Abstract
Two new 4’-acetylated analogs of chrysomycin were discovered during the screening for antitumor agents from the metabolites of actinomycetes. Their structures and physicochemical properties were determined by standard spectrometric analyses. Their cytotoxicities and antimicrobial activities were evaluated against a panel of cancer cell lines and microbes. While acetylation reinforced the cytotoxicity of chrysomycin B, it weakened the activity of chrysomycin A. Chrysomycin A and its acetylated analog showed high cytotoxicity toward most of the cancer cells with IC50s less than 10 ng ml−1. The 4’-acetyl-chrysomycin A was predominantly observed in nuclei at concentrations where the autofluorescence was observable. Chrysomycins were effective toward Gram-positive bacteria. The 4’-acetylated-chrysomycin A and B had MICs of 0.5–2 μg ml−1 and 2 to greater than 64 μg ml−1, respectively, toward Gram-positive bacteria including MRSA and VRE.
Similar content being viewed by others
Main
Chrysomycins are benzonaphthopyranone glycosides found in Streptomyces spp.1 Analogous compounds possessing the same chromophore, such as gilvocarcins, ravidomycins and polycarcins, are also known.2, 3, 4, 5 This group of compounds has strong antitumor properties and mild toxicities toward mouse. The antitumor activities of chrysomycins were evaluated in mice up to 256–512 mg kg−1 (i.p.) and elongation of survival times (~200%) were reported.6 The LD50 values of gilvocarcins were >300 mg kg−1 (i.v.) and >1000 mg kg−1 (p.o. and i.p.) and they showed similar effects on the elongation of the survival time of tumor-burdened mice.7, 8 Their inhibition of topoisomerase II and cross-linking of histone H3 and heat-shock protein GRP78 to DNA may be involved in the antitumor activity.9, 10 The related compound elsamitrucin was evaluated in several cancers in phase II studies.11, 12 Elsamitrucin showed moderate activity including 13% partial and 7% minor responses, and 30% stable disease in patients with refractory or relapsed non-Hodgkin’s lymphoma.12
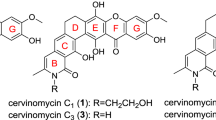
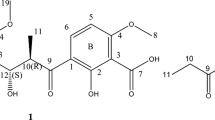
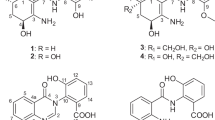
Chrysomycin is a classic antibiotic first reported in 1955.13 Although its antiphage and antibacterial activities and some physicochemical properties were described, the test samples contained a mixture of analogs1 with unknown structures. There were reports on the isolation and structure determination of chrysomycins A and B during the 1970s and 1980s, including studies of virenomycins as the synonym.1, 14, 15, 16, 17 Recently, chrysomycin C was reported as the reduced form of chrysomycin A and the existence of two other analogs, chrysomycins D and E, were indicated.18 Here, we describe the structures, physicochemical properties, cytotoxicity and antimicrobial activities of new 4’-acetylated analogs of chrysomycins A (1) and B (2) (Figure 1).
During our regular screening for antitumor compounds from the metabolites of actinomycetes using multiple human cancer cell lines and healthy mouse primary culture cells obtained from several organs, the culture broth extract of Streptomyces sp. strain MG271-CF2 showed about 100-fold stronger toxicity toward some cancer cells compared with normal cells. While the 16S ribosomal RNA gene sequence of the strain showed high homology with that of Streptomyces durmitorensis (strain MS405T, 1399 bp/1406 bp, 99.5%), S. aureus (strain B7319T, 1377 bp/1384 bp, 99.5%) and S. kanamyceticus (strain NBRC13414T, 1373 bp/1384 bp, 99.2%), the gene sequence showed rather lower homology with those of the other chrysomycin-producing species S. virens14 (strain NRRL B-24331, 1331 bp/1390 bp, 95.8%) and S. sporoverrucosus18 (strain NRRL B-16379, 1363 bp/1406 bp, 96.9%).
The small-scale separation of the culture extract by octadecyl-silica (ODS)-HPLC, HR-MS, and UV analyses indicated that chrysomycins A, B, and comparable amounts of two new analogous compounds 1 and 2 existed in the bioactive fractions. These two analogs are more hydrophobic, have similar UV spectra, and molecular weights of 42 Da larger (550 and 538 Da) than those of chrysomycins A and B (Supplementary Table 1), suggesting they are new monoacetylated analogs of chrysomycins. The new analogs are not the presumed analogs chrysomycins D and E because they are more hydrophilic than chrysomycins A and B and their molecular weights are 526 and 524 Da as determined by liquid chromatography-MS profiling, respectively.18 To enable structural determination and biological testing, larger-scale production and purification were attempted. The ethyl acetate extract of the producing strain culture was dissolved in methanol and left until sufficient lime precipitate mainly consisting of chrysomycins was formed. The precipitate was used as the source of the purification of 1 and 2. A previous report described that the separation of chrysomycins A and B was difficult and after subjecting the compounds to triacetylation, they were finally separated by HPLC.1 The new analogs 1 and 2 were also difficult to separate. As they were monoacetylated compounds and selective deacetylation could not be used after the separation, the triacetylation strategy could not be applied. Several liquid chromatography columns were tested in an attempt to purify each analog. While most of the columns were not suitable for separating the analogs, CHP-20 provided the best separation of 1 and 2. Because the analogs were not completely purified after a single separation, CHP-20 separations were performed repeatedly. ODS-HPLC was used to remove other impurities. Although there was still some material that was not purified (~20 mg each), 11.5 mg of 1 and 31.1 mg of 2 were isolated from the culture of 1620 g of barley medium (720 g grain). The physicochemical properties of 1 and 2 were analyzed as shown in Supplementary Table 1. The UV and IR spectra and optical rotation were consistent with those of chrysomycins and gilvocarcins2, 15, although the effect of 4’-acetylation on these physicochemical properties was unclear. The signs of optical rotations of 1 and 2 in acetic acid and methanol were opposite. A similar phenomenon was reported for chrysomycins, in which the signs were positive in acetic acid and negative in dimethyl sulfoxide (DMSO).15
The structures of 1 and 2 were determined by comparing their NMR and HR-MS data with that published for chrysomycins A and B1 (Figure 1, Table 1, Supplementary Figure 1 and Supplementary Table 1). The planar structures of 1 and 2 except for the acetylated moiety determined by 2D NMR correlations matched with those of chrysomycins A and B, respectively. The acetyl group was connected to the hydroxyl group at C-4’ (δC 77.1) of the sugar moiety as determined by the HMBC correlation from H-4’ to C-1” (δC 170.3). Some ravidomycins have a 4’-acetylation modification in their six-membered ring sugar.3 The structures of chrysomycin A and ravidomycins are similar in that they share the same chromophore and similar sugars. The difference is at C-3’, where the methyl group in chrysomycin A is changed to a N-dimethyl group and a hydroxyl group is deoxygenated in ravidomycins. It is expected that similar acetyltransferases may function to produce the new chrysomycin analogs and ravidomycins. The relative stereochemistry of the sugar moiety was determined by NOE, the optical rotation signs in acetic acid are same, and the absolute stereochemistry of the sugars of 1 and 2 were determined to be consistent with those of chrysomycins.
The cytotoxicities of chrysomycins toward cancer cells were very potent compared with the clinically used anticancer agent doxorubicin (Table 2 and Supplementary Table 2). Consistent with published reports,5 the analogs carrying a vinyl group, such as chrysomycin A and 1, showed especially strong cytotoxicity. Their IC50 values in sensitive cell lines were less than 0.1 ng ml−1. Because of the acetylation, the cytotoxicity of 1 was weaker than that of chrysomycin A. In contrast, the cytotoxicity of 2 was more than that of chrysomycin B. This discrepancy may be caused by the changes induced by the acetylation in the cellular permeation and in the binding to DNA and binding proteins. Some DNA intercalators not only bind to nucleic DNA but also to mitochondrial DNA either preferably or non-selectively. The localization of 1 and 2 in the culture cells was observed by harnessing their autofluorescence (Figure 2). The signal of 1 showed a similar pattern to that of 4',6-diamidino-2-phenylindole (DAPI) as it localized in the nuclei at high concentrations. The whole cells were obscurely bright and slightly stronger signals were observed around the nuclei at low concentrations. It seemed that it was more difficult for compound 2 to enter into the nuclei. Even at a high concentration, it showed similar patterns as that of the low concentration of 1. Compounds 1, 2, and DAPI were strongly observed in the rounded cells, which were probably in M phase. The collapse of the nuclear membrane might facilitate the access of the compounds to the nuclear DNA. The preference of mitochondrial DNA for the intercalator ethidium bromide (EtBr) led it to be localized in the nuclei at the high concentration but was localized in the mitochondria and nucleolus at the low concentration as previously reported.19 Such mitochondrial selective localization was not observed for 1, 2, and DAPI at the concentration at which their signals were observable.
Localization of 1 and 2 in culture cells. Human cancer cell lines ME-180 and HS-Os-1 were treated with 1, 2, 4',6-diamidino-2-phenylindole (DAPI), and ethidium bromide (EtBr) at a high (10 μg ml−1 for all compounds) and low (0.02 μg ml−1 for EtBr and 0.1 μg ml−1 for other compounds) concentration for 1.5 h and the compound autofluorescence was observed. Because the low concentration treatments emitted weak signals and exposed with stronger conditions, the backgrounds were brighter. PC and AF are phase contrast and autofluorescence, respectively.
While 1 and 2 showed no or low anti-fungal and anti-Gram-negative bacterial activity, they showed anti-Gram-positive bacterial activity (Supplementary Tables 3 and 4). Chrysomycin A and 1 showed strong anti-Gram-positive bacterial activity toward MRSA and VRE strains with MIC values of 0.5–2 μg ml−1. This trend is consistent with previously reported antimicrobial activities of chrysomycins.13, 16, 20 The only difference was noted for Mycobacterium smegmatis where a crude mixture of chrysomycins was used and a MIC value of 0.6 μg ml−1 was obtained, while the analogs reported herein had MIC values greater than 64 μg ml−1.13 This difference might be caused by some impurities in their chrysomycin mixture.
Because their toxicity to mouse is often low and their cytotoxicity to tumor cells and anti-Gram-positive bacterial activity are high, chrysomycins, including the new analogs found in this study, could be good drug candidates for the treatment of these illnesses.
References
Weiss, U. et al. The chemistry of the antibiotics chrysomycin A and B. Antitumor activity of chrysomycin A. J. Antibiot. 35, 1194–1201 (1982).
Takahashi, K., Yoshida, M., Tomita, F. & Shirahata, K. Gilvocarcins, new antitumor antibiotics. 2. Structural elucidation. J. Antibiot. 34, 271–275 (1981).
Findlay, J. A., Liu, J. S., Radics, L. & Rakhit, S. The structure of ravidomycin. Can. J. Chem 59, 3018–3020 (1981).
Findlay, J. A., Liu, J. S. & Radics, L. On the structure, chemistry, and C-13 nuclear magnetic-resonance of ravidomycin. Can. J. Chem. 61, 323–327 (1983).
Li, Y. Q. et al. Plasticity in gilvocarcin-type C-glycoside pathways: discovery and antitumoral evaluation of polycarcin V from Streptomyces polyformus. Org. Biomol. Chem. 6, 3601–3605 (2008).
Matson, J. A. et al. Antitumor activity of chrysomycins M and V. J. Antibiot. 42, 1446–1448 (1989).
Nakano, H. et al. Gilvocarcins, new antitumor antibiotics. 1. Taxonomy, fermentation, isolation and biological activities. J. Antibiot. 34, 266–270 (1981).
Morimoto, M., Okubo, S., Tomita, F. & Marumo, H. Gilvocarcins, new antitumor antibiotics. 3. Antitumor activity. J. Antibiot. 34, 701–707 (1981).
Lorico, A. & Long, B. H. Biochemical characterisation of elsamicin and other coumarin-related antitumour agents as potent inhibitors of human topoisomerase II. Eur. J. Cancer 29A, 1985–1991 (1993).
Matsumoto, A. & Hanawalt, P. C. Histone H3 and heat shock protein GRP78 are selectively cross-linked to DNA by photoactivated gilvocarcin V in human fibroblasts. Cancer Res. 60, 3921–3926 (2000).
Goss, G. et al. Phase II study of elsamitrucin in non-small cell lung cancer. Invest. New Drugs 12, 315–317 (1994).
Allen, S. L. et al. Phase II study of elsamitrucin (BMY-28090) for the treatment of patients with refractory/relapsed non-Hodgkin's lymphoma. Invest. New Drugs 14, 213–217 (1996).
Strelitz, F., Flon, H. & Acheshov, I. N. Chrysomycin: a new antibiotic substance for bacterial viruses. J. Bacteriol. 69, 280–283 (1955).
Gauze, G. F. et al. Formation of new antibiotic, virenomycin, by a culture of Streptomyces virens sp. nev. Antibiotiki. 22, 963–967 (1977).
Brazhnikova, M. G. et al. Physico-chemical characteristics of the new antitumor antibiotic virenomycin. Antibiotiki. 22, 967–970 (1977).
Kudinova, M. K. et al. Separation and characteristics of the components of the antibiotic virenomycin. Antibiotiki. 27, 507–511 (1982).
Brazhnikova, M. G. et al. Structure of the antibiotic virenomycin. Antibiotiki. 29, 884–892 (1984).
Jain, S. K. et al. Chrysomycins A-C, antileukemic naphthocoumarins from Streptomyces sporoverrucosus. RSC Adv. 3, 21046–21053 (2013).
Ashley, N. & Poulton, J. Mitochondrial DNA is a direct target of anti-cancer anthracycline drugs. Biochem. Biophys. Res. Commun. 378, 450–455 (2009).
Wei, T. T., Byrne, K. M., Warnick-Pickle, D. & Greenstein, M. Studies on the mechanism of actin of gilvocarcin V and chrysomycin A. J. Antibiot. 35, 545–548 (1982).
Acknowledgements
This work was supported by the Japan Society for the Promotion of Science (23701108, 15K08013 and 26450107). We acknowledge colleagues at the Institute of Microbial Chemistry for their support of this study and for helpful discussions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on The Journal of Antibiotics website
Rights and permissions
About this article
Cite this article
Wada, Si., Sawa, R., Iwanami, F. et al. Structures and biological activities of novel 4’-acetylated analogs of chrysomycins A and B. J Antibiot 70, 1078–1082 (2017). https://doi.org/10.1038/ja.2017.99
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2017.99
This article is cited by
-
Optimization of fermentation conditions and medium components for chrysomycin a production by Streptomyces sp. 891-B6
BMC Microbiology (2024)
-
Chrysomycin A inhibits the topoisomerase I of Mycobacterium tuberculosis
The Journal of Antibiotics (2022)
-
Functional Characterization of NgnL, an Alpha/beta-hydrolase Enzyme Involved in Biosynthesis of Acetylated Nodusmicin
Biotechnology and Bioprocess Engineering (2020)