Abstract
Dinoroseobacter shibae, a member of the Roseobacter clade abundant in marine environments, is characterized by a pronounced pleomorphism. Cell shapes range from variable-sized ovoid rods to long filaments with a high copy number of chromosomes. Time-lapse microscopy shows cells dividing either by binary fission or by budding from the cell poles. Here we demonstrate that this morphological heterogeneity is induced by quorum sensing (QS). D. shibae utilizes three acylated homoserine lactone (AHL) synthases (luxI1–3) to produce AHLs with unsaturated C18 side chains. A ΔluxI1-knockout strain completely lacking AHL biosynthesis was uniform in morphology and divided by binary fission only. Transcriptome analysis revealed that expression of genes responsible for control of cell division was reduced in this strain, providing the link between QS and the observed phenotype. In addition, flagellar biosynthesis and type IV secretion system (T4SS) were downregulated. The wild-type phenotype and gene expression could be restored through addition of synthetic C18-AHLs. Their effectiveness was dependent on the number of double bonds in the acyl side chain and the regulated trait. The wild-type expression level of T4SS genes was fully restored even by an AHL with a saturated C18 side chain that has not been detected in D. shibae. QS induces phenotypic individualization of D. shibae cells rather than coordinating the population. This strategy might be beneficial in unpredictably changing environments, for example, during algal blooms when resource competition and grazing exert fluctuating selective pressures. A specific response towards non-native AHLs might provide D. shibae with the capacity for complex interspecies communication.
Similar content being viewed by others
Introduction
The term ‘quorum sensing’ (QS) refers to a form of cell-to-cell communication that involves the production, excretion and detection of small diffusible signalling molecules called autoinducers (AI). The simplest QS systems consist of an enzyme for AI biosynthesis and a transcription factor that is activated by AI binding and induces the expression of a defined set of genes. Many bacteria make use of more complex systems involving two or more types of AI molecules and different network architectures (Waters and Bassler, 2005). QS has first been described in the marine bacterium Vibrio fischeri colonizing the squid light organ. When cell density reaches a certain threshold, the ‘quorum’, a burst of AI synthesis coordinates the population-wide activation of genes required for bioluminescence (Nealson and Hastings, 1979). Coordination of gene expression as a function of cell density was considered a paradigm for a long time but has been challenged in recent years.
The concept of QS activity being dependent on cell density has been extended to account for the fact that outside the shaking flask AI concentration is also influenced by diffusion (for example, enclosures, microniches, local gradients in biofilms, microcolonies) (Kaplan and Greenberg, 1985), its stability (for example, in dependence of pH) (Wang and Leadbetter, 2005) and various other parameters (Platt and Fuqua, 2010). One striking example comes from Schäfer et al. (2008) showing that Rhodopseudomonas palustris synthesizes an AI by incorporating a molecule from decaying plant material. They concluded that in this strain QS is dependent on both the cell density and the availability of an exogenously supplied substrate.
The concept of QS mediating the population-wide coordination of gene expression has been challenged, too, by the demonstration of a heterogeneous response towards QS signals in several organisms (Bassler and Losick, 2006). In Streptococcus pneumoniae, QS through a peptide AI leads to the induction of the competent state only in a fraction of cells, whereas the remainder of the population undergoes autolysis (Steinmoen et al., 2002), a phenomenon termed fratricide (Gilmore and Haas, 2005). A similar phenomenon was observed in Streptococcus mutans. Lemme et al. (2011) used fluorescence-activated cell sorting (FACS) to separate induced and uninduced cells, identifying major transcriptome differences between both subpopulations. For the Gram-negative bacterium Vibrio harveyi, it was shown that the wild type, displaying a heterogeneous QS response regarding bioluminescence, produces more biofilm than a constitutive QS-active mutant (Anetzberger et al., 2009). Recently, a high variability of other QS-regulated traits like secretion systems and exoproteolysis was demonstrated for the same organism (Anetzberger et al., 2012). Heterogeneous expression of QS genes and controlled traits have also been shown for Listeria monocytogenes (Garmyn et al., 2011) and Vibrio fischeri, the model organism for coordinated gene expression through communication (Perez and Hagen, 2010; Perez et al., 2011).
This paradigm shift is by far not restricted to QS. In the last decades, it became more and more apparent that physiological heterogeneity is a fundamental characteristic of isogenic bacterial populations (but not restricted to bacteria). It is mediated by various processes, namely stochastic gene expression, unequal distribution of molecules during cell division, ageing, and bi- or multistable gene-regulatory networks (reviewed in Kaern et al. (2005); (Avery, 2006; Smits et al., 2006)). Population heterogeneity is thought to be a survival strategy in fluctuating and unpredictable environments (Acar et al., 2008). For example, starved Sinorhizobium meliloti differentiates into cells with low and high poly-3-hydroxybutyrate levels that have higher competitiveness for resources and long-term survival capabilities, respectively (Ratcliff and Denison, 2010).
D. shibae is a representative of the Roseobacter clade, a large, diverse and ecologically important phylogenetic cluster of Alphaproteobacteria (Wagner-Döbler and Biebl, 2006; Brinkhoff et al., 2008), abundant in temperate and polar marine habitats (Selje et al., 2004; Giebel et al., 2011). It was isolated from the dinoflagellate Prorocentrum lima (Biebl et al., 2005) and lives in symbiosis with marine algae (Wagner-Döbler et al., 2010). Under optimal growth conditions a remarkable variability in cell size and morphology can be observed. The physiological role of the morphological heterogeneity and the mechanism by which it is controlled are unknown.
D. shibae, like many other Proteobacteria, relies on acylated homoserine-lactones (AHLs) for cell-to-cell communication. It produces C18en-HSL and C18dien-HSL with one and two unsaturations in the acyl side chain, respectively, which represent novel structures not described in any other bacterial species so far (Wagner-Döbler et al., 2005). Genome analysis of D. shibae revealed the presence of three LuxI type AHL synthase genes (termed luxI1, luxI2, luxI3). luxI1 and luxI2 are located on the chromosome downstream of a gene encoding a LuxR-type transcriptional regulator, whereas luxI3 is on the 86-kb plasmid without an adjacent luxR gene. In addition, three orphan LuxR-type transcriptional regulator genes were found in the genome (Wagner-Döbler et al., 2010).
Here, we studied the role of the novel long-chain AHLs (C18en-HSL and C18dien-HSL) produced by D. shibae. We constructed a ΔluxI1 knockout strain and found that it does not produce any AHLs; thus it represents a QS signal synthesis null mutant. Without the ability to communicate, D. shibae does not differentiate into morphologically distinct cell types. We investigated the link between QS and cellular heterogeneity by comparison of the wild type and the QS null strain as well as the genetically complemented mutant using electron microscopy, determination of chromosome copy number, time-lapse microscopy and cell-density-resolved transcriptome analysis. To study the role of distinct AHLs, we added C18-HSLs with different numbers of unsaturations to the culture of the ΔluxI1 mutant.
Materials and methods
Additional Materials and methods can be found in Supplementary Text S1.
Detection of AHLs
The production of AHLs was detected using the biosensor strains E. coli MT102 pJBA132 and P. putida F117 pKR-C12 as previously described (Wagner-Döbler et al., 2005), with the following modifications. D.shibae strains were grown in 100 ml defined minimal medium with 50 mM succinate and 2% of adsorber resins (Amberlite XAD-16, Rohm & Haas, Philadelphia, PA, USA) for 36 h at 30 °C with agitation. Adsorber resins were removed from the culture by filtration, added to a separating funnel together with 25 ml methylene chloride and 200 ml distilled water, and well shaken. The organic phase was removed. The methylene chloride extraction was repeated three times, with a total volume of 100 ml solvent. Extracts were concentrated to 2 ml using a rotary evaporator (Heidolph VV2001, Kelheim, Germany) and stored at −20 °C. For bioassays, 10 μl was pipetted into polypropylene microtitre plates (PlateOne, Starlab, Hamburg, Germany). When the methylene chloride had evaporated, 100 μl medium and 100 μl of the respective sensor strain were added. Microtitre plates were incubated at 30 °C with agitation for 30 h.
Microarray analysis
Processing of microarray data is described in Supplementary Text S1. For analysis of differential expression, only those genes with a false discovery rate-adjusted P-value <0.01 and an absolute log2-fold change >1 under at least one condition were taken into account. Raw and processed microarray data have been deposited at the gene expression omnibus database under the accession number GSE42013.
Flow cytometry and FACS
For flow cytometry and cell sorting, 1-ml samples were collected from cultures at the desired cell density, and cells were fixed for 15 min by addition of 2% glutaraldehyde. Fixed samples were transferred to liquid nitrogen and stored at −20 °C. Before measurements cells were diluted 100-fold when the optical density was below 0.4 and diluted 1000-fold when above 0.4 in sterile filtered PBS buffer (pH 7.4). For stoichiometric DNA staining, 10 μl 100 × SYBR Green solution (Molecular Probes, Leiden, The Netherlands) was added to 1 ml sample and incubated for 20 min in the dark (Marie et al., 1997). For each sample a minimum of 50 000 cells were analyzed on a FACSCanto flow cytometer (BD Bioscience) to follow SYBR Green fluorescence and a FACSAriaII (BD Bioscience, Heidelberg, Germany) for cell sorting. Fluorescent signals were collected logarithmically using an FITC filter (excitation 488 nm, emission 519 nm). The sorting strategy is shown in Supplementary Figure S3. Data processing and analysis were carried out using the ‘flowCore’ package (Hahne et al., 2009) of the R BioConductor project. As the exact number of chromosomes in D. shibae cells is not known, chromosome content was defined as chromosome equivalents in relation to the SybrGreen peak with the lowest intensity.
Time-lapse microscopy
Time-lapse microscopy was performed using an automated microscope (Zeiss Axiovert 200) using the heating system 6 incubator and controller (Ibidi, Martinsried, Germany). For live-cell microscopy of growing microcolonies the agarose pad method was applied as described in Young et al. (2012) using SWM medium pads in 35 mm μ-dishes (Ibidi) at an incubation temperature of 30 °C. Subsequent image analyses were performed with Axiovision (Zeiss, Jena, Germany) and the TLM-Tracker software (Braunschweig, Germany; Klein et al., 2012).
Results
We first investigated whether the three identified luxI genes of D. shibae encode functional autoinducer synthases capable of synthesizing AHLs. The constitutive heterologous expression of each synthase in a non-AHL-producing E. coli resulted in the production of long-chain AHLs (Supplementary Figure S1). GC–MS analysis (Neumann et al., 2013) showed that when expressed in E. coli, the main compound produced by LuxI1 was the wild-type signal C18en-HSL. In addition, small amounts of C16-HSL, C15-HSL and C14-HSL were found. LuxI2 expressed in E. coli yielded C14-HSL variants as well as C15-HSL. LuxI3 did not produce any AHLs when cloned into pBBR1MCS-2, but synthesized C14-HSL and 3-oxo-C14-HSL when cloned into the hyperexpression vector pTrcHis-TOPO. Compared with the wild type, the chain lengths and types of substitutions found through heterologous expression of the AHL synthases in E. coli clearly differed from those produced in D. shibae.
For example, the main wild-type signal C18dien-HSL was not produced in E. coli. By contrast, C15-HSL was never detected in D. shibae. These differences may be due to the different abundance of fatty acid precursors in the producing organism or lack of specific precursors.
In order to unravel the role of the autoinducer synthase LuxI1 and to identify QS-regulated traits in D. shibae, we constructed a ΔluxI1 strain by replacing 425 bp of the coding sequence with a gentamicin-resistance cassette via homologous recombination. Integration of the knockout cassette at the luxI1 locus was verified by PCR and sequencing. The luxI1 deletion was genetically complemented with the plasmid pDP1containing the luxI1 open reading frame controlled by the high-level-expression gentamicin promoter. Additionally, a control strain carrying the empty plasmid was generated.
Deletion of luxI1 completely eliminates AHL production in D. shibae
Previously it had been shown that D. shibae synthesizes C18en-HSL and C18dien-HSL in MB medium (Wagner-Döbler et al., 2005, 2010). This was now also confirmed in the minimal medium used for microarray analyses by GC–MS (Supplementary Figure S2). The previously detected C8-HSL was not found. Extracts of the ΔluxI1 strain showed no significant induction of fluorescence in the sensor strain used for the detection of long-chain AHLs (Figure 1a). GC–MS analysis of the extracts confirmed the complete absence of AHLs in the ΔluxI1 strain (data not shown). As the deletion of only one synthase leads to a complete loss of AHL production, the contribution of LuxI2 and LuxI3 to AHL synthesis in D. shibae appears to be dependent on the presence and level of the signal molecules provided by LuxI1. The wild-type signals C18en-HSL and C18dien-HSL were strongly overproduced in the D. shibae ΔluxI1 strain carrying pDP1, and thus it was possible to purify these novel signals and determine their absolute configuration (Neumann et al., 2013). Both C18en-HSL and C18dien-HSL were then chemically synthesized and their effect on the phenotype and transcriptome of the ΔluxI1 deletion strain was studied.
Production of long-chain AHLs (a) and growth (b) of D. shibae wild-type, ΔluxI1 and ΔluxI1 pDP1. (a) The maximum fluorescence induced in the sensor strain P. putida pKR-C12 after incubation with AHLs extracted from the indicated D. shibae strains and positive controls (pure AHLs at a concentration of 2.4 μM) is displayed. Error bars indicate the s.d. of triplicate measurements. (b) Cell counts of the indicated D. shibae strains in minimal medium at 30 °C with continuous shaking were determined in three biological replicates using flow cytometry.
QS affects the growth rate of D. shibae
The growth of the ΔluxI1 mutant differed from the wild type in various aspects: the mutant showed a shorter lag phase, higher growth rate and higher maximum cell density than the parent strain (Figure 1b and Table 1). Genetic complementation of the luxI1 deletion restored the wild-type growth behavior, whereas the presence of the ‘empty’ vector pBBR1MCS-2 had no significant impact on the growth of the mutant.
In order to study the response of the non-AHL-producing mutant ΔluxI1 to AHLs, synthetic signals C8-, C18en- and C18dien-HSL as well as C18-HSL were added to ΔluxI1 cultures at final concentrations of 500 nM. The C18-HSL has not been detected in D. shibae, neither in rich medium nor in minimal medium (Neumann et al., 2013). Moreover, all of the AHLs found in D. shibae cultures have one or two double bonds in the acyl side chain. We therefore assume that the saturated AHL C18-HSL is a non-native signal in D. shibae.
In the presence of C8-HSL we did not observe any effect on the mutant’s growth behavior (Supplementary Table S2). Growth rate and doubling time were identical to the ΔluxI1 culture. The data are in accordance with the lack of detection of C8-HSL, showing that D. shibae neither produces nor responds to this signal. The growth rate was affected in different ways by the different C18-HSLs (Supplementary Table S2). The non-native signal C18-HSL did not affect the growth rate and doubling time of the ΔluxI1 mutant. By contrast, C18en-HSL and C18dien-HSL reduced the growth rate even below that of the wild type.
Loss of QS signaling results in homogenous cell size, reduced chromosome content and cell division exclusively by binary fission
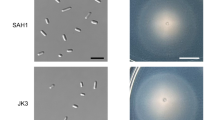
Through investigation of the ΔluxI1mutant by scanning electron microscopy, we observed altered cell morphology (Figure 2a). Wild-type cultures exhibited heterogeneous cell morphology with respect to cell shape and size. They were composed of ovoid and rod-shaped cells of different sizes and eye-catching elongated cells reaching up to 10 μm length. Cells of D. shibae ΔluxI1 were homogeneous in size and morphology. Genetic complementation restored the wild-type morphotypes. Closer examination suggested that wild-type cells were using different types of cell division (Figure 2b) that could not be observed in the mutant. To further investigate the heterogeneity in the wild-type population and to examine whether the elongated morphotype of the subpopulation might be caused by different growth and division behavior, we determined the relative chromosome content at various cell densities on the single-cell level using stoichiometric SybrGreen staining and subsequent flow cytometric analysis, assuming that chromosomes accumulate in elongated cells. This technique provides an elegant tool to study the cell cycle and the DNA replication pattern (Müller, 2007) given that the chromosome content correlates with the fluorescence intensity. As the absolute number of chromosomes per cell is not known, the term ‘chromosome equivalent’ is used.
Morphological characteristics of D. shibae wild-type, ΔluxI1 and ΔluxI1 pDP1. Scanning electron micrographs demonstrate the morphological heterogeneity in the wild-type and genetically complemented strain. D. shibae ΔluxI1 shows a uniform cell-size distribution. For each strain one overview (scale bar: 5 μm) (a) and two detailed pictures (of 10 μm width) are shown (b). Arrows indicate sites of beginning cell division. (c) Distribution of log2 SybrGreen intensity measured through the FITC channel of the flow cytometer (FITC-A) and displayed as the frequency of cells showing a distinct intensity. The stoichiometric DNA staining revealed two peaks for one (C1n) and two (C2n) chromosome equivalents, respectively, in all strains. The wild-type and genetically complemented strain showed a long tail of cells with higher fluorescence (Cxn), indicating more than 2 chromosome equivalents per cell.
D. shibae wild type showed two distinct peaks representing two different cell fractions with one (C1n) and two chromosome equivalents (C2n) per cell, respectively (Figure 2c). However, a small fraction contained multiple chromosome equivalents per cell (Cxn). Sorting of the wild-type cells according to their chromosome content and subsequent microscopic investigation (Supplementary Figures S3 and S4) confirmed that the C1n fraction consisted of small ovoid cells, the C2n fraction contained cells dividing by binary fission and the fraction containing more than two chromosome equivalents was comprised of elongated cells. Only cells with one or two chromosome equivalents were observed in the QS null mutant ΔluxI1 (Figure 2c). These data further suggest that subpopulations with different replication and cell division patterns coexist in one wild-type culture of D. shibae, which are dependent on the LuxI1-produced AHLs.
Flow cytometric investigation of the chromosome distribution at the mid-exponential growth phase (OD 0.4) in mutant populations supplemented with the different long-chain AHLs revealed a graduated complementation pattern, similar to that described above for growth. The results are presented in Figure 3. The D. shibae wild-type population was composed of 30.75% cells containing one chromosome equivalent (C1n), 50.5% harboring two equivalents (C2n) and 18.75% carrying multiple chromosome copies (Cxn). The homogeneous mutant culture consisted of 49.85% C1n cells, 49.1% C2n cells and 1.05% cells belonging to the Cxn fraction. The same distribution was observed in DMSO-treated mutant cells, which served as a negative control. The addition of saturated C18-HSL resulted in a slight increase in Cxn cells (2.4%). However, in the presence of C18en-HSL and C18dien-HSL, the distribution of chromosome equivalents of the ΔluxI1culture was shifted to the wild-type pattern almost completely.
Relative proportion of cells with one, two or more chromosome equivalents in D. shibae, the ΔluxI1 deletion mutant, the genetically complemented strain and chemically complemented cultures. Each bar represents the relative proportion of the three cell types (C1n, C2n, Cxn) in a culture of the indicated strains. D. shibae ΔluxI1, D. shibae ΔluxI1 cultivated with DMSO (negative control) or cultivated with the respective AHL, D. shibae wild-type and genetically complemented mutant strain D. shibae ΔluxI1 pDP1.
In vivo analysis using time-lapse microscopy confirmed that wild-type cells employ different modes of cell division. Figure 4a demonstrates that elongated cells divide by forming one substantially smaller daughter cell through polar growth. We define this type of cell division as budding. In contrast, small ovoid rods divide into two equally sized daughter cells; thus they employ binary fission. In Figure 4b we highlight a cell that buds from alternating cell poles before it divides into three daughter cells. The QS mutant employs exclusively binary fission (Figure 4c). The full movies of wild-type, mutant and complemented strain can be found as Supplementary Movies S1–S3.
Growth and division of individual D. shibae wild-type and ΔluxI1 cells visualized by time-lapse microscopy. (a) Symmetric (green) and asymmetric cell division (blue, red) co-occur in D. shibae wild-type cultures. (b) Example of a wild-type cell (highlighted blue in the lower schematic) showing alternating budding from both cell poles. (c) QS null mutant cells dividing by binary fission. Daughter cells are indicated by novel colors. Scale bar represents 5 μm. Corresponding movies can be found in Supplementary Movies S1–S3.
Comparative transcriptome analysis of D. shibae wild-type, ΔluxI1 and ΔluxI1 pDP1
To gain insights into the effects of QS on transcriptional control, two different experiments were performed: gene expression in ΔluxI1 and ΔluxI1 pDP1, respectively, was compared with the wild type, with samples being taken at different culture densities (OD600 0.1, 0.2, 0.4, 0.6 and 0.8) in the exponential phase as well as in the stationary phase (6 h after the strains reached their maximum OD600). To investigate in depth the capability of C18en-HSL and C18dien-HSL to act as signaling molecules responsible for specific regulation of gene expression, we analyzed the transcriptome profiles of mutant cultures supplemented with those AHLs in comparison with ΔluxI1and the wild type. Furthermore, the influence of non-native C18-HSL was studied on the transcriptome level. Cultures in the mid-exponential growth phase supplemented with 500 nM of AHL were used for this experiment. Compared with the wild type, 344 genes were differentially expressed in the ΔluxI1 mutant throughout growth. These genes were clustered into five groups according to their expression changes during growth (Supplementary Table S3 and Supplementary Figure S5). Interestingly, we did not observe strong density-dependent expression profiles for most of the genes. Instead, gene expression differed between wild type and mutant throughout exponential growth; additionally, large differences were observed in the stationary phase. One hundred and thirty-three genes showed a significant differential regulation in both exponential and stationary phase. In all 68 genes were differentially expressed exclusively in the exponential, whereas 143 genes exclusively in the stationary phase. The expression of only 59 genes was increased, whereas all other genes showed decreased expression in the QS null mutant.
D. shibae wild-type gene expression was also compared with that of the ΔluxI1 mutant complemented with pDP1 (Supplementary Table S3). Three hundred and twenty-six genes displayed a significant change in expression when all samples were taken into account. However, major changes occurred only in the late exponential and stationary phase, possibly reflecting overexpression of the luxI1 gene. The wild-type expression level of 255 out of 344 genes differentially expressed in the ΔluxI1 mutant was successfully restored in the complemented strain. Eighty-nine genes showed differential expression, in most cases an inverse regulation compared with the mutant, consistent with the overexpression of luxI1. Thus, the microarray data are consistent with the observed restoration of the wild-type phenotype in the genetically complemented strain.
Two alternative sigma factors, rpoHI (Dshi_2978) and rpoHII (Dshi_2609), were downregulated in the mutant, with the minimum at the beginning and in the stationary phase, respectively. Expression of one anti-sigma factor and its respective antagonist (Dshi_0072/73) was strongly reduced (Supplementary Table S3). Remarkably, 45% of all genes differentially regulated in the ΔluxI1 mutant encoded hypothetical proteins. This is a large fraction compared with 28% of all genes in the genome. Sixty-four genes encode proteins with a predicted signal peptide but no transmembrane domains; thus, they might represent secreted factors. Almost all of them are hypothetical proteins. In the following sections, the four major differentially regulated traits will be discussed in detail.
LuxI and luxR type QS genes
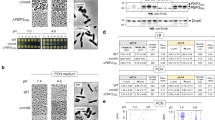
The gene expression of the cognate luxR1 regulator (Dshi_0311) was not affected by deletion of luxI1, indicating that it is independent from the AHL produced by the neighboring synthase. The second luxR2/I2 pair of genes (Dshi_2852/1) and – to a lesser extent – the orphan synthase luxI3 (Dshi_4152), however, were downregulated in the mutant, indicating that the AHLs synthesized by LuxI1 might be necessary for their activation (Figure 5a(1)). The three orphan LuxR type transcriptional regulators (Dshi_1550/1815/1819) in contrast displayed no significant change in expression. The overexpression of luxI1 in trans led to a slight overexpression of luxR2/I2 and restoration of luxI3 wild-type expression level (Figure 5a(2)). The addition of chemically synthesized AHLs to cultures of D. shibae ΔluxI1 re-established the wild-type expression level of luxR2/I2 for all three compounds tested. In contrast, only C18dien-HSL was able to restore the expression of luxI3 in the QS null mutant (Figure 5a(3)). These microarray data for representative samples were confirmed by qRT-PCR, which additionally showed complete lack of expression of luxI1 in the mutant (Supplementary Figure S6).
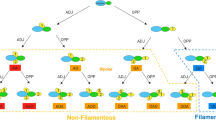
Transcriptome analysis of the quorum sensing null mutant. Heatmap visualization of log2 fold changes between samples. 1. D. shibae ΔluxI1 compared to wild type during exponential growth and in stationary phase. 2. D. shibae ΔluxI1 pDP1 compared to wild type (same sampling points). 3. D. shibae ΔluxI1 supplemented with 500 nM C18-HSL, C18en-HSL and C18dien-HSL and ΔluxI1 compared to wild type. Samples were taken at OD600 0.4. (a) LuxI autoinducer synthase and associated luxR-type transcriptional regulator genes. (b) Genes of the core cell-division-related control system in D. shibae. (c) Operons encoding flagella biosynthesis genes. (d) Plasmid-encoded Type IV secretion system genes.
Cell cycle-related genes
Cell cycle regulation has been exhaustively studied in the Alphaproteobacterium Caulobacter crescentus. This organism is characterized by a dimorphic lifestyle controlled by a complex gene-regulatory network with the histidine kinases CckA and ChpT (Biondi et al., 2006) and the transcription factor CtrA as the main components of the regulatory cascade (Purcell et al., 2008). A recent comparative genome analysis revealed that most Alphaproteobacteria share a common core set of regulators with differing accessory elements (Brilli et al., 2010). Like in Rodobacter sphaeroides, Roseobacter denitrificans and Ruegeria pomeroyi, a core of nine genes is also present in D. shibae. Only five of those were affected by alterations in the QS system (Figure 5b(1)). The luxI1 deletion led to a reduced expression of cckA (Dshi_1644), chpT (Dshi_1470) and ctrA (Dshi_1508). DivL (Dshi_3346), a target gene of CtrA with unknown function in Rhodobacterales, was downregulated too. The transcription factor DnaA, responsible for the initiation of DNA replication (Dshi_3373), was significantly downregulated during exponential growth and upregulated in the stationary phase; however, the log2-fold change (∼−0.7) was below the cutoff used. The two Clp proteases controlling the protein level of CtrA in C. crescentus (Dshi_1387/1388) did not change in expression in the ΔluxI1mutant. The transcription factor GcrA (Dshi_2616) and the DNA-methyltransferase CcrM (Dshi_0024) that activates DnaA promoter regionspriya through methylation were also unchanged. When luxI1 was overexpressed by introducing pDP1 into the ΔluxI1 mutant, the expression of these genes was fully restored (Figure 5b(2)). The chemical complementation with different AHLs revealed a graduated response (Figure 5b(3)). Only C18dien-HSL was able to fully restore the wild-type expression level. Interestingly, only ctrA and divL responded to the addition of C18-HSL and C18en-HSL. In summary, restoration of gene expression increased with the number of double bonds in the AHL side chain. To gain a better understanding how changes in cell cycle-related gene expression might act globally, we then searched for binding motives for CtrA in the promoters of D. shibae genes (Supplementary Figure S7). We identified 74 genes on the chromosome and 8 genes on the plasmids with CtrA binding sites. However, not all of them were differentially regulated in the ΔluxI1 mutant. The presence of two CtrA-binding sites in the promoter of the regulator luxR2 suggests a crosstalk between the QS and cell cycle control systems. Furthermore, a CtrA binding site was present in the promoter of rpoHII, indicating a link between cell cycle regulation and stress response. It seems plausible that the changes in the expression of the aforementioned genes may result in the observed differences in the chromosome content and cell division between wild-type and mutant strain.
Flagellar biosynthesis
D. shibae has a polar flagellum, which is encoded by three gene clusters (Dshi_3246-3268; 3358-3365, 3376-3380). Expression of the complete flagellar biosynthesis machinery was reduced in the ΔluxI1 mutant at all studied optical densities (Figure 5c(1)), with the maximum reduction occurring in the stationary phase. Overexpression of luxI1 in trans restored the wild-type expression level (Figure 5c(2)). Accordingly, flagella could not be detected in the QS null mutant using flagella staining or transmission electron microscopy, whereas flagellation was observed in wild-type cultures as well as in the genetically complemented strain (Supplementary Figure S8). Like for the cell cycle regulation genes, the flagellar synthesis gene expression showed a graduated response to exogenous AHLs dependent on the number of double bonds in the acyl side chain (Figure 5c(3)), and flagellation of ΔluxI1 was observed when AHLs were provided (Supplementary Figure S9).
Type IV secretion system
D. shibae contains two vir gene clusters, one on the 191 kb plasmid and the other on the 126 kb plasmid. Those two plasmids have been described as sister plasmids derived from a common ancestor (Wagner-Döbler et al., 2010). Accordingly, the two vir gene clusters are virtually identical, comprising the complete set of genes for the type IV secretion machinery for translocation of DNA or proteins (virB1 to virB11) (Christie et al., 2005). The 191-kb plasmid carries altogether 198 genes; in addition to the 14 genes of the vir-operon, only 11 other genes were regulated in the QS null mutant. Similarly, the 126-kb plasmid carries 136 genes. In addition to the 13 genes of the vir-operon, 6 other genes were differentially expressed in the mutant. All other genes on those two plasmids were unchanged, although many of them were expressed. Thus, in D. shibae not the copy number of the plasmid, but specifically the plasmid-localized vir gene clusters are controlled by QS, in contrast to the findings in Agrobacterium tumefaciens (Pappas and Winans, 2003). Both vir gene clusters (Dshi_3637–3650, 3972–3984) were among the only genes whose expression was constantly reduced throughout growth as well as in the stationary phase in the QS null mutant (Figure 5d(1)). In contrast to cell division-related genes and flagellar biosynthesis, both vir gene clusters were overexpressed in D. shibae ΔluxI1pDP1, indicating that the response of this trait is more sensitive towards changes in AHL concentration than others (Figure 5d(2)). This hypothesis is further confirmed by the finding that the addition of AHLs produced by D. shibae caused overexpression of these genes. By contrast, the addition of non-native C18-HSL only led to restoration of the wild-type vir cluster expression level (Figure 5d(3)). This demonstrates that QS molecules from other bacteria can affect the gene expression of specific traits in D. shibae.
Discussion
The C18en-HSL and C18dien-HSL produced by D. shibae represent structures whose signaling role is studied here for the first time. The data strongly suggest that QS in D. shibae controls the switch between two modes of life. In the absence of AHL signals, a fast-growing, morphologically homogenous population is found, which does not invest energy into the synthesis of T4SS and flagella. In the presence of AHL signals, a slower-growing population of remarkable morphological and cell division heterogeneity can be observed, with some of the cells being flagellated. We found that the QS system of D. shibae is not restricted to the autoinducers produced by the organism itself; it specifically responds to a non-self-produced structurally similar AHL by activating the T4SS.
Inactivation of the autoinducer synthase LuxI1 eliminated production of AHLs in D. shibae completely. This is in accordance with the microarray and qPCR data, which showed downregulation of luxR2I2 and luxI3. Thus, a hierarchical relationship appears to be present, with expression of LuxI2 and LuxI3 depending on the signal of LuxI1. Many QS systems show a similarly hierarchical structure (Frederix and Downie, 2011). In P. aeruginosa, two major autoinducers are produced; the long-chain C12-oxo-HSL (Las system) is the dominant one controlling the synthesis of the short-chain C4-HSL (Rhl system) through upregulation of the transcriptional regulator rhlR (Jimenez et al., 2012). One of the most complicated QS systems studied to date is that of Rhizobium leguminosarium biovar viciae, a root nodule-forming soil bacterium, which has four autoinducer synthases. They again display a hierarchical structure, with the dominant master regulator CinR being induced by the 3-OH-C14en-HSL, the product of the autoinducer synthase CinI (Wisniewski-Dye and Downie, 2002). The Roseobacter isolate Ruegeria sp. KLH11 has a QS system that is very similar to that of D. shibae, with two luxI/luxR pairs and one orphan luxI homolog. Like in D. shibae, knocking out its synthase ssaI (sponge-associated symbiont A) gene resulted in complete loss of AHL synthesis (Zan et al., 2012).
The expression of the luxI1 autoinducer synthase gene was constant throughout exponential growth in D. shibae wild type, and the cognate regulator LuxR1 was highly and constitutively expressed throughout growth. Expression of luxI1 under the control of a constitutive promoter restored the pleomorphic phenotype. This finding indicates that the luxR1I1 operon is involved in maintaining morphological heterogeneity but not heterogeneously expressed itself. Bistable expression of regulators acting downstream of luxR1I1, like ctrA or luxR2I2, could be responsible for the observed phenotype. As the hydrophobic long-chain AHLs are unlikely to diffuse freely through the membrane, variability of the transport rate could be another source of cellular heterogeneity.
The D. shibae QS null mutant responded to C18-HSL, a signal that has not been detected in culture supernatants of this bacterium. C18-HSL caused upregulation of the second QS system (luxR2/luxI2), and thus triggered the wild-type-like QS response. However, only a selected set of genes, in particular both plasmid-encoded vir operons, were re-activated (see below). LuxR-type transcriptional regulators accept structurally similar AHLs, a phenomenon that is widely exploited by using reporter strains to detect novel AHLs. Interspecies communication using the archetypical LuxRI system should therefore in fact be widespread. Such crosstalk has rarely been shown, one example being Burkholderia cepacia and Pseudomonas aeruginosa, which colonize the lung of cystic fibrosis patients (Riedel et al., 2001). To fully understand the QS network and signal integration, single-cell techniques will have to be employed, as demonstrated for Vibrio fischeri (Perez et al., 2011) and V. harveyi (Long et al., 2009).
Lack of AHLs affected the expression of 344 genes, representing 8% of the genome. This is comparable to the 6% of QS-controlled genes found in P. aeruginosa (Schuster et al., 2003). Interestingly, as in our study, the largest transcriptional changes occurred at the transition to stationary phase. Thus, QS and starvation sensing converge. In P. aeruginosa the QS regulon and the regulon of the alternative sigma factor RpoS controlling the general stress response showed a strong overlap (Schuster et al., 2004; Schuster and Greenberg, 2007). This overlap may also be present in D. shibae, as expression of two alternative sigma factors was reduced in the QS null mutant.
In D. shibae flagellar synthesis is controlled by AHL signaling. It has been calculated that marine bacteria may spend more than 10% of their total energy budget on movement, and the smaller the cell is, the larger is the amount of energy needed to stabilize it against Brownian movement (Mitchell, 2002; Mitchell and Kogure, 2006). Thus, the metabolic costs for flagellar synthesis are only worth spending in a diffusion-limited patchy microenvironment where motility might provide the chance to reach more optimal conditions or nutrients, a classical condition for QS. In Roseobacters, flagella have been shown to enable chemotaxis towards dimethylsulfono-propionate (DMSP), a storage compound and osmoprotectant synthesized by marine algae (Belas et al., 2009). Flagella mutants have been shown to be impaired in their ability to form biofilms on abiotic surfaces and were not able to attach to diatoms (Sonnenschein et al., 2012) or dinoflagellates (Miller and Belas, 2006).
The vir gene clusters of D. shibae encode a T4SS, which is highly conserved among Roseobacter strains (Wagner-Döbler et al., 2010). Its physiological function has not yet been unraveled. T4SS are the only secretion systems that can translocate not only proteins but also DNA (Christie et al., 2005). In contrast to flagellar biosynthesis and cell-cycle-related genes, the expression of the vir genes could be restored by all tested C18-HSLs, even by the saturated C18-HSL. The natural habitat of D. shibae, the phycosphere of marine algae, harbors microbial communities dominated by Roseobacters. They are known to produce a variety of long-chain AHLs (Wagner-Döbler and Biebl, 2006) and D. shibae may be able to respond to the prevailing AHLs in the community in a specific way.
It is increasingly becoming clear that bacterial cells within isogenic populations can display heterogeneous phenotypes. This so-called phenotypic variation can result from noise in gene expression that is most pronounced when the total number of the involved molecules, for example, transcription factors, is small. It can also be caused by control structures of gene-regulatory networks, in particular positive feedback loops resulting in bistability of gene expression. Variability in the phenotypic outcome of a bacterial population has been suggested to be beneficial especially in highly dynamic environments (Acar et al., 2008). In D. shibae, cell morphology is the most obvious trait showing heterogeneity in the population. We could show by time-lapse microscopy that different cell division types co-exist in this strain: binary fission and budding. The most exciting finding is that this variability is not simply the outcome of noise in the regulation of cell division but controlled by QS. Microarray analysis confirmed the microscopic and flow cytometric investigations. Given the diversity of the cell-cycle-control mechanisms in Alphaproteobacteria and insufficient knowledge of the control system in Rhodobacterales, it is at the moment not possible to speculate on how it functions in D. shibae. It remains to be elucidated if the various types of cell division observed here are connected through a regular cell cycle. In Rhodobacter capsulatus (Mercer et al., 2010) and Silicibacter sp. TM1040 (Belas et al., 2009) the growth rate of the culture was not affected by knockout of ctrA. However, in the latter strain the knockout leads to elongated cells. Polar growth has been described for representatives of Rhizobiales, Caulobacterales and Rhodobacterales and may be ancient in Alphaproteobacteria (Brown et al., 2012); the core genes of cell cycle control are conserved throughout the phylum (Brilli et al., 2010). This is the first time that the mode of cell division has been shown to be controlled by QS in Alphaproteobacteria. Strikingly, the luxI1 mutant loses morphological heterogeneity and shows a faster growth rate than the wild type. This is in contrast to previous work showing growth inhibition by an autoinducer (Gray et al., 1996). Recently, it was observed that QS can induce gas vesicle formation in Serratia sp. (Ramsay et al., 2011). Indeed, morphological differentiation processes are among the first examples that were recognized to involve cell–cell communication. In Bacillus subtilis sporulation and competence are induced by peptide pheromones through complex interconnected genetic circuits (Grossman, 1995). The frequency of sporulating cells is controlled by bistability of the isogenic population (Veening et al., 2008b). Fruiting body formation in Myxobacteria is another extremely complex developmental process, which is controlled by autoinducers in a density-dependent way and requires polar growth. These are in fact the earliest examples of cell–cell communication, going back to the end of the nineteenth century (Kaiser et al., 2010).
Conclusion
In D. shibae, QS induces morphological heterogeneity. Moreover, QS controls flagellation and the expression of the T4SS. It remains to be determined whether these traits are induced in a subpopulation only, and these subpopulations are distinct or overlapping.
Phenotypic variability results in a population with a reduced growth rate, thus representing a burden. We propose that QS-induced heterogeneity ensures that at least a subpopulation of cells maintains a high fitness under constantly changing conditions. This strategy has been described as ‘risk-spreading’ or ‘bet-hedging’ (Veening et al., 2008a). It has evolved to maximize the fitness of the population in an environment with unpredictable fluctuations (Veening et al., 2008a; de Jong et al., 2011). Such fluctuating selective pressures are likely to occur in plankton blooms and during the seasonal succession of microbial communities in the ocean. Size-selective grazing may favor the survival of the larger cells (Gonzalez et al., 1990; Sherr et al., 1992; Hansen, 2011). Moreover, the bacteria have their ears wide open, being able to respond also to long-chain AHLs produced by neighboring cells from different species. Finally, the heterogeneity maintained in the population by the produced QS signals calls for an in-depth investigation on the single-cell level.
Accession codes
References
Acar M, Mettetal JT, van Oudenaarden A . (2008). Stochastic Switching As a Survival Strategy in Fluctuating Environments. Nat Genet 40: 471–475.
Anetzberger C, Pirch T, Jung K . (2009). Heterogeneity in Quorum Sensing-Regulated Bioluminescence of Vibrio Harveyi. Mol Microbiol 73: 267–277.
Anetzberger C, Schell U, Jung K . (2012). Single Cell Analysis of Vibrio Harveyi Uncovers Functional Heterogeneity in Response to Quorum Sensing Signals. BMC Microbiol 12: 209.
Avery SV . (2006). Microbial Cell Individuality and the Underlying Sources of Heterogeneity. Nat Rev Microbiol 4: 577–587.
Bassler BL, Losick R . (2006). Bacterially Speaking. Cell 125: 237–246.
Belas R, Horikawa E, Aizawa S, Suvanasuthi R . (2009). Genetic Determinants of Silicibacter Sp. TM1040 Motility. J Bacteriol 191: 4502–4512.
Biebl H, Allgaier M, Tindall BJ, Koblizek M, Lünsdorf H, Pukall R et al (2005). Dinoroseobacter Shibae Gen. Nov., Sp. Nov., a New Aerobic Phototrophic Bacterium Isolated From Dinoflagellates. Int J Syst Evol Microbiol 55: (Pt 3) 1089–1096.
Biondi EG, Reisinger SJ, Skerker JM, Arif M, Perchuk BS, Ryan KR et al (2006). Regulation of the Bacterial Cell Cycle by an Integrated Genetic Circuit. Nature 444: 899–904.
Brilli M, Fondi M, Fani R, Mengoni A, Ferri L, Bazzicalupo M et al (2010). The Diversity and Evolution of Cell Cycle Regulation in Alpha-Proteobacteria: a Comparative Genomic Analysis. BMC. Syst Biol 4: 52.
Brinkhoff T, Giebel HA, Simon M . (2008). Diversity, Ecology, and Genomics of the Roseobacter Clade: a Short Overview. Arch Microbiol 189: 531–539.
Brown PJ, de Pedro MA, Kysela DT, Van der HC, Kim J, De BX et al (2012). Polar Growth in the Alphaproteobacterial Order Rhizobiales. Proc Natl Acad Sci USA 109: 1697–1701.
Christie PJ, Atmakuri K, Krishnamoorthy V, Jakubowski S, Cascales E . (2005). Biogenesis, Architecture, and Function of Bacterial Type IV Secretion Systems. Annu Rev Microbiol 59: 451–485.
de Jong IG, Haccou P, Kuipers OP . (2011). Bet Hedging or Not? A Guide to Proper Classification of Microbial Survival Strategies. Bioessays 33: 215–223.
Frederix M, Downie AJ . (2011). Quorum Sensing: Regulating the Regulators. Adv Microb Physiol 58: 23–80.
Garmyn D, Gal L, Briandet R, Guilbaud M, Lemaitre JP, Hartmann A et al (2011). Evidence of Autoinduction Heterogeneity Via Expression of the Agr System of Listeria Monocytogenes at the Single-Cell Level. Appl Environ Microbiol 77: 6286–6289.
Giebel HA, Kalhöfer D, Lemke A, Thole S, Gahl-Janssen R, Simon M et al (2011). Distribution of Roseobacter RCA and SAR11 Lineages in the North Sea and Characteristics of an Abundant RCA Isolate. ISME J 5: 8–19.
Gilmore MS, Haas W . (2005). The Selective Advantage of Microbial Fratricide. Proc Natl Acad Sci USA 102: 8401–8402.
Gonzalez JM, Sherr EB, Sherr BF . (1990). Size-Selective Grazing on Bacteria by Natural Assemblages of Estuarine Flagellates and Ciliates. Appl Environ Microbiol 56: 583–589.
Gray KM, Pearson JP, Downie JA, Boboye BE, Greenberg EP . (1996). Cell-to-Cell Signaling in the Symbiotic Nitrogen-Fixing Bacterium Rhizobium Leguminosarum: Autoinduction of a Stationary Phase and Rhizosphere-Expressed Genes. J Bacteriol 178: 372–376.
Grossman AD . (1995). Genetic Networks Controlling the Initiation of Sporulation and the Development of Genetic Competence in Bacillus Subtilis. Annu Rev Genet 29: 477–508.
Hahne F, LeMeur N, Brinkman RR, Ellis B, Haaland P, Sarkar D et al (2009). FlowCore: a Bioconductor Package for High Throughput Flow Cytometry. BMC Bioinformatics 10: 106.
Hansen PJ . (2011). The Role of Photosynthesis and Food Uptake for the Growth of Marine Mixotrophic Dinoflagellates. J Eukaryot Microbiol 58: 203–214.
Jimenez PN, Koch G, Thompson JA, Xavier KB, Cool RH, Quax WJ . (2012). The Multiple Signaling Systems Regulating Virulence in Pseudomonas Aeruginosa. Microbiol Mol Biol Rev 76: 46–65.
Kaern M, Elston TC, Blake WJ, Collins JJ . (2005). Stochasticity in Gene Expression: From Theories to Phenotypes. Nat Rev Genet 6: 451–464.
Kaiser D, Robinson M, Kroos L . (2010). Myxobacteria, Polarity, and Multicellular Morphogenesis. Cold Spring Harb Perspect Biol 2: a000380.
Kaplan HB, Greenberg EP . (1985). Diffusion of Autoinducer Is Involved in Regulation of the Vibrio Fischeri Luminescence System. J Bacteriol 163: 1210–1214.
Klein J, Leupold S, Biegler I, Biedendieck R, Munch R, Jahn D . (2012). TLM-Tracker: Software for Cell Segmentation, Tracking and Lineage Analysis in Time-Lapse Microscopy Movies. Bioinformatics 28: 2276–2277.
Lemme A, Gröbe L, Reck M, Tomasch J, Wagner-Döbler I . (2011). Subpopulation-Specific Transcriptome Analysis of Competence-Stimulating-Peptide-Induced Streptococcus Mutans. J Bacteriol 193: 1863–1877.
Long T, Tu KC, Wang Y, Mehta P, Ong NP, Bassler BL et al (2009). Quantifying the Integration of Quorum-Sensing Signals With Single-Cell Resolution. PLoS Biol 7: 3.
Marie D, Partensky F, Jacquet S, Vaulot D . (1997). Enumeration and Cell Cycle Analysis of Natural Populations of Marine Picoplankton by Flow Cytometry Using the Nucleic Acid Stain SYBR Green I. Appl Environ Microbiol 63: 186–193.
Mercer RG, Callister SJ, Lipton MS, Pasa-Tolic L, Strnad H, Paces V et al (2010). Loss of the Response Regulator CtrA Causes Pleiotropic Effects on Gene Expression but Does Not Affect Growth Phase Regulation in Rhodobacter Capsulatus. J Bacteriol 192: 2701–2710.
Miller TR, Belas R . (2006). Motility Is Involved in Silicibacter Sp. TM1040 Interaction With Dinoflagellates. Environ Microbiol 8: 1648–1659.
Mitchell JG . (2002). The Energetics and Scaling of Search Strategies in Bacteria. Am Nat 160: 727–740.
Mitchell JG, Kogure K . (2006). Bacterial Motility: Links to the Environment and a Driving Force for Microbial Physics. FEMS Microbiol Ecol 55: 3–16.
Müller S . (2007). Modes of Cytometric Bacterial DNA Pattern: a Tool for Pursuing Growth. Cell prolif 40: 621–639.
Nealson KH, Hastings JW . (1979). Bacterial Bioluminescence: Its Control and Ecological Significance. Microbiol Rev 43: 496–518.
Neumann A, Patzelt D, Wagner-Döbler I, Schulz S . (2013). Identification of New N-Acylhomoserine Lactone Signalling Compounds of Dinoroseobacter Shibae DFL-12 by Overexpression of LuxI Genes. Chembiochem submitted.
Pappas KM, Winans SC . (2003). A LuxR-Type Regulator From Agrobacterium Tumefaciens Elevates Ti Plasmid Copy Number by Activating Transcription of Plasmid Replication Genes. Mol Microbiol 48: 1059–1073.
Perez PD, Hagen SJ . (2010). Heterogeneous Response to a Quorum-Sensing Signal in the Luminescence of Individual Vibrio Fischeri. PLoS One 5: e15473.
Perez PD, Weiss JT, Hagen SJ . (2011). Noise and Crosstalk in Two Quorum-Sensing Inputs of Vibrio Fischeri. BMC Syst Biol 5: 153.
Platt TG, Fuqua C . (2010). What’s in a Name? The Semantics of Quorum Sensing. Trends Microbiol 18: 383–387.
Purcell EB, Boutte CC, Crosson S . (2008). Two-Component Signaling Systems and Cell Cycle Control in Caulobacter Crescentus. Adv Exp Med Biol 631: 122–130.
Ramsay JP, Williamson NR, Spring DR, Salmond GP . (2011). A Quorum-Sensing Molecule Acts As a Morphogen Controlling Gas Vesicle Organelle Biogenesis and Adaptive Flotation in an Enterobacterium. Proc Natl Acad Sci USA 108: 14932–14937.
Ratcliff WC, Denison RF . (2010). Individual-Level Bet Hedging in the Bacterium Sinorhizobium Meliloti. Curr Biol 20: 1740–1744.
Riedel K, Hentzer M, Geisenberger O, Huber B, Steidle A, Wu H et al (2001). N-Acylhomoserine-Lactone-Mediated Communication Between Pseudomonas Aeruginosa and Burkholderia Cepacia in Mixed Biofilms. Microbiology 147: (Pt 12) 3249–3262.
Schäfer AL, Greenberg EP, Oliver CM, Oda Y, Huang JJ, Bittan-Banin G et al (2008). A New Class of Homoserine Lactone Quorum-Sensing Signals. Nature 454: 595–599.
Schuster M, Greenberg EP . (2007). Early Activation of Quorum Sensing in Pseudomonas Aeruginosa Reveals the Architecture of a Complex Regulon. BMC Genomics 8: 287.
Schuster M, Hawkins AC, Harwood CS, Greenberg EP . (2004). The Pseudomonas Aeruginosa RpoS Regulon and Its Relationship to Quorum Sensing. Mol Microbiol 51: 973–985.
Schuster M, Lostroh CP, Ogi T, Greenberg EP . (2003). Identification, Timing, and Signal Specificity of Pseudomonas Aeruginosa Quorum-Controlled Genes: a Transcriptome Analysis. J Bacteriol 185: 2066–2079.
Selje N, Simon M, Brinkhoff T . (2004). A Newly Discovered Roseobacter Cluster in Temperate and Polar Oceans. Nature 427: 445–448.
Sherr BF, Sherr EB, McDaniel J . (1992). Effect of Protistan Grazing on the Frequency of Dividing Cells in Bacterioplankton Assemblages. Appl Environ Microbiol 58: 2381–2385.
Smits WK, Kuipers OP, Veening JW . (2006). Phenotypic Variation in Bacteria: the Role of Feedback Regulation. Nat Rev Microbiol 4: 259–271.
Sonnenschein EC, Abebew SD, Grossart HP, Ullrich MS . (2012). Chemotaxis of Marinobacter Adhaerens and Its Impact on Attachment to the Diatom Thalassiosira Weissflogii. Appl Environ Microbiol 78: 6900–6907.
Steinmoen H, Knutsen E, Havarstein LS . (2002). Induction of Natural Competence in Streptococcus Pneumoniae Triggers Lysis and DNA Release From a Subfraction of the Cell Population. Proc Natl Acad Sci USA 99: 7681–7686.
Veening JW, Smits WK, Kuipers OP . (2008a). Bistability, Epigenetics, and Bet-Hedging in Bacteria. Annu Rev Microbiol 62: 193–210.
Veening JW, Stewart EJ, Berngruber TW, Taddei F, Kuipers OP, Hamoen LW . (2008b). Bet-Hedging and Epigenetic Inheritance in Bacterial Cell Development. Proc Natl Acad Sci USA 105: 4393–4398.
Wagner-Döbler I, Ballhausen B, Berger M, Brinkhoff T, Buchholz I, Bunk B et al (2010). The Complete Genome Sequence of the Algal Symbiont Dinoroseobacter Shibae: a Hitchhiker's Guide to Life in the Sea. ISME J 4: 61–77.
Wagner-Döbler I, Biebl H . (2006). Environmental Biology of the Marine Roseobacter Lineage. Annu Rev Microbiol 60: 255–280.
Wagner-Döbler I, Thiel V, Eberl L, Allgaier M, Bodor A, Meyer S et al (2005). Discovery of Complex Mixtures of Novel Long-Chain Quorum Sensing Signals in Free-Living and Host-Associated Marine Alphaproteobacteria. Chembiochem 6: 2195–2206.
Wang YJ, Leadbetter JR . (2005). Rapid Acyl-Homoserine Lactone Quorum Signal Biodegradation in Diverse Soils. Appl Environ Microbiol 71: 1291–1299.
Waters CM, Bassler BL . (2005). Quorum Sensing: Cell-to-Cell Communication in Bacteria. Annu Rev Cell Dev Biol 21: 319–346.
Wisniewski-Dye F, Downie JA . (2002). Quorum-Sensing in Rhizobium. Antonie Van Leeuwenhoek 81: 397–407.
Young JW, Locke JC, Altinok A, Rosenfeld N, Bacarian T, Swain PS et al (2012). Measuring Single-Cell Gene Expression Dynamics in Bacteria Using Fluorescence Time-Lapse Microscopy. Nat Protoc 7: 80–88.
Zan J, Cicirelli EM, Mohamed NM, Sibhatu H, Kroll S, Choi O et al (2012). A Complex LuxR-LuxI Type Quorum Sensing Network in a Roseobacterial Marine Sponge Symbiont Activates Flagellar Motility and Inhibits Biofilm Formation. Mol Microbiol 85: 916–933.
Acknowledgements
This work was funded by the German Research Foundation (DFG) within the Collaborative Research Centre Transregio 51 Roseobacter. We thank the anonymous referees for helping to improve this manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Patzelt, D., Wang, H., Buchholz, I. et al. You are what you talk: quorum sensing induces individual morphologies and cell division modes in Dinoroseobacter shibae. ISME J 7, 2274–2286 (2013). https://doi.org/10.1038/ismej.2013.107
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.107
Keywords
This article is cited by
-
Combined inorganic nitrogen sources influence the release of extracellular compounds that drive mutualistic interactions in microalgal‒bacterial co-cultures
Journal of Applied Phycology (2022)
-
Physiological Advantage of Phenotypic Heterogeneity in a Quorum-Sensing Population
Journal of the Indian Institute of Science (2020)
-
Effect of the luxI/R gene on AHL-signaling molecules and QS regulatory mechanism in Hafnia alvei H4
AMB Express (2019)
-
Genome sequence of Planktotalea frisia type strain (SH6-1T), a representative of the Roseobacter group isolated from the North Sea during a phytoplankton bloom
Standards in Genomic Sciences (2018)
-
Bottleneck in secretion of α-amylase in Bacillus subtilis
Microbial Cell Factories (2017)