Abstract
Comparative genomic hybridization (CGH) analyses have detected gains of copy number on 13q, especially at 13q31-q32, in cell lines and primary cases of various types of lymphoma. Since amplification of chromosomal DNA is one of the mechanisms that can activate tumor-associated genes, and because 13q amplification had been reported in various other types of tumors as well, we attempted to define by fluorescence in situ hybridization (FISH) a common region at 13q31-q32 in which to explore genes that might be targets for the amplification events. Although the commonly amplified region we defined was relatively large (approximately 4 Mb), only one true gene, GPC5, was found there. GPC5 was over-expressed in lymphoma cell lines that had shown amplification, in comparison with those that had not. Our findings suggest that GPC5 is a likely target for amplification, and that over-expression of this gene may contribute to development and/or progression of lymphomas and other tumors.
Similar content being viewed by others
Introduction
Gene amplification is frequently observed in human tumors, and appears to be one of the mechanisms leading to activation of proto-oncogenes (Stark et al. 1989). Defining and characterizing genes within amplified chromosomal regions represents an excellent route toward identification of elements involved in tumorigenesis.
Lymphoma remains incurable even with current therapeutic modalities. Multiple genetic alterations that have been identified in various types of lymphoma (Neat et al. 2001; Allen et al. 2002; Mao et al. 2002; Mehra et al. 2002) may be linked in a sequence of events that parallels malignant progression. The REL and BCL2 proto-oncogenes are known amplification targets that are associated with malignant progression of lymphomas (Houldsworth et al. 1996; Rao et al. 1998). However, those two genes do not fully account for all of the genetic material present in amplified regions or for the variety of malignant phenotypes that lymphomas can exhibit. Indeed, cumulative results of comparative genomic hybridization (CGH) analyses have revealed additional amplified regions in lymphomas that contain novel target genes that could be associated with pathogenesis of this disease. (Neat et al. 2001; Allen et al. 2002; Mao et al. 2002). Identification of those target genes should yield new insights into the molecular pathogenesis of lymphoma, establish diagnostic markers, and suggest novel therapeutic targets.
Among the chromosomal regions showing recurrent amplification in CGH analyses of various types of lymphoma, 13q31-q32 is of particular interest because (1) gains of copy number are frequent in that region; e.g., in eight (18%) of 45 cases of follicular lymphoma (Neat et al. 2001) and four (29%) of 14 cases of primary cutaneous B-cell lymphoma (Mao et al. 2002); and (2) genomic alterations within or around this region have also been implicated in other malignancies such as small and non-small cell lung cancers, neuroglial tumors, and squamous cell carcinoma of the head and neck (Knuutila et al. 1998). Therefore, the 13q31-q32 region may harbor one or more genes which, when activated by amplification, might be associated with progression and/or specific phenotypes of lymphomas and other tumors. However, no actual or potential targets of the amplification at 13q31-q32 have yet been clarified. These observations prompted us to perform a detailed molecular characterization of the 13q31-q32 amplicon using lymphoma cell lines, with the goal of identifying gene(s) involved in the pathogenesis of this type of tumor.
Materials and methods
Cell lines
A total of six lymphoma cell lines, KARPAS1718 (splenic lymphoma with circulating villous lymphocyte), RAMOS (Burkitt lymphoma), AL-1 (Burkitt lymphoma), B46M (Burkitt lymphoma), OCI-ly1 (non-Hodgkin lymphoma), and OCI-ly3 (non-Hodgkin lymphoma) were employed in the present study. These cell lines were maintained in RPMI-1640 supplemented with 10% fetal calf serum and penicillin-streptomycin. Metaphase chromosome slides were prepared and used in FISH experiments in the manner described previously (Fukuda et al. 2000).
Fluorescence in situ hybridization (FISH)
We performed FISH analysis using 25 bacterial artificial chromosomes (BACs, 17 are listed in Fig. 1b), as described previously (Inazawa et al. 1993; Ariyama et al. 1999). The locations of BACs within the region of interest were compiled from information archived by the University of California at Santa Cruz (UCSC; http://genome.ucsc.edu/) or the National Center for Biotechnology Information (NCBI; http://www.ncbi.nlm.nih.gov/). Relative positions of the selected BACs on a map of the 13q31-q32 region are indicated in Fig. 1b. Probes were labeled by nick-translation with biotin-16-dUTP or digoxigenin (Roche Diagnostics, Tokyo, Japan) and hybridized to metaphase chromosomes. Precise localization of each BAC was confirmed using normal metaphase chromosomes.
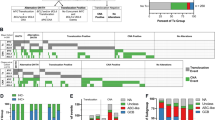
a Representative images of FISH analysis on metaphase chromosome from KARPAS1718, OCI-LY3 and AL-1 with BAC clone 487A02, containing the GPC5 gene, as a probe. KARPAS1718 shows remarkable increase in signals with HSR pattern and OCI-LY3 shows four signals, whereas AL-1 shows two signals (no amplification). b The amplicon map of the 13q31-q32 region between two makers, RH53181 and D13S197. The position of each STS and each of the 17 BACs was compiled from information archived by the UCSC and the NCBI (left). This region contains the only known gene, GPC5 (middle). Summarized results of DNA sequence copy-number analysis by FISH in four lymphoma cell lines showing amplification of this region (right). The horizontal axis shows the number of FISH signals achieved with the BAC probes indicated above. The number of signals was truncated at 15 because it was difficult to enumerate them above this level. The smallest region of overlap (SRO) with maximal amplification is indicated by the closed arrow. c Relative expression of GPC5 in six lymphoma cell lines, four with increased copy number of GPC5 and two with normal copy number of GPC5. The highest expression is found in the KARPAS1718 cell line exhibiting remarkable copy number gain of this gene (a, b)
Reverse transcriptase-polymerase chain reaction (RT-PCR), direct sequence, Northern blot, and database search
We performed reverse transcriptase-polymerase chain reaction (RT-PCR) using at least two primer sets for each of 15 uncharacterized, predicated transcripts within the smallest region of overlap within 13q31-q32 that we defined. Primer sequences are available on request. Single-stranded cDNAs were generated from total RNAs of cell lines using the SuperScript First-Strand Synthesis System (Invitrogen, Carlsbad, Calif.) according to the manufacturer's directions. RT-PCR was performed using FastStart Taq DNA polymerase (Roche Diagnostics, Tokyo, Japan) according to the manufacturer's directions. Every RT-PCR product was evaluated by electrophoresis, direct sequence using ABI377 Autosequencer, and/or Northern blotting (Human 12-lane MTN, Clontech, Palo Alto, Calif.) described elsewhere (Imoto et al. 2001). All 15 uncharacterized, predicated transcripts were evaluated using NCBI database, such as LocusLink with Evidence Viewer (http://www.ncbi.nlm.nih.gov/LocusLink) and BLAST (http://www.ncbi.nlm.nih.gov/BLAST/).
Real-time quantitative RT-PCR
The quantification of GPC5 mRNA was carried out using a real-time fluorescence detection method (Wittwer et al.1997). Single-stranded cDNAs were generated from total RNAs of each cell line. Real-time quantitative PCR was performed using LightCycler (Roche Diagnostics) with CYBR Green according to the manufacturer's protocol. The primer sequences for GPC5 gene were as followings: the forward primer, 5′-GGTCGTTGGAAGAACTCTCG-3′ and the reverse primer, 5′-GCGGCCACAAATCCTATTTA-3′. The glyceraldehyde-3-phosphate dehydrogenase gene (GAPDH; Roche Diagnostics) served as an endogenous control and each sample was normalized on the basis of its GAPDH content.
Results and discussion
Definition of the 13q31-32 amplicon by FISH
As the first step toward exploring candidate gene(s) we constructed a map of the 13q31-q32 amplicon by FISH, using available lymphoma cell lines, to define a relatively small chromosomal region that would make a positional search possible. The criterion we used to define the amplicon was based on the premise that the best candidates (i.e., targets that are likely to be selected during the amplification process) would be located within the smallest region of overlap among regions showing the highest copy numbers in a panel of cell lines or primary tumors (Albertson et al. 2000).
We first performed FISH analyses using ten BACs located around 13q31-q32, covering approximately 10 Mb, in the six lymphoma cell lines; four of those lines (KARPAS1718, RAMOS, OCI-ly1, and OCI-ly3) had shown copy-number gains of chromosome 13q in G-banding experiments (Mehra et al. 2002) or in CGH studies (Mehra et al. 2002; Dr. Masao Seto in Aichi Cancer Center Research Institute, personal communication) and we found that two BACs, 86C3 and 302C13, showed a consistent increase in the number of signals in these four cell lines (data not shown).
Then we performed FISH analyses using additional 15 BACs located around 86C3 and 302C13 (Fig. 1b). In KARPAS1718 cells, 12 of the 17 BACs used in the second round of FISH experiments (321E11, 275J18, 114G01, 86C3, 388D04, 392A19, 430K10, 487A0 2,95C14, 302C13, 66G15 and 266K03) produced strong signals as homogeneous staining regions (HSRs) (Fig. 1a, b), indicating that they each represented part of the amplicon. In contrast, the other five BACs failed to show HSR patterns in the same cell line, reflecting positions outside the amplicon (Fig. 1b). In RAMOS cells, 15 BACs produced five signals and two BACs each produced two signals (Fig. 1b). In OCI-LY3 and OCI-LY1, all 17 BACs produced four and three signals, respectively (Fig. 1a, b). Taking these findings together, we defined the smallest region of overlap (SRO) between BACs 321E11 and 266K03. We estimated its extent to be 4 Mb, on the basis of the genome database archived by the UCSC. According to the NCBI database, this region harbors the GPC5 gene and 15 uncharacterized, possibly transcribed sequences (Table 1).
Expression of GPC5 gene in lymphoma cell lines
As Table 1 indicates, the sequences and genomic structures of the database-predicted transcripts, as well as the RT-PCR results followed by direct sequencing and northern blotting (data not shown), demonstrated that none of the 16 predicted transcripts except GPC5 were true genes. For example, no expressed-sequence tags (ESTs) or evidence of exons had been filed in genome databases, and no intronic sequences were detected, indicating that the predicted transcripts were probably pseudogenes; moreover, no specific RT-PCR products with correct sequences were detected for any of the 15 predicted sequences, even though we used more than two primer sets for each of them (data not shown). On the basis of the database-derived and experimental evidence, we concluded that none of the 15 predicted sequences was a true gene, and that the 4-Mb SRO contained only one actual gene, GPC5. Therefore we focused on GPC5 as a potential target for amplification at 13q31-q32.
As a second step toward determining whether GPC5 was in fact the target for amplification in our SRO, we evaluated whether this gene was consistently over-expressed as a consequence of gains in copy number (Imoto et al. 2001). All six lymphoma cell lines showed detectable expression of GPC5, although the relative expression levels in cell lines with increased copy numbers of GPC5 were consistently higher than in the lines with only two copies (Fig. 1c). Those data suggested that this gene might indeed be activated by the amplification mechanism and represent an actual target of amplification in lymphoma cell lines. However, GPC5 expression levels did not always parallel the copy numbers, indicating that other genetic and/or epigenetic mechanisms may contribute to increased expression of this gene.
GPC5 is a member of the glypican family, a group of membrane-bound heparan sulfate proteoglycans (HSPGs) that are linked to the cell surface by a glycosylphosphatidylinositol (GPI) anchor (Filmus and Song 2000). Six members of this molecular family (GPC1–GPC6) have been identified in mammals (Filmus et al. 1988; David et al. 1990; Stipp et al. 1994; Watanabe et al. 1995; Veugelers et al. 1997; Saunders et al. 1997; Veugelers et al. 1999; Paine-Saunders et al. 1999). In general, glypicans are expressed predominantly during development and their expression levels change in a stage- and tissue-specific manner, suggesting that glypicans are involved in the regulation of morphogenesis (Litwack et al. 1994; Saunders et al. 1997; Pellegrini et al. 1998). Since glypicans can regulate the activity of growth and survival factors, changes in their expression could very well be associated with tumor progression. For example, GPC1 expression was significantly increased in a large proportion of pancreatic cancers (Kleef et al. 1998). GPC3 expression was down-regulated in a significant proportion of ovarian-cancer cell lines (Lin et al. 1999) and in mesothelioma cell lines (Murthy et al. 2000), but up-regulated in most hepatocellular carcinomas (Hsu et al. 1997) and a significant proportion of colorectal tumors (Filmus and Selleck 2001).
As GPC5 is expressed in various embryonic tissues but only in brain tissue in the adult, GPC5 expression may have a general role in the control of growth and differentiation during mammalian development (Veugelers et al. 1997). In Northern blotting experiments most cultured tumor-cell lines, including acute lymphoblastic leukemia, histiocytic lymphoma, and myeloma, have been negative for GPC5 (Veugelers et al. 1997). However, the expression patterns reported here, as well as functions of other members of the glypican family, strongly suggest that altered expression of GPC5, especially over-expression through gene amplification, might play an important role in the pathogenesis of lymphomas. Further studies will be necessary to evaluate the amplification and expression status of this gene in primary cases of lymphoma and to determine the functional roles of GPC5 in the pathogenesis of these tumors.
References
Albertson DG, Ylstra B, Segraves R, Collins C, Dairkee SH, Kowbel D, Kuo WL, Gray JW, Pinkel D (2000) Quantitative mapping of amplicon structure by array CGH identifies CYP24 as a candidate oncogene. Nat Genet 25:144–146
Allen JE, Hough RE, Goepel JR, Bottomley S, Wilson GA, Alcock HE, Baird M, Lorigan PC, Vandenberghe EA, Hancock BW, Hammond, DW (2002) Identification of novel regions of amplification and deletion within mantle cell lymphoma DNA by comparative genomic hybridization. Br J Haematol 116:291–298
Ariyama Y, Mori T, Shinomiya T, Sakabe T, Fukuda Y, Kanamaru A, Yamada Y, Isobe M, Seto M, Nakamura Y, Inazawa J (1999) Chromosomal imbalances in adult T-cell leukemia revealed by comparative genomic hybridization: gains at 14q32 and 2p16-22 in cell lines. J Hum Genet 44:357–363
David G, Lories V, Decock B, Marynen P, Cassiman JJ, Van den Berghe H (1990) Molecular cloning of a phosphatidylinositol-anchored membrane heparan sulfate proteoglycan from human lung fibroblasts. J Cell Biol 111:3165–3176
Filmus J, Church J, Buick RN (1988) Isolation of a cDNA corresponding to a developmentally regulated transcript in rat intestine. Mol Cell Biol 8:4243–4249
Filmus J, Selleck SB (2001) Glypicans: proteoglycans with a surprise. J Clin Invest 108:497–501
Filmus J, Song HH (2000) Glypicans. In: Iozzo RV (ed) Proteoglycans. Marcel Dekker, New York, USA, pp 161–176
Fukuda Y, Kurihara N, Imoto I, Yasui K, Yoshida M, Yanagihara K, Park JG, Nakamura Y, Inazawa J (2000) CD44 is a potential target of amplification within the 11p13 amplicon detected in gastric cancer cell lines. Genes Chromosomes Cancer 29:315–324
Houldsworth J, Mathew S, Rao PH, Dyomina K, Louie DC, Parsa N, Offit K, Chaganti RS (1996) REL proto-oncogene is frequently amplified in extranodal diffuse large cell lymphoma. Blood 87:25–29
Hsu HC, Cheng W, Lai PL (1997) Cloning and expression of a developmentally regulated transcript MXR7 in hepatocellular carcinoma: biological significance and temporospatial distribution. Cancer Res 57:5179–5184
Imoto I, Yang ZQ, Pimkhaokham A, Tsuda H, Shimada Y, Imamura M, Ohki M, Inazawa J (2001) Identification of cIAPI as a candidate target gene within an amplicon at 11q22 in esophageal squamous cell carcinomas. Cancer Res 61:6629–6634
Inazawa J, Saito H, Ariyama T, Abe T, Nakamura Y (1993) High-resolution cytogenetic mapping of 342 new cosmid markers including 43 RFLP markers on human chromosome 17 by fluorescence in situ hybridization. Genomics 17:153–162
Kleeff J, Ishiwata T, Kumbasar A, Friess H, Büchler MW, Lander AD, Korc M (1998) The cell surface heparan sulfate proteoglycan glypican-1 regulates growth factor in pancreatic carcinoma cells and is over-expressed in human pancreatic cancer. J Clin Invest 102:1662–1673
Knuutila S, Bjorkqvist AM, Autio K, Tarkkanen M, Wolf M, Monni O, Szymanska J, Larramendy ML, Tapper J, Pere H, El-Rifai W, Hemmer S, Wasenius VM, Vidgren V, Zhu Y (1998) DNA copy number amplifications in human neoplasms: review of comparative genomic hybridization studies. Am J Pathol 152:1107–1123
Lin H, Huber R, Schlessinger D, Morin PJ (1999) Frequent silencing of the GPC3 gene in ovarian cancer cell lines. Cancer Res 59:807–810
Litwack ED, Stipp CS, Kumbasar A, Lander AD (1994) Neuronal expression of glypican, a cell surface glycosylphosphatidylinositol-anchored heparan sulfate proteoglycan, in the adult rat nervous system. J Neurosci 14:3713–3724
Mao X, Lillington D, Child F, Russell-Jones R, Whittaker S (2002) Comparative genomic hybridization analysis of primary cutaneous B-cell lymphomas: identification of common genomic alterations in disease pathogenesis. Genes Chromosomes Cancer 35:144–155
Mehra S, Messner H, Minden M, Chaganti RSK (2002) Molecular cytogenetic characterization of non-Hodgkin lymphoma cell lines. Genes Chromosomes Cancer 33:225–234
Murthy SS, Shen T, Rienzo AD, Lee WC, Ferriola PC, Jhanwar SC, Mossman BT, Filmus J, Testa JR (2000) Expression of GPC3, an X-linked recessive overgrowth gene, is silenced in malignant mesothelioma. Oncogene 19:410–416
Neat MJ, Foot N, Jenner M, Goff L, Ashcroft K, Burford DH, Dunham A, Norton AT, Lister A, Fitzgibbon J (2001) Localisation of a novel region of recurrent amplification in follicular lymphoma to an ~6.8 Mb region of 13q32–33. Genes Chromosomes Cancer 32:236–243
Paine-Saunders S, Viviano BL, Saunders S (1999) GPC6, a novel member of the glypican gene family, encodes a product structurally related to GPC4 and is colocalized with GPC5 on human chromosome 13. Genomics 57:455–458
Pellegrini M, Pilia G, Pantano S, Lucchini F, Uda M, Fumi M, Cao A, Schlessinger D, Forabosco A (1998) GPC3 expression correlates with the phenotype of the Simpson-Golabi-Behmel syndrome. Dev Dyn 213:431–439
Rao PH, Houldsworth J, Dyomina K, Parsa NZ, Cigudosa JC, Louie DC, Popplewell L, Offit K, Jhanwar SC, and Chaganti RSK (1998) Chromosomal and gene amplification in diffuse large B-cell lymphoma. Blood 92:234–240
Saunders S, Paine-Saunders S, Lander AD (1997) Expression of the cell surface proteoglycan glypican-5 is developmentally regulated in kidney, limb, and brain. Dev Biol 190:78–93
Stark GR, Debatisse M, Giulott E, Wahl GM (1989) Recent progress in understanding mechanisms of mammalian DNA amplification. Cell 57:901–908
Stipp CS, Litwac ED, Lander AD (1994) Cerebroglycan: an integral membrane sulfate proteoglycan that is unique to the developing nervous system and expressed specifically during neuronal differentiation. J Cell Biol 124:149–160
Veugelers M, Vermeesch J, Reekmans G, Steinfeld R, Marynen P, David G (1997) Characterization of glypican-5 and chromosomal localization of human GPC5, a new member of the glypican gene family. Genomics 40:24–30
Veugelers M, De Cat B, Ceulemans H, Bruystens AM, Coomans C, Durr J, Vermeesch J, Marynen P, David G (1999) Glypican-6, a new member of the glypican family of cell surface proteoglycans. J Biol Chem 274:26968–26977
Watanabe K, Yamada H, Yamaguchi Y (1995) K-glypican: a novel GPI-linked heparan sulfate proteoglycan that is highly expressed in developing brain and kidney. J Cell Biol 130:1207–1218
Wittwer CT, Ririe KM, Andrew RV, David DA, Gundry RA, Balis UJ (1997) The Lightcycler: a microvolume multisample fluorimeter with rapid temperature control. Biotechniques 22:176–181
Acknowledgements
We thank Dr. Masao Seto (Aichi Cancer Center) for giving us information of CGH data in lymphoma cell lines used in the present study, and also thank Professor Yusuke Nakamura (Institute of Medical Science, The University of Tokyo) for his continuous encouragement. This work was supported by Grants-in-Aid for Scientific Research on Priority Areas (C) from the Japanese Ministry of Education, Culture, Sports, Science, and Technology, and by Core Research for Evolutional Science and Technology (CREST) of the Japan Science and Technology Corporation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yu, W., Inoue, J., Imoto, I. et al. GPC5 is a possible target for the 13q31-q32 amplification detected in lymphoma cell lines. J Hum Genet 48, 331–335 (2003). https://doi.org/10.1007/s10038-003-0026-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-003-0026-2
Keywords
This article is cited by
-
Blood vessel remodeling in the cerebral cortex induced by binge alcohol intake in mice
Toxicological Research (2023)
-
Prognostic value of Glypican family genes in early-stage pancreatic ductal adenocarcinoma after pancreaticoduodenectomy and possible mechanisms
BMC Gastroenterology (2020)
-
The Role of Cell Growth-Related Gene Copy Number Variation in Autoimmune Thyroid Disease
Biological Trace Element Research (2020)
-
Antithetic roles of proteoglycans in cancer
Cellular and Molecular Life Sciences (2012)
-
Genomic alterations in rectal tumors and response to neoadjuvant chemoradiotherapy: an exploratory study
Radiation Oncology (2011)