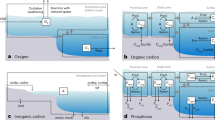
Abstract
Phosphonates (Pn) are diverse organic phosphorus (P) compounds containing C–P bonds and comprise up to 25% of the high-molecular weight dissolved organic P pool in the open ocean. Pn bioavailability was suggested to influence markedly bacterial primary production in low-P areas. Using metagenomic data from the Global Ocean Sampling expedition, we show that the main potential microbial contributor in Pn utilization in oceanic surface water is the globally important marine primary producer Prochlorococcus. Moreover, a number of Prochlorococcus strains contain two distinct putative Pn uptake operons coding for ABC-type Pn transporters. On the basis of microcalorimetric measurements, we find that each of the two different putative Pn-binding protein (PhnD) homologs transcribed from these operons possesses different Pn- as well as inorganic phosphite-binding specificities. Our results suggest that Prochlorococcus adapt to low-P environments by increasing the number of Pn transporters with different specificities towards phosphite and different Pns.
Similar content being viewed by others
Introduction
Dissolved inorganic P concentration of many ocean gyres is very low (Karl, 2000; Wu et al., 2000), near or below the detection limits of analytical methods (Karl and Tien, 1992), and it was shown at times to be the nutrient-limiting productivity (Wu et al., 2000; Sañudo-Wilhelmy et al., 2001; Thingstad et al., 2005; Mather et al., 2008; Lomas et al., 2010). Dissolved organic P concentrations in the open ocean are approximately five times higher than those of dissolved inorganic P, varying between different oceanic regions and depths (for example, 0.22 μM in the upper 100 m in the ALOHA station (Karl and Björkman, 2002)). Phosphonates (Pns) are organic P compounds containing C–P bonds that comprise up to 25% of the high-molecular-weight dissolved organic P pool in the open ocean (Clark et al., 1998). In contrast to the more labile N–P, S–P and O–P linkages, the C–P linkage is resistant to photolysis, thermal decomposition, chemical hydrolysis and phosphatases (Kononova and Nesmeyanova, 2002; Quinn et al., 2007). The chemical stability of Pns may explain their use in membrane lipids and exopolysaccharides, which might be susceptible to degradation owing to their extracellular location (Quinn et al., 2007; White and Metcalf, 2007). Pns bioavailability was suggested to influence markedly bacterial primary production (Karl and Björkman, 2002; Gilbert et al., 2009; Thomas et al., 2010), possibly providing an evolutionary advantage to species able to utilize Pn. The gene encoding the Pn-binding protein (phnD) of the ABC-type Pn transporter is found in genomes of different marine bacteria and is used as one of several proxies for the ability to assimilate Pns in natural microbial assemblages (Dyhrman et al., 2006; Ilikchyan et al., 2009, 2010).
Although different phosphonate (Pn) utilization pathways exist (McGrath et al., 1995; Kononova and Nesmeyanova, 2002; Quinn et al., 2007; Martinez et al., 2010; Gomez-Garcia et al., 2011), the uptake mediated by the ABC-type transporter, Pn-binding-protein (PhnD) is shared across different microbial phyla. The phnD gene is found in numerous marine bacterial genomes, including genomes of the globally abundant marine bacteria Trichodesmium (Dyhrman et al., 2006), Synechococcus, Prochlorococcus and Pelagibacter (the SAR11 group). Enrichment in SAR11 Pn utilization genes or peptides was reported in the P-depleted eastern Mediterranean Sea (via metagenomics (Feingersch et al., 2010)) and the Sargasso Sea (via metaproteomics (Sowell et al., 2009) and metagenomics (Coleman and Chisholm, 2010)), whereas cyanobacterial phnD genes were shown to be expressed in P-deficient media and in the environment (Dyhrman et al., 2006; Ilikchyan et al., 2009, 2010; Beversdorf et al., 2010).
In this work, we studied the distribution of organic P-utilizing microbes in different marine systems using the phnD gene, a gene-proxy indicator for organic Pn utilization. Prochlorococcus genes were found to be the dominant phnD genes in the environments and were therefore expressed in Escherichia coli and their binding to different Pn molecules was monitored in order to find the corresponding Pn substrates.
Materials and methods
Abundance and classification of microbial P and Pn utilization genes in different Global Ocean Sampling (GOS) stations
The ‘GOS: all ORF peptides (P)’ database, downloaded from CAMERA (https://portal.camera.calit2.net/gridsphere/gridsphere), was first BLASTp queried with six PhnD and seven PstS sequences from representative taxonomic phyla (-p blastp -e 1e-5 –F F). The result was a list of sequences that were blasted further against the NCBI non-redundant database in order to determine if they are PhnD and PstS (-p blastp -e 1e-5 –F F). Hits with E-value <1 × 10−20 were categorized and classified according to the top high-scoring sequence pair of a query as determined by the NCBI taxonomic identifier. The PhnD (315 aa average) and PstS (352 aa average) hits were normalized for each GOS station as the stations vary in sequencing depth and sampling methods. In order to compensate for this variation, the average copy number of the genes was calculated as previously described (Beszteri et al., 2010) and can be see in Equation (1):

where  is the average genome size, Im the gene length and
is the average genome size, Im the gene length and  the average copy number of the gene concerned.
the average copy number of the gene concerned.
The average genome size in every station was calculated with the method developed by Raes et al. (2007) (Equation (2)) using 35 COGs of single copy marker genes obtained from the STRING protein DB version. 9.0. (Szklarczyk et al., 2011).
Equation (2):

where Ls is the average read length of sample rm,s the count of reads annotated as marker m from sample s and Rs the total number of base pairs sequenced from sample s.
The pstS or phnD gene sequences from GOS that were similar to Psychroflexus torquis were classified as ‘other’, because the genome sequence of the P. torquis contains small fragments of unknown environmental DNA, as surmised from rDNA in unassembled reads (Howard et al., 2006).
PhnD protein expression
The two phnD genes from Prochlorococcus MIT9301 and the phnD gene from E. coli were synthesized and cloned in the expression vector pJexpress406 (T5 promoterA, Kanr, Ampr, Clmr, pUC originC) by DNA2.0 (Menlo Park, CA, USA) as His-tagged and signal sequence-less (Rizk et al., 2006) proteins, and the codon usage was optimized to fit E. coli usage. One Shot TOP10 E. coli cells (Invitrogen, Carlsbad, CA, USA) transformed with the expression vector carrying the appropriate PhnD inserts were plated on LB agar plates with ampicillin (100 μg ml−1), and grown overnight at 37 °C. Single colonies were suspended in 15 ml of LB with ampicillin (100 μg ml−1), grown overnight at 37 °C and used to inoculate TB medium (10 ml of overnight culture in 1 l) supplemented with ampicillin (50 μg ml−1). Cultures were grown at 210 r.p.m. and 37 °C until OD600=0.45–0.8. IPTG (0.1 mM) was then added and the cells were grown overnight at 210 r.p.m. and 18 °C for overnight expression.
Cells from 1 l cultures were harvested and resuspended in 45 ml binding buffer (30 mM imidazole, 0.5 M NaCl, 20 mM phosphate buffer, pH 7.4). The cells were disrupted by two passages through a French-Press at room temperature. Cell extract was centrifuged (30 500 g for 30 min at 4 °C), and supernatant was transferred to a new centrifuge tube. The soluble fraction containing the recombinant proteins was purified by Fast Protein Liquid Chromatography using the AKTA Explorer (Pharmacia, Uppsala, Sweden) system equipped with 5 ml His-trap column (HisTrap HP Columns, GE Healthcare, Uppsala, Sweden) and eluted with elution buffer (0.5 M imidazole, 0.5 M NaCl, 20 mM phosphate buffer, pH 7.4) according to the manufacturer's instructions.
Titrations of microcalorimetry measurements
Titrations of microcalorimetry measurements were performed with a VP-isothermal microcalorimetry (ITC) calorimeter (Microcal, Northampton, MA, USA). Purified PhnD protein solutions for ITC were dialyzed twice for 12 h, 5 ml against 3 lof buffer A (50 mM Tris HCl, pH 7.4, 500 mM NaCl and 0.02% NaN3). Pn solutions were prepared by dilution with buffer A. Aliquots (10 μl) of the Pn solutions at about 10 times the PhnD protein concentration were added by means of a 280-μl rotating stirrer-syringe to the reaction cell containing 1.41 ml of 0.1 μM of the different PhnD protein solutions. The heat of dilution was determined to be negligible in separate titrations of the Pn solutions into the buffer solution. Calorimetric data analysis was carried out using Origin 7.0 software (MicroCal).
Isothermal titration calorimetery between the PhnD proteins and the different Pn ligands can be described by the following direct relation shown in Equation (3):

where (qi) is the amount of heat released or observed proportional to the bound ligand, v is the volume of the reaction cell and ΔLi is the increase in the concentration of bound ligand after the ith injection.
The quantity ΔLi is the difference between the concentration of bound ligand in the ith and (i–1)th injections, and its functional form depends on the specific binding model. For the simplest case, in which the protein has one binding site, it becomes Equation (4):

where Ka is the binding constant and [L] is the concentration of free ligand (Leavitt and Freire, 2001).
Results and discussion
In order to account for the distribution and prevalence of organic P-utilizing microbes in different marine systems, we calculated the relative abundance of gene-proxy indicators for organic Pn utilization (the phnD gene) in the GOS project (Rusch et al., 2007), and compared it with inorganic P utilization (based on the pstS gene encoding for the ABC-type transporter phosphate-binding protein (Martiny et al., 2009)). GOS is currently the largest collection of metagenomic datasets that includes samples from the Atlantic, Pacific and Indian Oceans. Although different microbial groups show the potential for inorganic P utilization (Figure 1a), the BLAST results imply that the main factors in organic P utilization in surface waters are Prochlorococcus (Figure 1b in the Pacific and Indian Ocean stations and in one station in the Sargasso Sea) and to a lesser extent SAR11 bacteria (Figure 1b in several Atlantic Ocean stations).
Relative abundance of microbial inorganic P and Pn utilization genes at different GOS stations. (a) The relative quantity of pstS genes of various microbial groups (rows) at each station (columns) is represented by the area of the corresponding spot (if any). (b) The relative quantity of phnD genes of various microbial groups. The figure was constructed using the ggplot2 package (Wickham, 2009) in R. See station locations overlaid on a P concentrations map in Figure S3.
Prochlorococcus cell abundance alone could not explain the high abundance of Prochlorococcus phnD genes, as SAR11s are the dominant bacteria in these GOS stations (SAR11 is eight times more abundant over the entire GOS sets compared with Prochlorococcus based on 16S gene recruitments (see Table 8 in Rusch et al., 2007)). One possible explanation is that our phylogenetic group assignments are based on homology searches and are therefore prone to errors such as wrong affiliations as a result of possible horizontal gene transfers. Another possible explanation is the existence of SAR11 types that completely lack any copy of the phnD gene, or alternatively the existence of Prochlorococcus strains having more than one copy of the phnD gene. Indeed, a search through different SAR11 and Prochlorococcus genomes revealed the absence of the phnD gene from the genomes of two SAR11 isolates (‘Candidatus Pelagibacter ubique’ HTCC1062 (Rappé et al., 2002; Giovannoni et al., 2005) and HTCC1002 (Rappé et al., 2002); see Table 1), and the existence of two different phn operons (coding for the different components of ABC-type Pn transporters) in two Prochlorococcus genomes (the high-light (HL) adapted MIT9301 and the low-light (LL) adapted MIT9303 ecotypes; see Table 1). As already observed by Martiny et al. (2009), the same Prochlorococcus ecotypes also show an increase in the number of pstS genes in their genomes. Interestingly, this trend is also observed among marine Synechococcus strains (Moore et al., 2005). Moreover, not only does the gene organization of the two phn operons in Prochlorococcus differ (phn D CE and phnC D E), the proteins themselves are also distinct (possessing 27% aa identity) and cluster separately based on protein phylogeny (Figure 2). The Prochlorococcus phnCDE operon is found in the vicinity of another operon that contains two novel Pn utilization genes (phnY and phnZ) that were identified recently via functional screening in E. coli (Martinez et al., 2010). Both the phnCDE and phnYZ operons are located in a genomic island in Prochlorococcus MIT9301 (Kettler et al., 2007). It is important to note that the phnD primers previously used by Ilikchyan et al. (2009, 2010) to follow Prochlorococcus phnD expression will recognize only the phnD gene from the phn D CE and not the copy in the phnCDE operon.
Neighbor-joining tree of Prochlorococcus PhnD proteins. Evolutionary distances for PhnD proteins were determined from the alignment of 320 aa positions using a neighbor-joining analysis. Neighbor-joining and maximum parsimony analyses were conducted using PAUP. Bootstrap resampling (1000) of neighbor-joining and maximum parsimony trees were performed in all analyses to provide confidence estimates for the inferred topologies. Bootstrap values (neighbor-joining/maximum parsimony) greater than 50% are indicated above the branches. Prochlorococcus PhnD proteins are shown in green. HL – high-light adapted strains, LL – low-light adapted strains. Different phn operon organizations in Prochlorococcus MIT9301 and MIT9303 and in different reference genomes are shown on the right; gray ORF represents genes not related to the phn operon.
In order to determine Pn-binding specificities for the two different Prochlorococcus PhnD proteins (each one possessing less than 30% aa identity to the E. coli PhnD homolog), the phnD genes from the phn D CE (locus P9301_07261; phnD1) and phnC D E (locus P9301_12511; phnD2) operons of the HL-adapted Prochlorococcus MIT9301 (as MIT9301 often shares the highest sequence similarity with GOS sequences (Kettler et al., 2007)) were expressed as His-tagged proteins in E. coli and tested against a battery of different simple Pns, phosphite and phosphate using ITC measurements. ITC measures the heat capacity change upon the interaction between a macromolecule and a ligand. Titration of the ligand versus the macromolecule results in an isotherm-binding curve that can be fitted to a simple binding equation to calculate the equilibrium constant. As expected, the purified control His-tagged E. coli PhnD protein bound different Pns with the previously reported affinities (Rizk et al., 2006) (2-aminoethylphosphonate (2-AEPn) ≫ ethylphosphonate (EPn) > methylphosphonate (MPn) ⩾ phosphonoacetate (PnAc) ⩾ aminomethylphosphonate (AMPn); see Table 2 for Kd values and Figure S1 for raw data. It is important to note that using ITC measurements, we failed to detect any binding of the E. coli PhnD protein either to phosphite or to phosphate. These were previously suggested to be P substrates in E. coli (Metcalf and Wanner, 1991; Rizk et al., 2006)). Interestingly, both Prochlorococcus PhnD proteins showed a completely different specificity range: one (PhnD2) binds strongly only to MPn, EPn and inorganic phosphite, whereas the other (PhnD1) binds strongly to inorganic phosphite and with very weak affinities to MPn and phosphate (see Table S1 for dissociation constants). Although this suggests that simple Pns and phosphites could be utilized by Prochlorococcus possessing PhnDs similar to PhnD2 (observed in only few cultured ecotypes), it also implies that in the environment, Prochlorococcus possessing PhnDs similar to PhnD1 (observed in all cultured ecotypes isolated so far) use phosphites. Indeed, Martinez et al. (in press) were able to show that Prochlorococcus MIT9301 can grow on phosphite as a sole P source. However, under the growth condition tested, MIT9301 failed to grow on different Pns (2-AEPn, EPn or phosphonoalanine) as a P source. Based on these findings, Martinez et al. (in press) suggested that the second Pn operon phnCDE (re-annotated as ptxABC by the authors) is a phosphite utilization operon along with the neighboring phnY and phnZ genes and an adjacent newly identified phosphite dehydrogenase (ptxD). However, other Prochlorococcus strains tested, which lack the ptxABCD operon but contain the phnDCE operon, failed to grow on phosphite or Pns (2-AEPn, EPn or phosphonoalanine) under the growth conditions tested (Martinez et al., in press). Although our biochemical binding assay predicted that the transporter encoded by the phnCDE (ptxABC) operon would transport simple Pns (MPn and EPn) as well as phosphite, it also predicted that the transporter encoded by the phnDCE operon is a phosphite transporter. This could not be resolved by Martinez et al. (in press) under the growth conditions used in their experiments. Further experiments using different growth conditions are therefore needed in order to confirm whether the transporter encoded by the phnDCE operon transport phosphite and the transporter encoded by the phnCDE (ptxABC) operon could also transport simple Pns (MPn and EPn). No data currently exist on phosphite abundance in the marine environment (David Karl, personal communication). The current widely used methodologies (Lomas et al., 2010) of total dissolved P measurement and subsequent calculation of organic P by the difference between phosphate and total dissolved P would hide any phosphite as organic P.
Examination of publicly available marine metatranscriptomic datasets (Hewson et al., 2009; Stewart et al., 2010) revealed that both Prochlorococcus phnD1 and phnD2 genes are expressed in several Atlantic Ocean surface water sites but not in Pacific Ocean sites (Table 3). This is in agreement with recent observations of the enrichment of P utilization genes (Prochlorococcus and SAR11) in the BATS station (Bermuda Atlantic Time Series, North Atlantic) compared with the HOT station (Hawaii Ocean Time Series, North Pacific) (Coleman and Chisholm, 2010, 2011).
Besides the two different PhnD proteins from Prochlorococcus MIT9301, a phylogenetic tree constructed with all Prochlorococcus-like PhnDs from GOS revealed the existence of other potential phosphite or PhnD families (some of which having less than 70% identity to PhnDs from cultured Prochlorococcus (marked with black arrows in Figure S2)), with the majority (more than 90%) belonging to the PhnD1 enlarged family.
The marine cyanobacteria of the genera Prochlorococcus are globally important marine primary producers (Chisholm et al., 1988; Partensky et al., 1999) and have undergone extensive genome streamlining (Partensky and Garczarek, 2010). It seems that Prochlorococcus ecotypes adapted to low-P environments use at least three different strategies to deal with low-P availability: (i) they substitute their phospholipids with non-P membrane lipids such as sulphoquinovosyldiacylglycerol (Van Mooy et al., 2009) and therefore reduce the cellular demand for P by half; (ii) they increase the number of organic (Table 1) and inorganic P transporters (Martiny et al., 2009); and (iii) they have an arsenal of transporters with different affinities to phosphite and different Pns (Table 2) to be ready on demand. Future in vivo growth experiments using different Prochlorococcus strains are needed to test which Prochlorococcus cells can supplement growth using phosphite or simple Pns (like MPn and EPn) as P sources.
References
Beszteri B, Temperton B, Frickenhaus S, Giovannoni SJ . (2010). Average genome size: a potential source of bias in comparative metagenomics. ISME J 4: 1075–1077.
Beversdorf LJ, White AE, Björkman KM, Letelier RM, Karl DM . (2010). Phosphonate metabolism of Trichodesmium IMS101 and the production of greenhouse gases. Limnol Oceanogr 55: 1768.
Chisholm SW, Olson RJ, Zettler ER, Goericke R, Waterbury J, Welschmeyer N . (1988). A novel free-living prochlorophyte abundant in the oceanic euphotic zone. Nature 334: 340–343.
Clark LL, Ingall ED, Benner R . (1998). Marine phosphorus is selectively remineralized. Nature 393: 426–429.
Coleman ML, Chisholm SW . (2010). Ecosystem-specific selection pressures revealed through comparative population genomics. Proc Natl Acad Sci USA 107: 18634–18639.
Coleman ML, Chisholm SW . (2011). Reply to Luo and Konstantinidis: Phosphorus-related genes are enriched in Prochlorococcus populations from the North Atlantic. Proc Natl Acad Sci USA 108: E64–E66.
Dyhrman ST, Chappell PD, Haley ST, Moffett JW, Orchard ED, Waterbury JB et al. (2006). Phosphonate utilization by the globally important marine diazotroph Trichodesmium. Nature 439: 68–71.
Feingersch R, Suzuki MT, Shmoish M, Sharon I, Sabehi G, Partensky F et al. (2010). Microbial community genomics in eastern Mediterranean Sea surface waters. ISME J 4: 78–87.
Gilbert JA, Thomas S, Cooley NA, Kulakova A, Field D, Booth T et al. (2009). Potential for phosphonoacetate utilization by marine bacteria in temperate coastal waters. Environ Microbiol 11: 111–125.
Giovannoni SJ, Tripp HJ, Givan S, Podar M, Vergin KL, Baptista D et al. (2005). Genome streamlining in a cosmopolitan oceanic bacterium. Science 309: 1242–1245.
Gomez-Garcia MR, Davison M, Blain-Hartnung M, Grossman AR, Bhaya D . (2011). Alternative pathways for phosphonate metabolism in thermophilic cyanobacteria from microbial mats. ISME J 5: 141–149.
Hewson I, Poretsky RS, Beinart RA, White AE, Shi T, Bench SR et al. (2009). In situ transcriptomic analysis of the globally important keystone N2-fixing taxon Crocosphaera watsonii. ISME J 3: 618–631.
Howard EC, Henriksen JR, Buchan A, Reisch CR, Burgmann H, Welsh R et al. (2006). Bacterial taxa that limit sulfur flux from the ocean. Science 314: 649–652.
Ilikchyan IN, Mckay RM, Kutovaya OA, Condon R, Bullerjahn GS . (2010). Seasonal expression of the picocyanobacterial phosphonate transporter gene phnD in the Sargasso Sea. Front Microbiol 1: 135.
Ilikchyan IN, McKay RM, Zehr JP, Dyhrman ST, Bullerjahn GS . (2009). Detection and expression of the phosphonate transporter gene phnD in marine and freshwater picocyanobacteria. Environ Microbiol 11: 1314–1324.
Karl DM . (2000). Phosphorus, the staff of life. Nature 406: 31–32.
Karl DM, Björkman KM . (2002). Dynamics of DOP. In Hansell DA, Carlson CA (eds) Biogeochemistry of Marine Dissolved Organic Matter. Academic: London, pp 249–366.
Karl DM, Tien G . (1992). MAGIC: a sensitive and precise method for measuring dissolved phosphorus in aquatic environments. Limnol Oceanogr 37: 105–116.
Kettler G, Martiny AC, Huang K, Zucker J, Coleman ML, Rodrigue S et al. (2007). Patterns and implications of gene gain and loss in the evolution of Prochlorococcus. PLoS Genet 3: e231.
Kononova SV, Nesmeyanova MA . (2002). Phosphonates and their degradation by microorganisms. Biochemistry (Mosc) 67: 184–195.
Leavitt S, Freire E . (2001). Direct measurement of protein binding energetics by isothermal titration calorimetry. Curr Opin Struct Biol 11: 560–566.
Lomas MW, Burke AL, Lomas DA, Bell DW, Shen C, Dyhrman ST et al. (2010). Sargasso Sea phosphorus biogeochemistry: an important role for dissolved organic phosphorus (DOP). Biogeosciences 7: 695–710.
Martinez A, Osburne MS, Sharma AK, DeLong EF, Chisholm SW . (in press). Phosphite utilization by the marine picocyanobacterium Prochlorococcus MIT9301. Environ Microbiol.
Martinez A, Tyson GW, Delong EF . (2010). Widespread known and novel phosphonate utilization pathways in marine bacteria revealed by functional screening and metagenomic analyses. Environ Microbiol 12: 222–238.
Martiny AC, Huang Y, Li W . (2009). Occurrence of phosphate acquisition genes in Prochlorococcus cells from different ocean regions. Environ Microbiol 11: 1340–1347.
Mather RL, Reynolds SE, Wolff GA, Williams RG, Torres-Valdes S, Woodward EMS et al. (2008). Phosphorus cycling in the North and South Atlantic Ocean subtropical gyres. Nature Geosci 1: 439–443.
McGrath JW, Wisdom GB, McMullan G, Larkin MJ, Quinn JP . (1995). The purification and properties of phosphonoacetate hydrolase, a novel carbon-phosphorus bond-cleavage enzyme from Pseudomonas fluorescens 23F. Eur J Biochem 234: 225–230.
Metcalf WW, Wanner BL . (1991). Involvement of the Escherichia coli phn (psiD) gene cluster in assimilation of phosphorus in the form of phosphonates, phosphite, Pi esters, and Pi. J Bacteriol 173: 587–600.
Moore LR, Ostrowski M, Scanlan DJ, Feren K, Sweetsir T . (2005). Ecotypic variation in phosphorus-acquisition mechanisms within marine picocyanobacteria. Aquat Microb Ecol 39: 257–269.
Partensky F, Garczarek L . (2010). Prochlorococcus: advantages and limits of minimalism. Ann Rev Mar Sci 2: 305–331.
Partensky F, Hess WR, Vaulot D . (1999). Prochlorococcus, a marine photosynthetic prokaryote of global significance. Microbiol Mol Biol Rev 63: 106–127.
Quinn JP, Kulakova AN, Cooley NA, McGrath JW . (2007). New ways to break an old bond: the bacterial carbon-phosphorus hydrolases and their role in biogeochemical phosphorus cycling. Environ Microbiol 9: 2392–2400.
Raes J, Korbel JO, Lercher MJ, von Mering C, Bork P . (2007). Prediction of effective genome size in metagenomic samples. Genome Biol 8: R10.
Rappé MS, Connon SA, Vergin KL, Giovannoni SJ . (2002). Cultivation of the ubiquitous SAR11 marine bacterioplankton clade. Nature 418: 630–633.
Rizk SS, Cuneo MJ, Hellinga HW . (2006). Identification of cognate ligands for the Escherichia coli phnD protein product and engineering of a reagentless fluorescent biosensor for phosphonates. Protein Sci 15: 1745–1751.
Rusch DB, Halpern AL, Heidelberg KB, Sutton G, Williamson SJ, Yooseph S et al. (2007). The Sorcerer II Global Ocean Sampling expedition: I, the northwest Atlantic through the eastern tropical Pacific. PLoS Biol 5: e77.
Sañudo-Wilhelmy SA, Kustka AB, Gobler CJ, Hutchins DA, Yang M, Lwiza K et al. (2001). Phosphorus limitation of nitrogen fixation by Trichodesmium in the central Atlantic Ocean. Nature 411: 66–69.
Sowell SM, Wilhelm LJ, Norbeck AD, Lipton MS, Nicora CD, Barofsky DF et al. (2009). Transport functions dominate the SAR11 metaproteome at low-nutrient extremes in the Sargasso Sea. ISME J 3: 93–105.
Stewart FJ, Ottesen EA, DeLong EF . (2010). Development and quantitative analyses of a universal rRNA-subtraction protocol for microbial metatranscriptomics. ISME J 4: 896–907.
Szklarczyk D, Franceschini A, Kuhn M, Simonovic M, Roth A, Minguez P et al. (2011). The STRING database in 2011: functional interaction networks of proteins, globally integrated and scored. Nucleic Acids Res 39: D561–D568.
Thingstad TF, Krom MD, Mantoura RF, Flaten GA, Groom S, Herut B et al. (2005). Nature of phosphorus limitation in the ultraoligotrophic eastern Mediterranean. Science 309: 1068–1071.
Thomas S, Burdett H, Temperton B, Wick R, Snelling D, McGrath JW et al. (2010). Evidence for phosphonate usage in the coral holobiont. ISME J 4: 459–461.
Van Mooy BA, Fredricks HF, Pedler BE, Dyhrman ST, Karl DM, Koblizek M et al. (2009). Phytoplankton in the ocean use non-phosphorus lipids in response to phosphorus scarcity. Nature 458: 69–72.
White AK, Metcalf WW . (2007). Microbial metabolism of reduced phosphorus compounds. Annu Rev Microbiol 61: 379–400.
Wickham H . (2009). ggplot: Elegant Graphics for Data Analysis. Springer: New York, 2009.
Wu J, Sunda WG, Boyle EA, Karl DM . (2000). Phosphate depletion in the Western North Atlantic Ocean. Science 289: 759–762.
Acknowledgements
We would like to thank Orly Tabachnikov for her technical help and Arnon Henn, Nurit Kress, David Karl and Debbie Lindell for their encouragement and discussions. This work was supported by grant 580/10 from the Israel Science Foundation (OB).
Author Contributions
RF and OB designed the project and the experiments; RF, AP, FG and OB performed the bioinformatics experiments; TM conducted the chemistry experiments; RF, OA and YS conducted the biochemistry and molecular biology experiments; OB wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Feingersch, R., Philosof, A., Mejuch, T. et al. Potential for phosphite and phosphonate utilization by Prochlorococcus. ISME J 6, 827–834 (2012). https://doi.org/10.1038/ismej.2011.149
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.149
Keywords
This article is cited by
-
Marine picocyanobacterial PhnD1 shows specificity for various phosphorus sources but likely represents a constitutive inorganic phosphate transporter
The ISME Journal (2023)
-
Differential global distribution of marine picocyanobacteria gene clusters reveals distinct niche-related adaptive strategies
The ISME Journal (2023)
-
Phosphorus as an integral component of global marine biogeochemistry
Nature Geoscience (2021)
-
Transporter characterisation reveals aminoethylphosphonate mineralisation as a key step in the marine phosphorus redox cycle
Nature Communications (2021)
-
Phosphite binding by the HtxB periplasmic binding protein depends on the protonation state of the ligand
Scientific Reports (2019)