten polymorphisms also allowed us to identify the LD block within ADAM33 (Fig. 1), indicating that there is less historical recombination in the Japanese population. In contrast, the extent of LD estimated with r 2 statistics was very low throughout the gene, with the exception of several pairs, including V-2 and V-3, V5, V-1 and ST+7, and T+1, T2, and T1.
Linkage disequilibrium structure in ADAM33 Pair-wise LD between all pairs of SNPs in ADAM33 was evaluated by D′ and r 2 statistics. Pair-wise LD was determined in all subjects, for a total of 1,014 chromosomes, and is described as a block structure. In the schematic block, pairs of LD (0.8<D′, r 2) are shown as black boxes, LD (0.6<D′, r 2<0.8) as dark gray boxes, and LD (0.5<D′, r 2<0.6) as light gray boxes
Haplotype-based association of ADAM33
SNP, ST+7, V-1 and V5 showed significant associations with the AIA phenotype and were tight in LD (D′=0.91–0.99) and were, therefore, used for haplotype construction. Four major haplotypes, frequencies over 1%, were selected for evaluation in a haplotype-based association study, and the distributions of these frequencies are shown in Table 4. The haplotype-based association was tested with 10,000 iterated permutations. Frequency of the haplotype G/C/A at the ST+7/V-1/V5 sites of AIA showed significantly higher frequency than that of CTR and ATA (χ2=8.25, df=1, P=0.004; permutation P=0.001 and χ2=6.44, df=1, P=0.011; permutation P=0.008, respectively). A second haplotype, A/A/G, an opposite of haplotype G/C/A, also showed significant differences between AIA and ATA and between AIA and CTR. Accordingly, the G/C/A haplotype was regarded as an at-risk haplotype for AIA (Table 4).
Discussion
The work by Van Eerdewegh et al. (2002) that identified ADAM33 as an asthma-susceptibility gene, using positional cloning and an association study, had a profound impact on researchers investigating the genetics of bronchial asthma. Polymorphisms of ADAM33 were originally shown to be associated with BHR. In the present association study, we focused on AIA patients who tend to show severe BHR and found, at the ST+7, V-1 and V5 sites, significant associations with AIA. Correction for multiple tests (such as Bonferroni’s correction) was not performed in this study, because the investigated SNPs were tight in LD in our sample population and so were not independent.
In previous studies the V-1 SNP of ADAM33 was reported to be associated with disease in a mixed UK/USA population and in a UK-only population (Van Eerdewegh et al. 2002), and in a Hispanic population (Howard et al. 2003), and the ST+7 SNP with asthma in UK/USA (Van Eerdewegh et al. 2002), Dutch white (Howard et al. 2003), and German (Werner et al. 2004) populations. Recently, significant associations, at the S+1, ST+4 and T2 sites and at the T1, T2, S2 and V-3 sites, were reported in pediatric and adult Japanese populations, respectively (Noguchi et al. 2006; Hirota et al. 2006), but these associations were not found in our sample. In contrast, no association was found between asthma and ADAM33 in Puerto Rican and Mexican populations (Lind et al. 2003), the Korean population (Lee et al. 2004), or a North American pediatric population (Raby et al. 2004). Comparison of our results with those studies indicates that no single SNP was universally associated with the asthma phenotype. One explanation for these differences might be the heterogeneity of asthma. There seem to be genetic differences among subtypes of asthma, such as pediatric asthma, adult-onset asthma, AIA, and so forth. BA was not defined by common criteria in these investigations, and, in our study, the emphasis was on aspirin sensitivity. Further, there are structural differences in the LD pattern among populations. For example, there are large differences in LD pattern and haplotype structure among populations in the genomic interleukin 4 and interleukin 13 regions (Sakagami et al. 2004) and in the angiotensinogen region (Nakajima et al. 2004). Likewise, if there is an undetected causal polymorphism in the vicinity of the ADAM33 region, each study could detect associations with different polymorphisms due to ethnically diverse LD patterns of ADAM33. Although we cannot exclude the possibility that one of the three SNPs ST+7, V-1, or V5 may be related to a functional variation (affecting, for example, transcriptional activity and/or stability of RNA) leading to the AIA phenotype, it is more likely that the underlying cause lies in the haplotype. And the other, allelic heterogeneity, by which different mutations in the same gene combine in a single phenotype, might be responsible for the discrepancies among populations.
The severity of asthma is known to correlate with PC20 (Murray et al. 1981); severely asthmatic patients show severe BHR. So our data confirm previous reports that ADAM33 is a gene particularly associated with BHR (Van Eerdewegh et al. 2002). Accordingly, we classified our asthmatic patients on the basis of clinical severity and analyzed the allelic associations among the groups. At the same three sites, significant differences in allelic frequencies between severe asthma and CTR were observed. In terms of clinical manifestations of AIA, it has been reported that aspirin sensitivity is associated with increased asthmatic severity and airway remodeling. The TENOR study group reported that aspirin-exacerbated respiratory disease (AERD) was associated with increased asthma severity and fixed airway obstruction, which was considered a manifestation of airway remodeling, in an epidemiologic cohort study (Mascia et al. 2005). It was concluded that ADAM33 causes increasing asthma severity and progressive airway remodeling and, consequently, contribute to the pathogenesis of AIA.
ADAM33, an asthma-associated gene, was studied by analysis of genotype–phenotype associations in Japanese asthmatic patients. Polymorphisms of ADAM33 were associated with AIA and severe forms of asthma. Our results should contribute to functional studies of ADAM33 as well as furthering understanding of the pathophysiology of AIA.
References
Asakura M, Kitakaze M, Takashima S, Liao Y, Ishikura F, Yoshinaka T, Ohmoto H, Node K, Yoshino K, Ishiguro H, Asanuma H, Sanada S, Matsumura Y, Takeda H, Beppu S, Tada M, Hori M, Higashiyama S (2002) Cardiac hypertrophy is inhibited by antagonism of ADAM12 processing of HB-EGF: metalloproteinase inhibitors as a new therapy. Nat Med 8:35–40
Black RA, Rauch CT, Kozlosky CJ, Peschon JJ, Slack JL, Wolfson MF, Castner BJ, Stocking KL, Reddy P, Srinivasan S, Nelson N, Boiani N, Schooley KA, Gerhart M, Davis R, Fitzner JN, Johnson RS, Paxton RJ, March CJ, Cerretti DP (1997) A metalloproteinase disintegrin that releases tumor-necrosis factor-alpha from cells. Nature 385:729–733
Garton KJ, Gough PJ, Blobel CP, Murphy G, Greaves DR, Dempsey PJ, Raines EW (2001) Tumor necrosis factor-alpha-converting enzyme (ADAM17) mediates the cleavage and shedding of fractalkine (CX3CL1). J Biol Chem 276:37993–38001
Hedman J, Kaprio J, Poussa T, Nieminen MM (1999) Prevalence of asthma, aspirin intolerance, nasal polyposis and chronic obstructive pulmonary disease in a population-based study. Int J Epidemiol 28:717–722
Hill WG, Robertson A (1968) Linkage disequilibrium infinite populations. Theor Appl Genet 38:226–231
Hirota T, Hasegawa K, Obara K, Matsuda A, Akahoshi M, Nakashima K, Shirakawa T, Doi S, Fujita K, Suzuki Y, Nakamura Y, Tamari M (2006) Association between ADAM33 polymorphisms and adult asthma in the Japanese population. Clin Exp Allergy 36:884–891
Howard TD, Postma DS, Jongepier H, Moore WC, Koppelman GH, Zheng SL, Xu J, Bleecker ER, Meyers DA (2003) Association of a disintegrin and metalloprotease 33 (ADAM33) gene with asthma in ethnically diverse populations. J Allergy Clin Immunol 112:717–722
Japanese Society of Allergology (1995) Guidelines for the diagnosis and management of bronchial asthma. Committee on the Definition, Treatment, and Management of Bronchial Asthma. Allergy 50 [27 Suppl]:1–42
Jenkins C, Costello J, Hodge L (2004) Systematic review of prevalence of aspirin induced asthma and its implications for clinical practice. BMJ 328:434
Jinnai N, Sakagami T, Sekigawa T, Kakihara M, Nakajima T, Yoshida K, Goto S, Hasegawa T, Koshino T, Hasegawa Y, Inoue H, Suzuki N, Sano Y, Inoue I (2004) Polymorphisms in the prostaglandin E2 receptor subtype 2 gene confer susceptibility to aspirin-intolerant asthma: a candidate gene approach. Hum Mol Genet 13:3203–3217
Jongepier H, Boezen HM, Dijkstra A, Howard TD, Vonk JM, Koppelman GH, Zheng SL, Meyers DA, Bleecker ER, Postma DS (2004) Polymorphisms of the ADAM33 gene are associated with accelerated lung function decline in asthma. Clin Exp Allergy 34:757–760
Kasper L, Sladek K, Duplaga M, Bochenek G, Liebhart J, Gladysz U, Malolepszy J, Szczeklik A. (2003) Prevalence of asthma with aspirin hypersensitivity in the adult population of Poland. Allergy 58:1064–1066
Kawagishi Y, Mita H, Taniguchi M, Maruyama M, Oosaki R, Higashi N, Kashii T, Kobayashi M, Akiyama K. (2002) Leukotriene C4 synthase promoter polymorphism in Japanese patients with aspirin-induced asthma. J Allergy Clin Immunol 109:936–942
Lee JH, Park HS, Park SW, Jang AS, Uh ST, Rhim T, Park CS, Hong SJ, Holgate ST, Holloway JW, Shin HD (2004) ADAM33 polymorphism: association with bronchial hyper-responsiveness in Korean asthmatics. Clin Exp Allergy 34:860–865
Lewontin RC (1964) The interaction of selection and linkage. 1. General considerations, heterotic models. Genetics 49:49–67
Lind DL, Choudhry S, Ung N, Ziv E, Avila PC, Salari K, Ha C, Lovins EG, Coyle NE, Nazario S, Casal J, Torres A, Rodriguez-Santana JR, Matallana H, Lilly CM, Salas J, Selman M, Boushey HA, Weiss ST, Chapela R, Ford JG, Rodriguez-Cintron W, Silverman EK, Sheppard D, Kwok PY, Gonzalez Burchard E (2003) ADAM33 is not associated with asthma in Puerto Rican or Mexican populations. Am J Respir Crit Care Med 168:1312–1316
Mascia K, Haselkorn T, Deniz YM, Miller DP, Bleecker ER, Borish L (2005) Aspirin sensitivity and severity of asthma: evidence for irreversible airway obstruction in patients with severe or difficult-to-treat asthma. J Allergy Clin Immunol 116:970–975
Murray AB, Ferguson AC, Morrison B (1981) Airway responsiveness to histamine as a test for overall severity of asthma in children. J Allergy Clin Immunol 68:119–124
Nakajima T, Wooding S, Sakagami T, Emi M, Tokunaga K, Tamiya G, Ishigami T, Umemura S, Munkhbat B, Jin F, Guan-Jun J, Hayasaka I, Ishida T, Saitou N, Pavelka K, Lalouel JM, Jorde LB, Inoue I (2004) Natural selection and population history in the human angiotensinogen gene (AGT): 736 complete AGT sequences in chromosomes from around the world. Am J Hum Genet 74:898–916
Noguchi E, Ohtsuki Y, Tokunaga K, Yamaoka-Sageshima M, Ichikawa K, Aoki T, Shibasaki M, Arinami T (2006) ADAM33 polymorphisms are associated with asthma susceptibility in a Japanese population. Clin Exp Allergy 36:602–608
Raby BA, Silverman EK, Kwiatkowski DJ, Lange C, Lazarus R, Weiss ST (2004) ADAM33 polymorphisms and phenotype associations in childhood asthma. J Allergy Clin Immunol 113:1071–1078
Sakagami T, Witherspoon DJ, Nakajima T, Jinnai N, Wooding S, Jorde LB, Hasegawa T, Suzuki E, Gejyo F, Inoue I (2004) Local adaptation and population differentiation at the interleukin 13 and interleukin 4 loci. Genes Immun 5:389–397
Sanak M, Simon HU, Szczeklik A (1997) Leukotriene C4 synthase promoter polymorphism and risk of aspirin-induced asthma. Lancet 350:1599–1600
Sousa AR, Parikh A, Scadding G, Corrigan CJ, Lee TH (2002) Leukotriene-receptor expression on nasal mucosal inflammatory cells in aspirin-sensitive rhinosinusitis. N Engl J Med 347:1493–1499
Szczeklik A, Nizankowska E, Duplaga M (2000) Natural history of aspirin-induced asthma. AIANE Investigators. European Network on Aspirin-Induced Asthma. Eur Respir J 16:432–436
Vally H, Taylor ML, Thompson PJ (2002) The prevalence of aspirin-intolerant asthma (AIA) in Australian asthmatic patients. Thorax 57:569–574
Van Diemen CC, Postma DS, Vonk JM, Bruinenberg M, Schouten JP, Boezen HM (2005) A disintegrin, metalloprotease 33 polymorphisms and lung function decline in the general population. Am J Respir Crit Care Med 172:329–333
Van Eerdewegh P, Little RD, Dupuis J, Del Mastro RG, Falls K, Simon J, Torrey D, Pandit S, McKenny J, Braunschweiger K, Walsh A, Liu Z, Hayward B, Folz C, Manning SP, Bawa A, Saracino L, Thackston M, Benchekroun Y, Capparell N, Wang M, Adair R, Feng Y, Dubois J, FitzGerald MG, Huang H, Gibson R, Allen KM, Pedan A, Danzig MR, Umland SP, Egan RW, Cuss FM, Rorke S, Clough JB, Holloway JW, Holgate ST, Keith TP (2002) Association of the ADAM33 gene with asthma and bronchial hyperresponsiveness. Nature 418:426–430
Van Sambeek R, Stevenson DD, Baldasaro M, Lam BK, Zhao J, Yoshida S, Yandora C, Drazen JM, Penrose JF (2000) 5′ Flanking region polymorphism of the gene encoding leukotriene C4 synthase does not correlate with the aspirin-intolerant asthma phenotype in the United States. J Allergy Clin Immunol 106:72–76
Werner M, Herbon N, Gohlke H, Altmuller J, Knapp M, Heinrich J, Wjst M (2004) Asthma is associated with single-nucleotide polymorphisms in ADAM33. Clin Exp Allergy 34:26–31
Yoshinaka T, Nishii K, Yamada K, Sawada H, Nishiwaki E, Smith K, Yoshino K, Ishiguro H, Higashiyama S (2002) Identification and characterization of novel mouse and human ADAM33s with potential metalloprotease activity. Gene 282:227–236
Zhao JH, Curtis D, Sham PC (2000) Model-free analysis and permutation tests for allelic associations. Hum Hered 50:133–139
Acknowledgment
This work was supported by the Future Program Grant of the Japan Society for the Promotion of Science (II). We thank all DNA donors and supporting medical staff for making this study possible, particularly M. Sasagawa and K. Kurashige (Nanbugo General Hospital), M. Haraguchi and Y. Kobayashi (Niigata Citizen Hospital), F. Yoshimine (Muikamachi Hospital), K. Fujimori (Shibata Hospital), T. Koya (Saiseikai Kawaguchi Hospital), A. Takagi (Toyosaka Hospital), K. Ito (Saiseikai Niigata Hospital) and K. Ikarashi. We are grateful to M. Kakihara, K. Eguchi, E. Nakamura, Y. Miwa, Y. Sakamoto, K. Otaka and Y. Terada for their technical assistance.
Author information
Authors and Affiliations
Corresponding author
Additional information
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Sakagami, T., Jinnai, N., Nakajima, T. et al. ADAM33 polymorphisms are associated with aspirin-intolerant asthma in the Japanese population. J Hum Genet 52, 66–72 (2007). https://doi.org/10.1007/s10038-006-0081-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-006-0081-6
Keywords
This article is cited by
-
Association analysis of ILVBL gene polymorphisms with aspirin-exacerbated respiratory disease in asthma
BMC Pulmonary Medicine (2017)
-
Association of ADAM33 gene polymorphisms with asthma in Indian children
Journal of Human Genetics (2011)
-
A possible association of EMID2 polymorphisms with aspirin hypersensitivity in asthma
Immunogenetics (2011)
-
Association of ADAM33 gene polymorphisms with adult-onset asthma and its severity in an Indian adult population
Journal of Genetics (2011)
-
Association of ADAM33 gene with susceptibility to COPD in Tibetan population of China
Molecular Biology Reports (2011)