Abstract
Pseudomonas aeruginosa and Aspergillus fumigatus are the two microorganisms responsible for most of the chronic infections in cystic fibrosis patients. P. aeruginosa is known to produce quorum-sensing controlled rhamnolipids during chronic infections. Here we show that the dirhamnolipids secreted from P. aeruginosa (i) induce A. fumigatus to produce an extracellular matrix, rich in galactosaminogalactan, 1,8-dihydroxynaphthalene (DHN)- and pyo-melanin, surrounding their hyphae, which facilitates P. aeruginosa binding and (ii) inhibit A. fumigatus growth by blocking β1,3 glucan synthase (GS) activity, thus altering the cell wall architecture. A. fumigatus in the presence of diRhls resulted in a growth phenotype similar to that upon its treatment with anjpegungal echinocandins, showing multibranched hyphae and thicker cell wall rich in chitin. The diRhl structure containing two rhamnose moieties attached to fatty acyl chain is essential for the interaction with β1,3 GS; however, the site of action of diRhls on GS is different from that of echinocandins, and showed synergistic anjpegungal effect with azoles.
Similar content being viewed by others
Introduction
Aspergillus fumigatus and Pseudomonas aeruginosa are the two microorganisms responsible for most of the chronic infections encountered in cystic fibrosis adult patients. A. fumigatus is indeed the major filamentous fungus reported to colonize the airways of these patients, with a prevalence of 12–57% (Paugam et al., 2010), and major risks in developing allergic bronchopulmonary aspergillosis in 15% of patients (Knutsen et al., 2012; Baxter et al., 2013). The most common bacterium isolated from the sputum of patients with cystic fibrosis is P. aeruginosa, which chronically colonizes cystic fibrosis airways in up to 75% of adult patients (Hill et al., 2005; Moree et al., 2012; Baxter et al., 2013). The co-colonization by A. fumigatus and P. aeruginosa is very deleterious for the patients, characterized by a high decline in cystic fibrosis pulmonary functions and the worst prognosis with no access to lung transplantations (Ferreira et al., 2015).
Several recent studies have shown that P. aeruginosa and A. fumigatus interact during lung colonization. The first evidence for crosstalk’s in vivo between A. fumigatus and P. aeruginosa emerged from the fact that antibacterial therapy targeting P. aeruginosa colonization is followed by a significant reduction in A. fumigatus colonization (Baxter et al., 2013). Two studies have shown a direct interaction between P. aeruginosa and A. fumigatus (Blyth, 1971; Mowat et al., 2010). P. aeruginosa secretes molecules such as homoserine-lactones, phenazines, siderophores, quinolones, which interfere with the behavior of A. fumigatus (Mowat et al., 2010; Moree et al., 2012; Briard et al., 2015). Homoserine-lactones reduce the growth of A. fumigatus, whereas phenazines have a dual effect on A. fumigatus. At low concentrations such as in an iron-starved environment, pyocyanin, phenazine-carboxamide and phenazine-carboxylic acid could stimulate the growth of A. fumigatus by reducing the Fe(III). However, at high concentrations, all phenazines (including 1-hydroxy-phenazine) penetrated into the cells and induced the production of reactive-oxygen species and reactive-nitrogen species leading to fungal death (Briard et al., 2015). Such interactions were not demonstrated in vivo; however, phenazines were detected in the sputum of cystic fibrosis patients at concentrations which stimulated the growth of A. fumigatus in vitro (Wilson et al., 1988; Bjarnsholt et al., 2010; Briard et al., 2015). Regarding quinolones, their effect on A. fumigatus or on other fungi was never studied (Diggle et al., 2003).
The present study is centered on rhamnolipids (Rhls), which are secreted by P. aeruginosa and other Pseudomonas species (Abalos et al., 2001; Moree et al., 2012). To date these molecules have been shown to display only tension active properties. Our study demonstrated that the growth inhibition of A. fumigatus and an increased thickness of A. fumigatus cell wall following the adhesion of P. aeruginosa to A. fumigatus is due to the secretion of dirhamnolipids (diRhls) which inhibit the fungal β1,3 glucan synthase (GS).
Materials and methods
Strains and culture conditions
The A. fumigatus reference strain used in this study is CEA17ΔakuBKU80 (ku80) deficient in non-homologous end joining (da Silva Ferreira et al., 2006), which originates from the clinical isolate CBS 144-89, and is as pathogenic as CBS 144-89 in experimental murine aspergillosis models.
CEA17ΔakuBKU80 was used to generate the ΔpksP, ΔhppD, Δsph3 deletion strains and EMFR S678P mutant. ΔpksP is a 1,8-dihydroxynaphthalene (DHN)-melanin-deficient mutant (Jahn et al., 2000); ΔhppD is a pyo-melanin-deficient mutant and is a kind gift of AA Brakhage (Hans Knöll Institute, Jena, Germany) (Schmaler-Ripcke et al., 2009); Δsph3 is a spherulin-4-like-deleted mutant, deficient in the production of galactosaminogalactan (GAG), constructed as described in the Supplementary Information (Supplementary Figure 1). EMFR S678P is resistant to caspofungin, due to a point mutation for Serine678 in proline in the unique FKS gene coding for the β1,3 GS (Rocha et al., 2007), and is a kind gift from DS Perlin (Public Health Research Institute, Newark, NJ, USA).
All strains were conserved on 2% (w/v) malt agar slants. One-week-old conidia were recovered from the slants by vortexing with 0.05% (v/v) aqueous Tween 20 solution and used for inoculation in 2YT or Minimal Medium (Briard et al., 2015), RPMI (Clavaud et al., 2012) or Sabouraud (Lamarre et al., 2009) media.
The P. aeruginosa used in this study is the reference strain PAO1 (Holloway et al., 1994). This strain was used to generate the deletion strain, ΔrhlA, which do not produce Rhls and is a kind gift of Niels Hǿiby (Department of clinical microbiology, Rigshospitalet, Copenhagen) (Jensen et al., 2007). P. aeruginosa strains were conserved in 2YT medium containing 50% glycerol at −80 °C.
Co-culture of A. fumigatus and P. aeruginosa
To determine the best conditions, preliminary experiments were done with different culture agar-media in Petri dishes (90 mm diameter) as described in the Supplementary Information. The selected co-interaction model is 2 × 107 ku80 conidia inoculated on RPMI plates spotted at the same time with 5 μl of 2.5 × 105 PAO1 and incubated for 16 h at 37 °C.
Culture of A. fumigatus in presence of culture filtrate or dirhamnolipids of P. aeruginosa
PAO1 (A600=0.05) was inoculated in 50 ml 2YT for 24 h at 37 °C. The bacteria were separated by centrifugation at 15 000 r.p.m. for 10 min and the culture filtrate (CF) was filtrated through 0.22 μm Minisart filter (Sigma-Aldrich, Saint-Quentin Fallavier, France). A. fumigatus conidia (2 × 107) suspended in 3 ml 0.4% agar containing PAO1 CF was overlaid on RPMI-agar and incubated for 16 h at 37 °C. The mycelia grown were observed by transmission electron microscopy (TEM) as described below.
To screen for the CF-active fraction during purification (described below), 1.5 × 104 A. fumigatus conidia ml−1 were inoculated on eight-well glass bottom Ibidi μ-slides in 2YT medium reconstituted using the PAO1 CF instead of water or in presence of purified diRhls (obtained from CF as described below). The culture was incubated at 37 °C for 24 h. Mycelium was observed under light microscopy.
Analysis of the binding of P. aeruginosa to A. fumigatus polysaccharides, melanin and hyphae
P. aeruginosa was incubated with A. fumigatus cell wall α1,3 glucan, β1,3 glucan, galactomannan, GAG, melanin, chitin (Sigma-Aldrich), and ku80 or Δsph3 hyphae as described in the Supplementary Information.
Scanning and transmission electron microscopy preparation
The zone of partial fungal–bacterial inhibition (Figure 1a) from A. fumigatus and P. aeruginosa co-culture was cut in the Durapore filter, fixed in 2.5% glutaraldehyde in 0.1 m sodium cacodylate buffer for 24 h at 4 °C and prepared for scanning electron microscopy and TEM (Lamarre et al., 2009; Briard et al., 2015). As a control, the same culture was prepared without P. aeruginosa.
Interaction between A. fumigatus ku80 and P. aeruginosa PAO1 strains. (a) Zone of partial inhibition (dotted square) between A. fumigatus and P. aeruginosa. (b) Scanning (b1,2 and5) and transmission (b3,4 and6) electron microscopy views of the zone of partial inhibition; (b1–4) ku80-PAO1; (b4) enlargement of the interaction zone showed in b3; (b5 and6) control ku80 in the absence of PAO1. White arrow, bacteria; black arrow, fungus. Note the presence of an increased amount of fungal ECM in presence of P. aeruginosa (b1 compared to b5 and b3 compared to b6). Scale bars, 1 μm.
The thickness of the cell wall was calculated from 50 hyphae with six measures per hyphae from TEM pictures using the ImageJ/fiji analysis (Schneider et al, 2012).
Extraction and isolation of the active molecules from P. aeruginosa 24 h culture filtrate
The PAO1 24 h CF was extracted twice with ethyl acetate (1:1) and the organic phase was evaporated and dried. A mixed fraction, F-diRhls, containing two to three diRhls depending on the CF batches, was purified from the organic extract as described in the Supplementary Information. Each fraction was idenjpegied by high-resolution mass spectrometry, which correlated with the published data (Sharma et al., 2007; Abdel-Mawgoud et al., 2010; Arutchelvi and Doble, 2010).
Chemical synthesis of dirhamnolipid analogs diRhamnose-C3 and diRhamnose-C8
The synthesis of two diRhls analogs, diRhamnose-C3 (diRha-C3) and diRhamnose-C8 (diRha-C8) bearing a simpler aglycon—a propyl and an octyl chains, respectively, is described in the Supplementary Information.
Method for susceptibility testing and biomass quanjpegication by crystal violet
Minimal effective concentration (MEC) of the molecules was determined according to the Clinical Laboratory Standards Institute M38-A2 protocol (NCCM) by microdilution method in 96-well plates using crystal violet biomass quanjpegication as previously described (Briard et al., 2015).
Determination of the fractional inhibitory concentration (FIC) index (FICi)
The FICi of drug interactions was determined via checkerboard titration assays (Clavaud et al., 2012). The concentrations of the anjpegungal agents ranged from 0.008 to 1 μg ml−1, 0.004 to 0.5 μg ml−1 and 0.031 to 4 μg ml−1 for caspofungin, voriconazole and itraconazole, respectively. The concentrations of F-diRhls ranged from 0.03 to 4 mm. Serial twofold dilutions of anjpegungals were prepared with the assay culture in 2YT in 96-wells flat bottom plates as described above. The microtiter plates were incubated at 37 °C for 20 h and the growth biomass was assessed using crystal violet and resazurin methods (Clavaud et al., 2012; Briard et al., 2015). The pharmacodynamic interaction analysis was done as described (Mavridou et al., 2015). Briefly, the synergistic, additive or antagonistic effect of paired combinations of drugs was captured by the FICi: (a) FIC=FICdrugA+FICdrugB; (b) FICdrugA=MICdrugsA+B/MICdrugA; and (c) FICdrugB=MICdrugsA+B/MICdrugB. If the FIC ∑FICmin value is lower than 1, this indicates synergistic interactions between two drugs; if FIC ∑FICmax value is higher than 1.25, then an antagonistic interaction exists and between these two values, the interactions between the two drugs are additive.
Effect of diRhl on cell wall
Glucosamine and galactosamine determination in ku80 cell wall in presence and absence of F-diRhls and preparation of cell-free glucan and chitin synthases extracts are described in the Supplementary Information.
Statistical analysis
Data are reported as means±s.e.m. Comparisons, performed with Graph Pad Prism 3.0 software and analysis of variance statistical test.
Results
Morphological modifications of A. fumigatus mycelium in presence of P. aeruginosa
Preliminary experiments determined the best culture conditions allowing the fungus and the bacterium to grow together: 2 × 107 ku80 A. fumigatus conidia were inoculated on an RPMI agar plates and at the same time 5 μl of 2.5 × 105 PAO1 P. aeruginosa was spotted in the center of the Petri-dish. Under these experimental conditions, a zone of partial fungal inhibition was observed at the junction between the bacterial and the fungal colonies (Figure 1a). The zone of partial inhibition was observed under scanning electron microscopy and TEM (Figure 1b). The first observation under scanning electron microscopy showed an electron-dense extracellular matrix (ECM) embedding the hyphae of A. fumigatus which was more abundant in the presence of the bacteria than in its absence (Figures 1b1). PAO1 cells adhered to the hyphae, mainly at their apex and to the ECM (Figures 1b1). To idenjpegy the fungal cell wall molecule to which P. aeruginosa was binding, the A. fumigatus cell wall polysaccharides α1,3 glucan, β1,3 glucan, galactomannan, chitin, GAG and melanin (Latgé, 2010) were incubated with the PAO1 cells and scored the binding under light microscopy. The bacteria only bound to GAG (Figure 2; Supplementary Figure 2 and data not shown). Addition of 1.2 m NaCl prevented the binding of the bacteria to GAG showing that PAO1 binding was due to ionic interactions (Figures 2a and b). To confirm that GAG is responsible for the binding of P. aeruginosa to A. fumigatus ku80 hyphae, the spherulin-4 like gene, coding for Sph3 involved in the synthesis of GAG in A. fumigatus (Bamford et al., 2015) was deleted in ku80 (Supplementary Figure 1) and the Δsph3 strain was incubated with P. aeruginosa. As shown in Figures 2c and d, P. aeruginosa did not bind to Δsph3 hyphae.
Binding of P. aeruginosa to A. fumigatus GAG. (a) incubation of PAO1 with purified GAG. (b) Incubation of PAO1 with purified GAG in presence of 1.2 m NaCl. (c) Adhesion of PAO1 to A. fumigatus ku80 hyphae. (d) Absence of adhesion of PAO1 to AfΔsph3 mutant hyphae, unable to produce GAG. Note that the binding of PAO1 to GAG is inhibited under high salt concentration (b). Scale bars, 25 μm.
The interaction between bacterium and fungus induced formation of an electron-dense ECM around the fungal hyphae. This electron-dense material was absent in the control cultures (in absence of bacteria) as well as at the contact point between the fungal cell wall and P. aeruginosa PAO1 (Figures 1b3). This electron-dense material is reminiscent of melanin observed around hyphae in the A. fumigatus biofilm (Beauvais et al., 2007). Three A. fumigatus deletion mutants ΔpksP, ΔhppD and ΔpksPΔhppD affected in DHN-, pyo- or both melanin synthesis, respectively (Jahn et al., 2000; Schmaler-Ripcke et al., 2009), were grown in with the presence of PAO1. As shown in the Figure 3, this electron-dense material was partially lost in ΔpksP and ΔhppD, and was absent in ΔpksPΔhppD. Therefore, the electron-dense material present in the ECM in presence of PAO1 was composed of both DHN- and pyo-melanin.
Presence of melanin in the ECM of A. fumigatus ku80 during co-cultivation with P. aeruginosa PAO1 in RPMI. (a) Parental strain ku80. (b) AfΔpksP mutant unable to produce DHN-melanin. (c) AfΔhppD mutant unable to produce pyo-melanin. (d) Double AfΔpksPΔhppD mutant. Note the absence of melanin in AfΔpksPΔhppD, showing that melanin in ECM is a mixture of DHN and pyo-melanin. Scale bars, 250 nm.
TEM also showed that A. fumigatus presented a thicker cell wall (104 nm±3.2) when grown in the presence of P. aeruginosa (Supplementary Figure 3), compared to that in its absence (72 nm±1.19). This observation suggested that bacterial co-culture induced modification of the fungal cell wall structure.
P. aeruginosa secreted dirhamnolipids are responsible for the A. fumigatus cell wall modification
To test whether P. aeruginosa induced cell wall modifications in A. fumigatus requires its direct contact or due to molecules secreted from bacteria, A. fumigatus conidia were incubated in the presence of bacterial CF. The CF from either 2YT or RPMI media promoted the formation of an ECM around fungal hyphae and induced thickening of the fungal cell wall (Figure 4a). Increase in the fungal cell wall thickness (106 nm) was comparable to that observed when bacteria are directly in contact with the fungus. In addition, A. fumigatus growth was reduced with multibranched hyphae, the phenotype similar when this fungus is grown in the presence of anjpegungal echinocandins (Figure 4b; Perlin, 2011). These results suggested that diffusible molecules in P. aeruginosa PAO1 CF were responsible for the fungal cell wall modifications.
A. fumigatus hyphal morphology grown in presence of 24 h PAO1 CF. (a) Thickness of A. fumigatus ku80 cell wall in presence of CF in RPMI, calculated from 50 hyphae with six measures per hyphae from TEM pictures using the ImageJ/fiji analysis (Schneider et al, 2012). (b) Light microscopy images showing the increased branching of the fungal hyphae in presence of CF. Scale bar 50 μm.
The molecules responsible for the cell wall phenotype were then purified from a 2YT CF of P. aeruginosa after 24 h of growth. Only one high-performance liquid chromatography fraction gave the cell wall phenotype mentioned above when incubated with A. fumigatus. MS analyses (data not shown) showed that this fraction contained a mixture of diRhls (F-diRhls). Following purification, three rhamnobiosides differing by the length of their fatty acid chains were idenjpegied: diRha-C10-C10 eluting at tR: 7.29 min had m/z 650.39 (HRMS-ESI+ for C32H58O13 ([M+Na]+, 673.3775), diRha-C10-C11 eluting at tR: 8.93 min had m/z 664.4 (HRMS-ESI+ for C33H60O13 ([M+Na]+, 687.3932), diRha-C10-C12:1 and diRha-C12:1-C10 eluting at tR: 9.10 min and 9.47 min, respectively, had m/z 676.4 (HRMS-ESI+ for C34H60O13 ([M+Na]+, 699.3932) (Figure 5). F-diRhls was majorly composed of diRha-C10-C10 (60–100% depending of the CF batches). Each diRhls was either tested individually or in combination (F-diRhls) on A. fumigatus.
Analytical RP-HPLC profiles of the dirhamnolipids isolated following RP-HPLC purification of the 24 h culture filtrate of P. aeruginosa. Elution from a C18 Kromasil column (4.6 × 150 mm, 5 μm, 100 Å) at 1.0 ml min−1 with CH3CN in 0.08% aqueous trifluoroacetic acid (60→100% over 15 min.), detection with an evaporative light scattering detector and idenjpegication according to the literature (Sharma et al., 2007; Abdel-Mawgoud et al., 2010; Arutchelvi and Doble, 2010). (a) diRha-C10-C10. (b) diRha-C10-C10-CH3 (diRha-C10-C11) corresponding to the methyl ester of diRha-C10-C10. (c) diRha-C10-C12:1 and diRha-C12:1-C10 bearing an unsaturated fatty acid.
TEM observation of the hyphae after treatment with 4 mm diRha-C10-C10 showed an increased thickness of the cell wall (215 nm±5,6) and ECM as observed previously with P. aeruginosa PAO1 and its CF (Supplementary Figures 4a and c). This increased thickness of the cell wall could be correlated to an increased concentration of the GAG adhesin at the cell surface. This hypothesis was verified biochemically by determination of the amount of galactosamine monomers, representative of GAG, in the extracted cell wall in absence and presence of 4 mm F-diRhls; 1.74 μg (±0.44) galactosamine per mg mycelium dry weight was found in A. fumigatus cell wall in the absence of F-diRhls which was significantly increased to 11.26 μg (±0.14) galactosamine per mg mycelium dry weight in the presence of F-diRhls.
To confirm that F-diRhls were responsible for the morphological changes in A. fumigatus cell wall, co-culturing of the fungus and the P. aeruginosa ΔrhlA mutant, which is unable to synthesize Rhls, was undertaken. The ΔrhlA mutant bound to ku80 hyphae similar to P. aeruginosa PAO1 but no melanin was present as in the control without bacteria (Figure 1b6; Supplementary Figure 4b). Moreover, the cell wall thickness was not modified in presence of ΔrhlA (Supplementary Figure 4c). These results confirmed that the cell wall phenotype induced by the bacterium was due to the presence of diRhls secreted by PAO1.
Fungal growth phenotype is due to the inhibition of β1,3 glucan synthase by diRhls
A. fumigatus had a reduced growth and altered morphology in presence of diRhls similar to that observed with CF (Figures 4b and 6a). The effect of F-diRhls on A. fumigatus was never fungicidal whatever concentration used. The short hyperbranched morphology was reminiscent of the echinocandin effect. Accordingly, only an MEC, which is the lowest drug concentration resulting in aberrant hyphal growth, can be measured (Arikan et al., 2002). The MEC was 4 mm, and resulted in 90% inhibition of A. fumigatus growth (Figure 6b). Calcofluor white treatment showed an increase in hyphal fluorescence compared to untreated control after treatment with purified diRhls and F-diRhls at MEC, suggesting an increase in the concentration of cell wall chitin (Figures 6a6 and 7). This hypothesis was verified biochemically by determination of the amount of glucosamine monomers, representative of chitin, in the extracted cell wall treated and untreated with F-diRhls at MEC; in the absence of F-diRhls, 9.15 μg (±1.55) glucosamine per mg mycelium dry weight was found, which was significantly increased (31.4 μg (± 0.38) glucosamine per mg mycelium dry weight) in the presence of F-diRhls. An increase in the cell wall chitin amount was also observed after incubation of the fungus with echinocandins (Walker et al., 2010). These data suggested that diRhls may have a direct effect on the β1,3 GS activity. Indeed, the F-diRhls inhibited the GS activity in a dose–response manner (Figure 7a). To determine if the inhibition of the GS activity was due to the detergent characteristic of the F-diRhls, other neutral nonionic detergent glycolipids such as octylglucoside and maltoside were tested. These detergents disrupt protein–lipid and lipid–lipid interactions in the membrane, which could disturb their activity. Figure 7a showed a slight statistically insignificant inhibition of GS activity by octylglucoside and maltoside, showing that the dose-dependent inhibition of GS by F-diRhs was mainly due to their GS-specific effect. Moreover, we verified the specificity of F-diRhls on the GS activity by testing the chitin synthase activity in presence of 1 mm F-diRhls, octylglucoside and maltoside. All compounds induced a similar statistically insignificant decrease of the chitin synthase activity (Supplementary Figure 5), showing that in contrast to GS inhibition, their minimal effect on the chitin synthase was not specific, and was due to biosurfactant activity on the membranes.
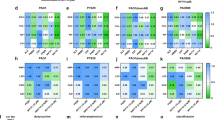
Impact of the diRhls on the A. fumigatus mycelial growth. (a1–4) Light microscopy of A. fumigatus grown for 24 h in the presence of diRha-C10-C10, diRha-C10-C11, [diRha-C12:1-C10 - diRha-C10-C12:1] or F-diRhls at 4 mM. (a5) A. fumigatus grown in the absence of diRhls. Note the increased branching of mycelium after growth with diRhls. (a6 and7) UV microscopy of calcofluor white labeling of A. fumigatus in the absence or presence of diRhls. Note the increased amount of fluorescence in the hyphae in presence of F-diRhls, which is representative of an increase in the chitin. (b) Reduction of mycelial growth by different concentrations of F-diRhls. Scale bars, 50 μm (a1–a5); 10 μm (a6 and a7).
Specific inhibition of A. fumigatus ku80 β1,3 glucan synthase activity by dirhamnolipids (F-diRhls). (a) β1,3 glucan synthase activity (**P=0.0036; ***P<0.0001); 0.5–4 mm F-diRhls, 5 mm rhamnobiose (diRha), 1 mm octylglucoside and maltoside, 4 mm propyl- and octyl-rhamnobioside analogs (diRha-C3 and diRha-C8, respectively) were added in the glucan synthase reaction mixture (P>0.05). (b) Structure of the chemically synthesized diRha-C3 and diRha-C8. (c) phenotype of A. fumigatus EMFR S678P in absence and presence of 0.5 mm F-diRhls.
To investigate the active moiety of F-diRhls on GS activity, we tested the inhibitory effect of rhamnobiose and the propyl- and octyl-rhamnobioside analogs, equipped with a linear three carbons (diRha-C3) and eight carbons (diRha-C8) aglycon (Figures 7a and b). None of these molecules inhibited GS activity, suggesting that the branched lipidic tail with β-hydroxy fatty acids of F-diRhls is essential for its activity against GS.
The A. fumigatus EMFR S678P strain, which is a mutant resistant to caspofungin consecutively to a point mutation in the binding site of this echinocandin to GS, has a MEC⩾16 μg caspofungin ml−1, while the MEC of caspofungin for parental ku80 strain is 0.25 μg ml−1 (Rocha et al., 2007). Surprisingly, the MEC of F-diRhls on the EMFR S678P strain was 0.5 mm compared to the 4 mm value with the A. fumigatus ku80 strain (Figure 7c). This result indicated that the diRhls and the echinocandin do not bind to the same GS site and that the S678P mutation favored the inhibitory effect of F-diRhls towards GS activity. When F-diRhls and caspofungin were tested in combination on the ku80 strain, the combined effect of two molecules was antagonist with a FICi ∑FICmax of 4.25 (Table 1). In contrast to the previously reported additive effect of caspofungin with voriconazole (Elefanti et al., 2013), the results presented in Table 1 showed that the combined effect of F-diRhls and voriconazole or itraconazole was synergistic with a FICi ∑FICmin of 0.5 and 0.56, respectively. These data supported the fact that diRhls and caspofungin target the same component of A. fumigatus, the GS.
This study shows that the F-diRhls induce several morphological effects: (i) they modify the nature of fungal ECM surrounding the A. fumigatus hyphae; (ii) they inhibit the growth of A. fumigatus and induce the formation of short multibranched hyphae, which results from the specific inhibition of the fungal GS activity; (iii) they induce a thickening of the cell wall due to compensatory reactions resulting from an increased chitin concentration to palliate the anjpegungal effect of dirhamnolipids; and (iv) the site of action of diRhls on GS is different from that of echinocandins, which facilitates its synergistic activity with azoles.
Discussion
This study has shown that P. aeruginosa binds strongly to the A. fumigatus hyphae. We previously reported that the polysaccharides composing A. fumigatus ECM were α1,3 glucan, GAG and galactomannan (Beauvais et al., 2007). In the binding experiments, in presence of P. aeruginosa, only GAG was found to be the molecule responsible for P. aeruginosa to A. fumigatus binding. GAG has been reported previously to be a major adhesin of A. fumigatus with many functions associated to the invasion of the host tissue (Gresnigt et al., 2014; Robinet et al., 2014). The current study also demonstrated that the production of GAG is increased in response to bacterial assault against the fungus.
Interestingly, the bacteria bind to the fungus principally at the apex that is at the site of the metabolic fungal activity. Increased fixation at the tip of A. fumigatus hyphae, where intense production of exudates takes place (Toljander et al., 2007), may be a reason for P. aeruginosa to obtain nutrients from the fungus. Similarly, P. aeruginosa binding to the hyphal (but not to the yeast) form of Candida albicans suggests an interaction via hyphal specific glycoproteins, lectins or adhesins (Hogan and Kolter, 2002). The ligand has not been idenjpegied yet.
The present and previous studies from our laboratory and others showed that P. aeruginosa produces two types of molecules which can influence A. fumigatus growth. The first ones are the toxins which directly kill A. fumigatus. Examples of those are phenazines and homoserine lactones (Mowat et al., 2010; Moree et al., 2012; Briard et al., 2015), which display different toxic effects. Phenazines kill the fungus by inducing intracellular reactive-oxygen species and reactive-nitrogen species production (Briard et al., 2015). Homoserine lactones repressed C. albicans filamentation (Hogan et al., 2004) and reduced A. fumigatus growth, but their mode of action is not yet studied (Mowat et al., 2010).
The second type of compounds is stress molecules which induces a typical anti-stress response from A. fumigatus. An example is the diRhls. A. fumigatus responds to diRhls in a way which is very similar to any response to environmental molecules including anjpegungals targeting cell wall such as echinocandins, calcofluor white or congo red (Ram and Klis, 2006; Perlin, 2011; Lee et al., 2012). Interestingly, use of diRhls led to the discovery of a new cell wall compensatory mechanism not previously described, which is the melanin synthesis.
Melanin synthesis is associated with reduced fungal susceptibility to a variety of aggressions. Infection increases the expression of melanin synthesis genes by various fungi and the more virulent strains produce the more melanin (Eisenman and Casadevall, 2012). In the host, melanin interferes with the normal function of phagocytic cells and scavenges reactive-oxygen species (Eisenman and Casadevall, 2012; Chamilos et al., 2016). In A. fumigatus, melanin production has been described in biofilm (Beauvais et al., 2007) or in vivo the production of DHN-melanin was suggested during experimental aspergillosis under conditions which can be considered to be starved stress conditions (Langfelder et al., 2001). The origin of melanin in A. fumigatus biofilm is still unknown. It was hypothesized that melanin resulted from air-oxidation of melanin precursors in the biofilm channels. The use of two A. fumigatus mutants unable to produce DHN- and pyo-melanin when co-cultured with P. aeruginosa indicated that ECM synthesized by parental A. fumigatus in presence of P. aeruginosa is composed of DHN- and pyo-melanin (Figure 3). Interestingly the melanin produced by the fungus in response to PAO1 disappeared at the contact region with the bacterium. Klebsiella aerogenes synthesizes a dopamine precursor from l-tyrosine that was used by Cryptococcus neoformans for melanin production, which required a fungal laccase (Frases et al., 2006). A. fumigatus expressed laccases during vegetative growth and a 4-hydroxyphenylpyruvate dioxygenase (HppD) responsible for the synthesis of pyo-melanin from l-tyrosine present in different media (Sugareva et al., 2006; Schmaler-Ripcke et al., 2009). Moreover, A. fumigatus interacting with P. aeruginosa ΔrhlA, which does not produce any Rhl, did not synthesize melanin demonstrating that the diRhls were inducing the synthesis of the melanin protecting layer by the fungus. However, the antibacterial function of the fungal melanin and especially the role of the two different melanins have not been investigated.
In P. aeruginosa, because of their metabolic synthesis, Rhls are usually produced as a mixture of homologs di- and mono-Rhls diverging in terms of lipid chain length. To our knowledge, diRha-C10-C10 is always the major component (Abalos et al., 2001; Sha et al., 2012; Christova et al., 2013; Singh et al., 2013; Gogoi et al., 2016). DiRha-C10-C10 showed higher anjpegungal activities on plant pathogens such as Penicillium juniculosum and Bothytis cinerea than Rhls (Sha et al., 2012). DiRha-C10-C10 was also active against C. albicans with a MIC>0,15 mm (Singh et al., 2013). At 7 mm, diRha-C10-C10 disrupted the biofilms and their biosurfactant activity inhibited cell adhesion. The presence of diRhls after 24 h in PAO1 CF is in accordance with the kinetics of Rhls synthesis by P. aeruginosa after the growth ceased (24–96 h) (Haba et al., 2003; Laabei et al., 2014; Schmidberger et al., 2014). Rhamnolipids are overproduced in vitro by P. aeruginosa during nutrient limitation such as in iron starvation (Schmidberger et al., 2014). In vivo in the host, iron concentration is limited, and 2.4–15.3 μg ml−1 Rhls were found in sputa of all cystic fibrosis patients chronically infected by P. aeruginosa (Kownatzki et al., 1987). Mucoid isolates from chronically infected patients produced more Rhls than non-mucoid isolates (Bjarnsholt et al., 2010). Indeed, they are components of P. aeruginosa biofilm and protect the bacteria against phagocytosis by polymorphonuclear neutrophils by inducing necrosis of the latter (Van Gennip et al., 2009).
Until now the effect of all Rhls, including diRha-C10-C10 and its homologs, was attributed only to their biosurfactant activity, which destabilizes plasma membrane resulting to quantitative changes in phospholipid headgroup (Sha et al., 2012; Sotirova et al., 2012; Singh et al., 2013). Similarly to the role of dihydromaltophilin on plasma membrane sphingolipid biosynthesis (Li et al., 2009), the disorganization of the plasma membrane by diRhls may stimulate cell wall synthesis by activating cell wall integrity signaling pathways, which could explain the thickening of the cell wall. The combination of the rhamnobiose moiety and of the branched aglycon in the diRhls is essential for the interaction with the GS since neither rhamnobiose nor diRha-C3 and diRha-C8 analogs showed an inhibitory activity. Similar inhibition of the GS and growth phenotype in A. fumigatus has been observed using echinocandins, such as caspofungin, micofungin or anidulafungin, which are lipopeptides differentiated by their aliphatic tails (Perlin, 2011; Clavaud et al., 2012). Enzymatic removal of the aliphatic tails of the drugs renders them inactive (Perlin, 2011). The antagonistic effect of caspofungin and F-diRhls on A. fumigatus confirmed that both lipid conjugates had GS as target. It has been speculated that the tail of the drugs may intercalate into the bilayer resulting in inhibition. Such drug–target interactions would not require the drugs to enter the cell and they may act on the enzyme from the extracellular face of the cell membrane. At the same time, the fact that echinocandin-resistant strain (EMFR S678P) showed high susceptibility to F-diRhls suggested that the inhibitory binding site of F-diRhls differs from that of echinocandin and that the S678P mutation enhances the affinity of F-diRhls for GS. In contrast, the combination of azoles and F-diRhls on A. fumigatus was synergic as it was found previously for azoles and echinocandins, which have different inhibitory binding sites (Mavridou et al., 2015).
Other studies showed that in vitro P. aeruginosa inhibited the growth of A. fumigatus, the cystic fibrosis isolates being more inhibitory than non-cystic fibrosis isolates (Ferreira et al., 2015). Moreover, until now it is not known how A. fumigatus colonized better in the respiratory tract of cystic fibrosis patients when they got an infection with P. aeruginosa and whether the fungus is in close contact with the bacteria. We showed previously that at low ‘in vivo’ concentrations, phenazines stimulated the growth of A. fumigatus by providing iron uptake (Briard et al., 2015). It is particularly conceivable that in vivo A. fumigatus establishes in the same niches as P. aeruginosa taking advantage of favorable growth conditions due to lung damages. A. fumigatus may also benefit from the P. aeruginosa biofilm for host immune protection due to the role of Rhls and alginate in the reduction of the host immune responses (Baxter et al., 2013; Briard et al., 2015). Moreover, recent studies from our group also showed that these interactions can be observed at distance. Fungi and bacteria do not have to be in contact and volatiles from P. aeruginosa can be used as nutrients by the fungus (Briard et al., 2016). These recent data point out the need to investigate in depth the pulmonary microbiota and the cross-talks between bacteria and fungi. In cystic fibrosis patients, the microbiota is completely disorganized and becomes particularly rich in the pathogenic bacteria P. aeruginosa and A. fumigatus (Delhaes et al., 2012; Baxter et al., 2013; Armstead et al., 2014). Equilibrium in the different partners of the microbiota may be important to avoid allergic bronchopulmonary aspergillosis, A. fumigatus bronchitis and other allergic manifestations as well as controlling cystic fibrosis infections and chronic obstructive pulmonary disorders (Whiteson et al., 2014).
References
Abalos A, Pinazo A, Infante MR, Casals M, García F, Manresa A . (2001). Physicochemical and antimicrobial properties of new rhamnolipids produced by Pseudomonas aeruginosa AT10 from soybean oil refinery wastes. Langmuir 17: 1367–1371.
Abdel-Mawgoud AM, Lépine F, Déziel E . (2010). Rhamnolipids: diversity of structures, microbial origins and roles. Appl Microbiol Biotechnol 86: 1323–1336.
Arikan S, Lozano-Chiu M, Paetznick V, Rex JH . (2002). In vitro synergy of caspofungin and amphotericin B against Aspergillus and Fusarium spp. Antimicrob Agents Chemother 46: 245–247.
Armstead J, Morris J, Denning DW . (2014). Multi-country estimate of different manifestations of aspergillosis in cystic fibrosis. PloS One 9: e98502.
Arutchelvi J, Doble M . (2010). Characterization of glycolipid biosurfactant from Pseudomonas aeruginosa CPCL isolated from petroleum-contaminated soil. Lett Appl Microbiol 51: 75–82.
Bamford NC, Snarr BD, Gravelat FN, Little DJ, Lee MJ, Zacharias CA et al. (2015). Sph3 Is a glycoside hydrolase required for the biosynthesis of galactosaminogalactan in Aspergillus fumigatus. J Biol Chem 290: 27438–27450.
Baxter CG, Rautemaa R, Jones AM, Webb AK, Bull M, Mahenthiralingam E et al. (2013). Intravenous antibiotics reduce the presence of Aspergillus in adult cystic fibrosis sputum. Thorax 68: 652–657.
Beauvais A, Schmidt C, Guadagnini S, Roux P, Perret E, Henry C et al. (2007). An extracellular matrix glues together the aerial-grown hyphae of Aspergillus fumigatus. Cell Microbiol 9: 1588–1600.
Bjarnsholt T, Jensen PØ, Jakobsen TH, Phipps R, Nielsen AK, Rybtke MT et al. (2010). Quorum sensing and virulence of Pseudomonas aeruginosa during lung infection of cystic fibrosis patients. PloS One 5: e10115.
Blyth W . (1971). Modifications in the ultrastructure of Aspergillus fumigatus due to the presence of living cells of Pseudomonas aeruginosa. Sabouraudia 9: 283–286.
Briard B, Bomme P, Lechner BE, Mislin GLA, Lair V, Prévost M-C et al. (2015). Pseudomonas aeruginosa manipulates redox and iron homeostasis of its microbiota partner Aspergillus fumigatus via phenazines. Sci Rep 5: 8220.
Briard B, Heddergott C, Latgé J-P . (2016). Volatile compounds emitted by Pseudomonas aeruginosa stimulate growth of the fungal pathogen Aspergillus fumigatus. mBio 7: e00219.
Chamilos G, Akoumianaki T, Kyrmizi I, Brakhage A, Beauvais A, Latge J-P . (2016). Melanin targets LC3-associated phagocytosis (LAP): a novel pathogenetic mechanism in fungal disease. Autophagy 12: 888–889.
Christova N, Tuleva B, Kril A, Georgieva M, Konstantinov S, Terziyski I et al. (2013). Chemical structure and in vitro antitumor activity of rhamnolipids from Pseudomonas aeruginosa BN10. Appl Biochem Biotechnol 170: 676–689.
Clavaud C, Beauvais A, Barbin L, Munier-Lehmann H, Latgé J-P . (2012). The composition of the culture medium influences the β-1,3-glucan metabolism of Aspergillus fumigatus and the anjpegungal activity of inhibitors of β-1,3-glucan synthesis. Antimicrob Agents Chemother 56: 3428–3431.
Delhaes L, Monchy S, Fréalle E, Hubans C, Salleron J, Leroy S et al. (2012). The airway microbiota in cystic fibrosis: a complex fungal and bacterial community—implications for therapeutic management. PloS One 7: e36313.
Diggle SP, Winzer K, Chhabra SR, Worrall KE, Cámara M, Williams P . (2003). The Pseudomonas aeruginosa quinolone signal molecule overcomes the cell density-dependency of the quorum sensing hierarchy, regulates rhl-dependent genes at the onset of stationary phase and can be produced in the absence of LasR. Mol Microbiol 50: 29–43.
Eisenman HC, Casadevall A . (2012). Synthesis and assembly of fungal melanin. Appl Microbiol Biotechnol 93: 931–940.
Elefanti A, Mouton JW, Verweij PE, Tsakris A, Zerva L, Meletiadis J . (2013). Amphotericin B- and voriconazole-echinocandin combinations against Aspergillus spp.: effect of serum on inhibitory and fungicidal interactions. Antimicrob Agents Chemother 57: 4656–4663.
Ferreira JAG, Penner JC, Moss RB, Haagensen JAJ, Clemons KV, Spormann AM et al. (2015). Inhibition of Aspergillus fumigatus and its biofilm by Pseudomonas aeruginosa is dependent on the source, phenotype and growth conditions of the bacterium. PloS One 10: e0134692.
Frases S, Chaskes S, Dadachova E, Casadevall A . (2006). Induction by Klebsiella aerogenes of a melanin-like pigment in Cryptococcus neoformans. Appl Environ Microbiol 72: 1542–1550.
Gogoi D, Bhagowati P, Gogoi P, Bordoloi NK, Rafay A, Dolui SK et al. (2016). Structural and physico-chemical characterization of a dirhamnolipid biosurfactant purified from Pseudomonas aeruginosa: application of crude biosurfactant in enhanced oil recovery. RSC Adv 6: 70669–70681.
Gresnigt MS, Bozza S, Becker KL, Joosten LAB, Abdollahi-Roodsaz S, van der Berg WB et al. (2014). A polysaccharide virulence factor from Aspergillus fumigatus elicits anti-inflammatory effects through induction of Interleukin-1 receptor antagonist. PLoS Pathog 10: e1003936.
Haba E, Pinazo A, Jauregui O, Espuny MJ, Infante MR, Manresa A . (2003). Physicochemical characterization and antimicrobial properties of rhamnolipids produced by Pseudomonas aeruginosa 47T2 NCBIM 40044. Biotechnol Bioeng 81: 316–322.
Hill D, Rose B, Pajkos A, Robinson M, Bye P, Bell S et al. (2005). Antibiotic susceptibilities of Pseudomonas aeruginosa isolates derived from patients with cystic fibrosis under aerobic, anaerobic, and biofilm conditions. J Clin Microbiol 43: 5085–5090.
Hogan DA, Kolter R . (2002). Pseudomonas-Candida interactions: an ecological role for virulence factors. Science 296: 2229–2232.
Hogan DA, Vik A, Kolter R . (2004). A Pseudomonas aeruginosa quorum-sensing molecule influences Candida albicans morphology. Mol Microbiol 54: 1212–1223.
Holloway BW, Römling U, Tümmler B . (1994). Genomic mapping of Pseudomonas aeruginosa PAO. Microbiol Read Engl 140: 2907–2929.
Jahn B, Boukhallouk F, Lotz J, Langfelder K, Wanner G, Brakhage AA . (2000). Interaction of human phagocytes with pigmentless Aspergillus conidia. Infect Immun 68: 3736–3739.
Jensen PØ, Bjarnsholt T, Phipps R, Rasmussen TB, Calum H, Christoffersen L et al. (2007). Rapid necrotic killing of polymorphonuclear leukocytes is caused by quorum-sensing-controlled production of rhamnolipid by Pseudomonas aeruginosa. Microbiology 153: 1329–1338.
Knutsen AP, Bush RK, Demain JG, Denning DW, Dixit A, Fairs A et al. (2012). Fungi and allergic lower respiratory tract diseases. J Allergy Clin Immunol 129: 280–291-293.
Kownatzki R, Tümmler B, Döring G . (1987). Rhamnolipid of Pseudomonas aeruginosa in sputum of cystic fibrosis patients. Lancet 1: 1026–1027.
Laabei M, Jamieson WD, Lewis SE, Diggle SP, Jenkins ATA . (2014). A new assay for rhamnolipid detection-important virulence factors of Pseudomonas aeruginosa. Appl Microbiol Biotechnol 98: 7199–7209.
Lamarre C, Beau R, Balloy V, Fontaine T, Wong Sak Hoi J, Guadagnini S et al. (2009). Galactofuranose attenuates cellular adhesion of Aspergillus fumigatus. Cell Microbiol 11: 1612–1623.
Langfelder K, Philippe B, Jahn B, Latgé JP, Brakhage AA . (2001). Differential expression of the Aspergillus fumigatus pksP gene detected in vitro and in vivo with green fluorescent protein. Infect Immun 69: 6411–6418.
Latgé J-P . (2010). Tasting the fungal cell wall. Cell Microbiol 12: 863–872.
Lee KK, Maccallum DM, Jacobsen MD, Walker LA, Odds FC, Gow NAR et al. (2012). Elevated cell wall chitin in Candida albicans confers echinocandin resistance in vivo. Antimicrob Agents Chemother 56: 208–217.
Li S, Calvo AM, Yuen GY, Du L, Harris SD . (2009). Induction of cell wall thickening by the anjpegungal compound dihydromaltophilin disrupts fungal growth and is mediated by sphingolipid biosynthesis. J Eukaryot Microbiol 56: 182–187.
Mavridou E, Meletiadis J, Rijs A, Mouton JW, Verweij PE . (2015). The strength of synergistic interaction between posaconazole and caspofungin depends on the underlying azole resistance mechanism of Aspergillus fumigatus. Antimicrob Agents Chemother 59: 1738–1744.
Moree WJ, Phelan VV, Wu C-H, Bandeira N, Cornett DS, Duggan BM et al. (2012). Interkingdom metabolic transformations captured by microbial imaging mass spectrometry. Proc Natl Acad Sci USA 109: 13811–13816.
Mowat E, Rajendran R, Williams C, McCulloch E, Jones B, Lang S et al. (2010). Pseudomonas aeruginosa and their small diffusible extracellular molecules inhibit Aspergillus fumigatus biofilm formation. FEMS Microbiol Lett 313: 96–102.
Paugam A, Baixench M-T, Demazes-Dufeu N, Burgel P-R, Sauter E, Kanaan R et al. (2010). Characteristics and consequences of airway colonization by filamentous fungi in 201 adult patients with cystic fibrosis in France. Med Mycol 48: S32–S36.
Perlin DS . (2011). Current perspectives on echinocandin class drugs. Future Microbiol 6: 441–457.
Ram AFJ, Klis FM . (2006). Idenjpegication of fungal cell wall mutants using susceptibility assays based on Calcofluor white and Congo red. Nat Protoc 1: 2253–2256.
Robinet P, Baychelier F, Fontaine T, Picard C, Debré P, Vieillard V et al. (2014). A polysaccharide virulence factor of a human fungal pathogen induces neutrophil apoptosis via NK cells. J Immunol 192: 5332–5342.
Rocha EMF, Garcia-Effron G, Park S, Perlin DS . (2007). A Ser678Pro substitution in Fks1p confers resistance to echinocandin drugs in Aspergillus fumigatus. Antimicrob Agents Chemother 51: 4174–4176.
Schmaler-Ripcke J, Sugareva V, Gebhardt P, Winkler R, Kniemeyer O, Heinekamp T et al. (2009). Production of pyomelanin, a second type of melanin, via the tyrosine degradation pathway in Aspergillus fumigatus. Appl Environ Microbiol 75: 493–503.
Schneider CA, Rasband WS, Eliceiri KW . (2012). NIH Image to ImageJ: 25 years of image analysis. Nat Met 9: 671–675.
Schmidberger A, Henkel M, Hausmann R, Schwartz T . (2014). Influence of ferric iron on gene expression and rhamnolipid synthesis during batch cultivation of Pseudomonas aeruginosa PAO1. Appl Microbiol Biotechnol 98: 6725–6737.
Sha R, Jiang L, Meng Q, Zhang G, Song Z . (2012). Producing cell-free culture broth of rhamnolipids as a cost-effective fungicide against plant pathogens. J Basic Microbiol 52: 458–466.
Sharma A, Jansen R, Nimtz M, Johri BN, Wray V . (2007). Rhamnolipids from the rhizosphere bacterium Pseudomonas sp. GRP(3) that reduces damping-off disease in Chilli and tomato nurseries. J Nat Prod 70: 941–947.
da Silva Ferreira ME, Kress MRVZ, Savoldi M, Goldman MHS, Härtl A, Heinekamp T et al. (2006). The akuB(KU80) mutant deficient for nonhomologous end joining is a powerful tool for analyzing pathogenicity in Aspergillus fumigatus. Eukaryot Cell 5: 207–211.
Singh N, Pemmaraju SC, Pruthi PA, Cameotra SS, Pruthi V . (2013). Candida biofilm disrupting ability of di-rhamnolipid (RL-2) produced from Pseudomonas aeruginosa DSVP20. Appl Biochem Biotechnol 169: 2374–2391.
Sotirova A, Avramova T, Stoitsova S, Lazarkevich I, Lubenets V, Karpenko E et al. (2012). The importance of rhamnolipid-biosurfactant-induced changes in bacterial membrane lipids of Bacillus subtilis for the antimicrobial activity of thiosulfonates. Curr Microbiol 65: 534–541.
Sugareva V, Härtl A, Brock M, Hübner K, Rohde M, Heinekamp T et al. (2006). Characterisation of the laccase-encoding gene abr2 of the dihydroxynaphthalene-like melanin gene cluster of Aspergillus fumigatus. Arch Microbiol 186: 345–355.
Toljander JF, Lindahl BD, Paul LR, Elfstrand M, Finlay RD . (2007). Influence of arbuscular mycorrhizal mycelial exudates on soil bacterial growth and community structure. FEMS Microbiol Ecol 61: 295–304.
Van Gennip M, Christensen LD, Alhede M, Phipps R, Jensen PØ, Christophersen L et al. (2009). Inactivation of the rhlA gene in Pseudomonas aeruginosa prevents rhamnolipid production, disabling the protection against polymorphonuclear leukocytes. APMIS Acta Pathol Microbiol Immunol Scand 117: 537–546.
Walker LA, Gow NAR, Munro CA . (2010). Fungal echinocandin resistance. Fungal Genet Biol 47: 117–126.
Whiteson KL, Bailey B, Bergkessel M, Conrad D, Delhaes L, Felts B et al. (2014). The upper respiratory tract as a microbial source for pulmonary infections in cystic fibrosis. Parallels from island biogeography. Am J Respir Crit Care Med 189: 1309–1315.
Wilson R, Sykes DA, Watson D, Rutman A, Taylor GW, Cole PJ . (1988). Measurement of Pseudomonas aeruginosa phenazine pigments in sputum and assessment of their contribution to sputum sol toxicity for respiratory epithelium. Infect Immun 56: 2515–2517.
Acknowledgements
Research in the Aspergillus Unit was supported by the Association Vaincre La Mucoviscidose (RF20140501052/1/1/141). We thank V Aimanianda (Aspergillus Unit, Institut Pasteur, Paris, France) for language editing. We thank A Mallet (Ultrapole, Institut Pasteur, Paris, France) for her help in scanning electron microscopy.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Briard, B., Rasoldier, V., Bomme, P. et al. Dirhamnolipids secreted from Pseudomonas aeruginosa modify anjpegungal susceptibility of Aspergillus fumigatus by inhibiting β1,3 glucan synthase activity. ISME J 11, 1578–1591 (2017). https://doi.org/10.1038/ismej.2017.32
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.32
This article is cited by
-
Influence of relevant cystic fibrosis bacteria on Scedosporium apiospermum and Scedosporium boydii growth and viability
Brazilian Journal of Microbiology (2021)
-
Galactosaminogalactan activates the inflammasome to provide host protection
Nature (2020)
-
Fungal ligands released by innate immune effectors promote inflammasome activation during Aspergillus fumigatus infection
Nature Microbiology (2018)
-
Small Colony Variants of Pseudomonas aeruginosa Display Heterogeneity in Inhibiting Aspergillus fumigatus Biofilm
Mycopathologia (2018)