Abstract
We developed a broad-ranging method for identifying key hydrogen-producing and consuming microorganisms through analysis of hydrogenase gene content and expression in complex anaerobic microbial communities. The method is based on a tiling hydrogenase gene oligonucleotide DNA microarray (Hydrogenase Chip), which implements a high number of probes per gene by tiling probe sequences across genes of interest at 1.67 × –2 × coverage. This design favors the avoidance of false positive gene identification in samples of DNA or RNA extracted from complex microbial communities. We applied this technique to interrogate interspecies hydrogen transfer in complex communities in (i) lab-scale reductive dehalogenating microcosms enabling us to delineate key H2-consuming microorganisms, and (ii) hydrogen-generating microbial mats where we found evidence for significant H2 production by cyanobacteria. Independent quantitative PCR analysis on selected hydrogenase genes showed that this Hydrogenase Chip technique is semiquantitative. We also determined that as microbial community complexity increases, specificity must be traded for sensitivity in analyzing data from tiling DNA microarrays.
Similar content being viewed by others
Introduction
Molecular hydrogen (H2) is a key metabolic intermediate in many anaerobic microbial communities, being produced by microbes during both the fermentation of organic compounds and consumed by microbes coupling the oxidation of H2 to the reduction of oxidized compounds (Wolin and Miller, 1982; Hoehler, 2005). In many microbial ecosystems, including phototrophic microbial mats and reductively dehalogenating microbial consortia present in organohalide-contaminated groundwater aquifers, hydrogen metabolism has a critical role for the systems-level performance and stability of the respective ecosystem; yet hydrogen flux between the community members has been difficult to determine (Yang and McCarty, 1998; Hoehler et al., 2001, 2002; Smidt and de Vos, 2004). Many interspecies hydrogen transfer interactions are syntrophic, and therefore only present in the complex microbial communities rather than in pure cultures (Bryant et al., 1967; Stams and Plugge, 2009). To our knowledge, there are currently no cultivation-independent molecular methods capable of comprehensively determining which microbes are producing or consuming hydrogen in the complex microbial communities through characterization of the hydrogenase gene presence and expression.
Microbial hydrogen production and consumption is catalyzed by hydrogenases. Hydrogenases can be placed into three broad categories based on the metal cofactors found at their active sites (Vignais and Billoud, 2007; Vignais, 2008; Thauer et al., 2010). (NiFe) hydrogenases (including (NiFeSe) hydrogenases) are implicated in both H2 production and consumption. (FeFe) hydrogenases typically produce H2, often have a higher turnover rate and are more often active in environments with higher H2 partial pressure than (NiFe) hydrogenases. (Fe) hydrogenases are so far found only in methanogenic archaea without cytochromes (Thauer et al., 2010). Hydrogenase genes are too diverse to be characterized through PCR-based methods similar to those used to characterize the microbial communities involved in processes like methanogenesis (Ohkuma et al., 1995) or sulfate reduction (Karkhoff-Schweizer et al., 1995). Non-targeted metagenomic and metatranscriptomic sequencing may identify and quantify the hydrogenase genes, but not in a cost-effective manner. Previous studies characterizing the hydrogenase genes present in microbial communities have only targeted specific subgroups of hydrogenases (Roeselers et al., 2008; Xing et al., 2008; Boyd et al., 2009). In this study, we focused on the most widespread hydrogenases, the (NiFe) (including (NiFeSe)) and (FeFe) hydrogenases, and demonstrated the most broadly targeted approach to date to characterize diverse hydrogenase genes present and expressed in a microbial community.
In order to characterize diverse hydrogenase genes in a comprehensive manner, we opted to use high-density oligonucleotide DNA microarrays. Although various microarray-based techniques have been applied to the study of microbial ecosystems in the past (Bodrossy et al., 2003; Taroncher-Oldenburg et al., 2003; Palmer et al., 2006; DeSantis et al., 2007; He et al., 2007, 2010; Miller et al., 2008; Pozhitkov et al., 2008; Rich et al., 2008; Dugat-Bony et al., 2011), this is the first attempt to broadly target a single class of genes with a high degree of sensitivity and specificity without first enriching the gene of interest using PCR. To achieve this, we developed a tiling functional gene DNA microarray technique using an in-situ ink-jet synthesized oligonucleotide DNA microarray (Hughes et al., 2001; Wolber et al., 2006), where probes are designed based on an even tiling pattern across each hydrogenase gene resulting in a complete 1.67 × –2 × coverage of each gene. The resulting high number of probes targeting each gene enables us to accurately characterize hydrogenase genes present or expressed in a given sample. The risk of false positive gene identification by cross hybridization to a single or small number of probes is minimized by the requirement that 90% or more of the ∼30–50 probes targeting a given gene are ‘bright’. We examined the usefulness and limitations of this tiling functional gene DNA microarray on samples derived from reductively dehalogenating laboratory microcosms, and complex phototrophic hydrogen-producing microbial mats.
Materials and methods
Microarray design
To ensure the most complete set of hydrogenase and the other genes employed for each experiment described in this study, the microarray design was revised four times in the 3-year span of the reported experiments to reflect changes in up-to-date genomic and metagenomic databases (Table 1). Furthermore, this enabled non-hydrogenase genes relevant to the different study systems to be added to the microarray design when space on the array allowed.
Sequences for the Test Microarray were obtained using BLAST on the NCBI non-redundant database (Altschul et al., 1990), with hydrogenase gene sequences listed for (NiFe)-hydrogenase large subunits and (FeFe)-hydrogenase sequences from Vignais et al., (2001) as query sequences. Resulting hydrogenase genes and genes similar to hydrogenases were clustered to 97% using CD-HIT (Li and Godzik, 2006), and the longest sequence for each cluster was selected as the representative sequence for use on the array. Overlapping 60-mer probes for each gene were designed to 2 × coverage. To investigate the effect of mismatched probes, several mismatches were introduced for a probe encoding the hydB gene from Shewanella oneidensis MR-1 (IMG 637345681). A total of 19 probes with a series of single mismatches (98% sequence identity with the true sequence) in different positions at either end of the probe sequence were included, and an 11-nucleotide (82% sequence identity with the true sequence) mismatch section from the center of the probe was also included, with 9 different random mismatch sequences on 9 different probes (see Supplementary Table S1 for mismatch-probe sequences).
All protein and nucleic acid sequences for the Hydrogenase Chip versions 1, 2, and 3 were retrieved from the Integrated Microbial Genomes and Microbiomes database (IMG/M) versions 2.5, 2.8 and 2.9, respectively (Markowitz et al., 2008). The protein sequences were screened for the hydrogenases based on PROSITE sequence signatures for all the (NiFe)- and (FeFe)-hydrogenase groups previously determined (Vignais and Billoud, 2007) using ScanProsite (Gattiker et al., 2002). The non-hydrogenase gene sequences were removed from the resulting gene set based on annotation, and genes were clustered to 97% nucleic-acid sequence identity using CD-HIT (Li and Godzik, 2006). Extra genes were added to each Hydrogenase Chip version based on the available space and experimental questions; these are described in the Supplementary Methods. Even-spaced tiling 60-mer probes for each gene were designed to 2 × (versions 1 and 2) and 1.67 × (version 3) coverage.
Probes for all arrays were randomly positioned on an Agilent 4 × 44 K oligonucleotide microarray format by eArray software (Agilent Technologies, Santa Clara, CA, USA), then synthesized by Agilent with in-situ ink-jet technology.
Pure-culture DNA extraction and sensitivity analysis
DNA for the Test Microarray hybridization was extracted using the DNeasy Blood and Tissue Kit (Qiagen, Hamburg, Germany) according to the manufacturer's instructions for bacterial genomic DNA extraction and purification, then quantified using the Qubit Fluorometer and broad-range double-stranded DNA quantification kit (Invitrogen, San Diego, CA, USA). A total of 1.5 μg of genomic DNA from each of Escherichia coli, Bacillus subtilis, Shewanella oneidensis MR-1, Pseudomonas aeruginosa and Shewanella sediminis HAW-EB3 was mixed for subsequent labeling and hybridization. The same DNA extraction protocol was used to obtain DNA for the sensitivity analysis hybridization of DNA from Escherichia coli, Shewanella oneidensis MR-1, Azotobacter vinelandii and Methanococcus maripaludis. Different quantities of DNA from each organism (see Supplementary Table S11) were mixed together then amplified by multiple displacement amplification using the REPLI-g Mini Kit (Qiagen) according to the manufacturer's instructions.
Reductive dechlorinating soil column operation and DNA extraction
The reductive dechlorinating soil column and the resultant DNA samples used in this study were the same as those used by Azizian et al. (2010). This column was maintained with lactate, propionate or formate as an electron donor. The inoculum culture for the soil column and liquid chemostat has been previously described (Yu et al., 2005). DNA was amplified by the multiple displacement amplification (MDA) for microarray applications using the REPLI-g Mini Kit (Qiagen), according to the manufacturer's instructions using 5 μl (approximately 5–10 ng) of starting DNA in solution.
Reductive dechlorinating chemostat and batch cultures
A reductive dechlorinating microbial consortium was grown in a chemostat amended with tetrachloroethene (PCE) and lactate. Material from this chemostat was removed and incubated in batch with H2 and various electron acceptor combinations, before being harvested for RNA extraction and analysis with Hydrogenase Chip version 3. The Supplementary Methods detail chemostat and batch culture maintenance.
Phototrophic microbial mats
Cores were taken from microbial mat pieces subjected to a full-diel cycle on 12 and 13 November 2009. Extensive molecular and biogeochemical investigations of this phototrophic microbial mat will be discussed in detail in a future publication. Experimental details for mat collection, incubation and sampling are provided in the Supplementary Microbial Mat methods section.
DNA/RNA co-isolation
Details of DNA and RNA co-extraction are described in Supplementary Methods.
DNA labeling and hybridization
DNA was labeled and hybridized to DNA microarrays using a method similar to TIGR protocol M009 (Kim et al., 2002). For details, see the Supplementary Methods.
RNA amplification, labeling and hybridization
The RNA amplification and labeling protocol was based on the whole-community RNA amplification protocol (Gao et al., 2007). Details are provided in the Supplementary Methods.
PCR, cloning and sequencing of dsrA and Desulfovibrio sp. hynA-1
Fragments of dsrA for all bacteria (Leloup et al., 2009) and the (NiFe)-hydrogenase hynA-1 for Desulfovibrio sp. were amplified by PCR, cloned and sequenced. Sequence analysis was carried out using BLAST (Altschul et al., 1990), MUSCLE alignment (Edgar, 2004) and PHYML (Guindon and Gascuel, 2003). This procedure is delineated in the Supplementary Methods.
Reverse transcriptase—quantitative PCR analysis of Dehalococcoides hupL
Refer to the Supplementary Methods for details of the qPCR method used to assess Dehalococcoides hupL transcript abundance.
DNA microarray data analysis
The DNA microarrays were analyzed using Feature Extraction software version 9.5.3 and included protocol GE1-v5_95_Feb07 (Agilent). Numeric spot intensity data were processed using the R Foundation for Statistical Computing (2008) package TilePlot version 1.2.1 developed as part of this study for analysis of the functional gene-tiling microarrays. This package has been deposited to the Comprehensive R Archive Network (CRAN—http://cran.r-project.org/). The bright-probe cutoff used in TilePlot was three times the median intensity of all spots on the array, with spots brighter than this cutoff defined as bright and spots dimmer than this cutoff defined as dim. The bright-probe fraction (BPF) for each gene was defined as the number of bright probes for the gene divided by the total number of gene probes. For multiple array comparisons, median intensities for each probe on the array were fed into the tileplot.double() function, with each sample loess-normalized to a common reference sample in a fashion similar to conventional microarray analysis (Smyth and Speed, 2003). For the five-genome hybridization in the Test Array and other experiments, for which no quantitative comparison between the arrays was performed, the tileplot.single() function (no normalization or multiple array comparison) was used. Further details about which arrays were used as the normalization standards and how significant gene abundance differences between the samples were determined, are described in the Supplementary Methods section.
Results
Test microarray
The Test Microarray was used to examine the specificity of the tiling DNA microarray approach. This array was intended only to broadly assess whether a tiling DNA microarray could detect functional gene sequences in a mixed microbial community while avoiding false positive gene identification. For this reason, both hydrogenase genes and genes similar to hydrogenase genes were included in the array design when these genes were identified by BLAST.
In order to determine the sensitivity of the tiling DNA microarray technique to false positives, we hybridized a mixture of genomic DNA from five different bacteria to the Test Microarray (the Gammaproteobacteria Escherichia coli, Shewanella oneidensis, Shewanella sediminis, Pseudomonas aeruginosa and the Gram-positive Bacillus subtilis). These genomes were selected to represent both closely and distantly related microorganisms. Hybridization signals observed from the genes printed on the Test Microarray were then ranked according to their respective bright-probe fraction (BPF), and plotted as a ‘BPF rank curve’ (Figure 1a, Supplementary Table S2). We expected to accurately detect only 49 out of 845 genes on the array, because only those 49 genes were present in the genomes of the sample mixture. As Table 2 shows, all 49 expected genes yielded BPF values greater than 90%. Almost all unexpected array genes yielded BPF values below 90%. Two unexpected genes were identified with BPF values greater than 90%, and thus were considered as false positives. These were two Salmonella enterica nuoC genes, which had 89% DNA sequence identity with Escherichia coli nuoC. This result is consistent with the observations of mismatch probe hybridization intensities, which demonstrated insignificant changes in fluorescence intensity between 100% and 98% sequence identity, but much poorer intensities for probes with 82% sequence identity (Supplementary Table S1). This suggests that cross hybridization is possible for target sequences greater than some undetermined identity threshold above 82%. No significant effect of mismatch position was observed. These results show that genes known to be present in a microbial community can be accurately and unambiguously detected in a moderately complex microbial community using the tiling DNA microarray approach.
BPF rank curves for all microarray hybridizations in this study. (a) Test array hybridized with a mixture of five bacterial genomes, (b) reductive dechlorinating soil column with genes shared from Hydrogenase Chip versions 2 and 3, (c) reductive dechlorinating PM5L chemostat and batch cultures P, S and SP and (d) phototrophic microbial mat DNA and RNA (3 × cutoff multiplier).
Sensitivity analysis
To determine the lowest possible quantity of DNA necessary for the identification of a gene, DNA from four different genomes were mixed together at concentrations from 0.1 to 100 ng, then subjected to MDA and hybridized to the Hydrogenase Chip version 3. We found that the lowest abundance at which a gene was confidently detected with a BPF >90% was in the range between 1 and 10 ng of genomic DNA, or between 0.9% and 9% of the total DNA added to the MDA reaction (see Supplementary Table S11).
Tracing hydrogenase genes in a reductive dechlorinating soil column
To test whether the hydrogenase genes present in an undefined microbial community can be identified by the Hydrogenase Chip, DNA samples from three different time points of a previously described reductive dechlorinating soil column (Azizian et al., 2010) were analyzed using the Hydrogenase Chip. Each time point represented the steady state of a different amendment of electron donor to the soil column, with formate in March 2008, lactate in July 2008 and propionate in January 2009. The soil column operated over 1050 days, corresponding to two versions of the Hydrogenase Chip. Time points representing lactate and formate amendment were analyzed using version 1, and version 2 was used for the time point representing propionate amendment. This ensured that the sample was evaluated using the most up-to-date set of hydrogenase gene sequences according to the genomic and metagenomic databases. For comparative analyses of hydrogenase genes present in multiple samples, only the subset of 20 957 probes (targeting 458 genes) common to both the Hydrogenase Chip versions were analyzed (Supplementary Table S3).
Of the 458 genes represented by the probes common to both the microarray designs, 14 genes show hybridizations with a bright-probe fraction (BPF) >90% in at least one of the three samples (Table 3, Supplementary Table S3). Log intensity ratios for all the samples are shown in Figure 2. For the detected Dehalococcoides sp. hydrogenase genes, some showed significant differences between the probe intensities for the propionate and formate amendments. In general, genes from the Dehalococcoides strain CBDB1 genome were enriched in the propionate amendment, genes from the genome of Dehalococcoides ethenogenes strain 195 showed no significant difference between the two treatments and some Dehalococcoides strain VS genes showed enrichment while others did not. Although there is bound to be significant cross hybridization between all the Dehalococcoides sp. strains due to 90%+ sequence identity, the consistent abundance difference for the hydrogenase genes of strains 195 and CBDB1 compellingly suggests differences in the Dehalococcoides community structure between the time points representative of formate and propionate amendment. The lactate time point showed generally diminished probe intensities for all the Dehalococcoides hydrogenase genes, consistent with a diminished fraction of electrons contributing toward reductive dehalogenation with lactate as an electron donor. Chemical measurements of the column effluent showed that propionate and formate resulted in 10% and 14% of electron equivalents contributing to chloroethene reduction, respectively, while only 6.5% of the electron equivalents contributed to chloroethene reduction under lactate-oxidizing conditions (Azizian et al., 2010). There was a significant increase in the intensity of Geobacter hydrogenase genes in samples from the lactate- and propionate-amended time points relative to the formate-amended time point. This is consistent with an increase in the fraction of electrons partitioned to Fe(III) reduction in this system during the propionate or lactate amendment. Measurement of soluble Fe(II) in the column effluent accounted for 2.0% and 1.6% of electron equivalents under propionate- and lactate-oxidizing conditions, respectively, while accounting for 1.1% of the electron equivalents under formate-oxidizing conditions (Azizian et al., 2010). As the majority of Fe(II) was likely bound in the solid phase and thus would not have entered the effluent, these figures should be interpreted in relative terms. The detected Desulfitobacterium hafniense (NiFe) hydrogenase showed significantly reduced probe intensities under propionate amendment relative to formate, but no significant difference between the lactate and formate amendments. This is consistent with the fact that to date no pure culture isolate of Desulfitobacterium sp. has been shown to use propionate as an electron donor substrate, in contrast to formate and lactate (Utkin et al., 1994; Bouchard et al., 1996; Christiansen and Ahring, 1996; Sanford et al., 1996; Finneran et al., 2002).
Log intensity ratios for hydrogenase genes with BPF >90% observed in the reductive dechlorinating soil column. Positive values (to the right) signify greater abundance in lactate or propionate, negative values (to the left) signify greater abundance in formate relative to either propionate or lactate. Error bars show median absolute deviation, P-values show the probability that the two compared samples are equal to one another according to the binomial test.
Notably, no hydrogen-producing fermenting microorganisms were detected in any of the samples analyzed from the soil column. However, Geobacter sp. detected in the lactate- and propionate-amended soil column may be fermenting in this environment, as Geobacter isolates have been shown to syntrophically produce hydrogen during the fermentation of organic compounds in the past (Cord-Ruwisch et al., 1998).
These analyses also showed that gene richness trends between the different treatments were reflected by the species richness trends as observed within the 16S rRNA gene clone libraries (Azizian et al., 2010). A ranking in the hydrogenase gene richness in the order lactate>propionate>formate was revealed by the array in terms of the relative positions of BPF rank curves (Figure 1b) and numbers of genes with a BPF>90% (14 genes for lactate>13 genes for propionate>12 genes for formate, see Table 3, Supplementary Table S3). Notably, all the hydrogenase genes identified using the Hydrogenase Chip belonged to genera that were also identified in the 16S rRNA gene clone libraries.
We also detected genes that were not common to both microarray designs. Of the 481 hydrogenase and formate metabolism genes unique to the Hydrogenase Chip version 1 used to analyze the formate and lactate time points, 27 genes had BPF values >90% (Supplementary Table S4). In all, 19 of these genes were from Desulfitobacterium sp. genomes, five from Dehalococcoides sp. genomes and three from Geobacter sp. genomes. Of the 540 hydrogenase and reductive dehalogenase genes unique to the Hydrogenase Chip version 2 used for the propionate time point, 19 had BPF values >90% (Supplementary Table S5). These were hydrogenase and reductive dehalogenase genes exclusively from the Dehalococcoides sp. and Desulfitobacterium sp genomes. The detection of genes other than hydrogenase genes, such as those involved in the formate metabolism and reductive dehalogenation, from the same genera as the found hydrogenase genes suggests that the tiling platform can be used for gene categories other than hydrogenase genes.
Tracing hydrogenase gene expression in reductive dehalogenating batch cultures
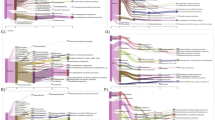
To more accurately characterize the community of hydrogen-producing and consuming microorganisms in reductive dehalogenating systems, we decided to determine which hydrogenase genes were not only present as DNA, but also transcribed. Thus, for evaluating the performance of the Hydrogenase Chip for detecting gene expression, we used undefined reductively dehalogenating mixed cultures in chemostat and batch reactor experiments. This is because preliminary data showed that abundant high-quality RNA could not be extracted from the Biosep beads used to sample the soil column (data not shown). The long-term anoxic chemostat was amended with lactate and tetrachloroethene (PCE), while the batch cultures derived from it were maintained for 44 days with H2 and one of the three electron acceptor conditions hypothesized to correlate with different hydrogenase gene expression patterns. Batch sample P was amended with PCE only, sample S with sulfate only and sample SP with both sulfate and PCE. On day 44, every bottle was incubated with both PCE and sulfate for 1 day before being harvested for molecular analysis, in order to simulate three different moderately complex microbial ecosystems undergoing simultaneous sulfate and PCE reduction, each optimized for different rates of both sulfate and PCE reduction (Figure 3).
To examine the hydrogenase genes expressed under these three conditions, we first used our DNA microarray to identify the hydrogenase genes present in the ancestral chemostat and acclimated batch cultures. Of the 1324 hydrogenase genes printed on the array, 36 yielded BPF >90% in at least one of the samples (see Table 4 and Supplementary Table S6). In all, 35 of these were from Dehalococcoides sp. and one from Desulfitobacterium hafniense. The apparent absence of hydrogenase genes from known sulfate-reducers was noteworthy, considering the observation of sulfate reduction in the derived batch cultures under H2-oxidizing conditions. This could be subsequently explained by a dsrA clone library of sample S and hydrogenase sequencing to determine sulfate-reducing members of the community. We found that all dsrA sequences appeared to be derived from Desulfovibrio sp. relatives (Supplementary Figure S1). This led us to sequence the Desulfovibrio hynA-1 hydrogenase gene from the sample S, which we found to be only 67% identical to the hydrogenase gene on the array with which it shares the highest identity. This explains the apparent absence of hydrogenase genes from the sulfate reducers.
We then examined the quantitative capabilities of the Hydrogenase Chip technique through measurement of the relative abundances of Dehalococcoides hupL in the acclimated batch cultures, using both the Hydrogenase Chip and reverse-transcriptase quantitative PCR (RT-qPCR). As expected, when examining the rates of electron acceptor consumption, sample P had become acclimated to high-PCE reduction rates and low-sulfate reduction rates, sample S to low-PCE reduction rates and high-sulfate reduction rates and sample SP to both sulfate and PCE reduction (Figure 3). We then investigated whether the observed rates correlated with shifts in abundance of Dehalococcoides hupL hydrogenase mRNA using the Hydrogenase Chip version 3. As Figure 4a shows, trends in Dehalocococcoides hupL median probe intensity followed trends in PCE transformation rate. This is consistent with earlier work showing hupL transcript abundance correlates with PCE respiration rates in Dehalococcoides (Rahm and Richardson, 2008). To independently assess hupL mRNA abundance to determine the accuracy of the Hydrogenase Chip, RT-qPCR targeting this gene was performed. As Figure 4b shows, shifts in mRNA abundance observed using the Hydrogenase Chip correlated with qPCR quantification of the abundance of Dehalococcoides hupL hydrogenase. The high median absolute deviation for the sample SP was likely caused by cross hybridization from a Dehalococcoides type more closely related to strain 195, as the median probe intensity strain 195 hupL increased significantly in sample SP relative to samples S and P. Apart from this aberration, the Hydrogenase Chip quantification appeared to correlate linearly with reverse transcriptase quantitative PCR measurements from the same samples.
hupL expression level as indicator for Dehalococcoides dehalogenation activity. Log intensity ratios of Dehalococcoides hupL with RNA hybridized to the Hydrogenase Chip version 3 compared with mean PCE transformation rates (a) and RT-qPCR (b). Gene abundance measurements show the natural logarithm of batch copy number (qPCR) or median probe intensity (Hydrogenase Chip) divided by the corresponding measurement in the chemostat sample. PCE transformation rates in (a) were used without normalizing to chemostat rates, but the natural logarithm is shown for consistency with microarray and qPCR data. Vertical error bars show median absolute deviation from the array, horizontal error bars show standard deviation from three qPCR replicates or from PCE degradation rates measured in two replicates for each batch culture.
Tracing hydrogen production in phototrophic microbial mats
In order to test the Hydrogenase Chip on a much more complex microbial community, we used the array to address the question of H2 production in phototrophic microbial mats. During nighttime hours when oxygenic photosynthesis could not occur, the top layer of these mats became sufficiently anoxic to allow significant H2 production to take place; however, the ecological basis of this H2 release remained unclear. In order to test whether ecological, microbial or mechanistic insights into this observation can be gained by using the Hydrogenase chip, both DNA and RNA were extracted from samples taken from the upper 2 mm layer of a phototrophic microbial mat on 12 November 2009 under daylight conditions with minimal measurable hydrogen production (4.2±0.2 nmol H2 accumulated per cm3 of mat material at the 1200-hours time point) and under dark conditions with considerable hydrogen production (25.5±8.5 nmol H2 accumulated per cm3 of mat material at the 2000-hours time point, with the peak measured H2 concentration at 0700 hours the following day with 144.2±64.2 nmol H2 accumulated per cm3 of mat material), and hybridized to the Hydrogenase Chip version 3. In the subsequent analysis explained below, we learnt that gene identification stringency must be adjusted in highly diverse samples.
When total DNA was hybridized, four genes had a BPF >90% in either of the two samples (Table 5, Supplementary Table S8). However, none of these genes were shown to have RNA BPF values >90% (Table 4, Supplementary Table S7). Paradoxically, it is most likely the highly diverse nature of the microbial mat ecosystem (Ley et al., 2006) that leads to the apparently low richness in hydrogenase genes. A more complex community should lead to a greater degree of non-target cross hybridization and a higher median probe intensity for the entire array than for the reductive dehalogenating samples analyzed in this study. In order to compensate for the loss in sensitivity in highly diverse samples, we lowered the gene identification stringency by reducing the TilePlot median cutoff multiplier from 3 × to 2 × . This increased the number of genes identified in DNA to 31 and in RNA to 2 (Supplementary Tables S9 and S10). Many of these genes would be expected in the microbial mat environment based on their associated physiologies. However, lowering the cutoff multiplier below 3 × produced false positive results from the Test Microarray experiment described in this study, and therefore one would expect that genes identified in the mat with a cutoff multiplier of 2 × are likely to include false positives.
In order to determine changes in gene transcript abundance relative to organismal abundance, relative median probe intensities comparing the 2000-hours time point with the 1200-hours time point were calculated for DNA and RNA measurements for hydrogenase genes identified (Figure 5). Notably, the gene encoding the (NiFe) hydrogenase from Microcoleus chthonoplastes PCC 7420 was the only hydrogenase gene to show a significant intensity change between the 1200-hours and 2000-hours samples. The low H2 production 1200-hours time point was characterized by lower RNA abundance and higher DNA abundance, and the high H2 production 2000-hours time point was characterized by higher RNA abundance and lower DNA abundance. This upregulation of a (NiFe) hydrogenase provides evidence that Microcoleus sp. may be responsible for H2 production in this microbial mat system. If confirmed by further studies, this is the first demonstration of Microcoleus sp. producing significant amounts of H2 in a phototrophic microbial mat. The lowered DNA hybridization intensity at night is probably due to some aerobic microbes having migrated in the night from deeper layers into the upper layers toward atmospheric O2, thereby diluting the cyanobacteria in the uppermost layer of the mat, or due to downwards migration by the cyanobacteria themselves. Such diel migration has been previously described in similar mats (Bebout and Garcia-Pichel, 1995; Fourçans et al., 2006; Dillon et al., 2009).
RNA and DNA log intensity ratios for genes with DNA BPF >90% from the phototrophic microbial mat. Positive values (to the right) signify greater abundance in the 2000-hours time point (H2 producing) compared with the 1200-hours time point, negative values (to the left) show greater abundance in the 1200-hours time point. Error bars show median absolute deviation, P-values show the probability that the two compared samples are equal to one other according to the binomial test.
Discussion
We have developed a tiling DNA microarray technique for assessing the functional gene content and expression status of a mixed microbial community and evaluated this approach through the characterization of hydrogen production and consumption in several different microbial communities. We have demonstrated here that this tiling approach is resilient against false positives, although with the trade-off of increased potential for false negative results in microbial communities of greater complexity. The method is also semiquantitative, showing trends similar to those observed by the established qPCR methods. The Hydrogenase Chip is the most broadly targeted attempt to characterize the hydrogenase gene content of a microbial community to date. The tiling functional gene microarray technique may also prove useful for characterizing other gene categories. With a simple tiling approach to design and ink-jet printing, it is possible to rapidly and inexpensively re-design and adapt tiling functional gene microarrays for specific environments and focused questions.
Resilience against false positives
Through the labeling and hybridization of a defined genomic DNA mixture, we showed that the tiling approach is generally free of false positive gene detection. A BPF >90% was demonstrated as a robust measure of positive gene identification, with the clearly recognizable sharp drop off in the BPF rank curve around BPF=90% shown for all the microarrays in this study (Figures 1a–d). The Test Microarray did reveal an exception to this robustness where target genes share sequence identity with probe sequences at some threshold above 82% identity. This is an acceptable false positive threshold for an application like the Hydrogenase Chip, where the goal is to define the broader biogeochemical results of changes in hydrogen metabolism. Most relevant physiologically distinct groups of microorganisms are differentiated by greater than 18% nucleotide sequence difference.
Due to the complexity involved in constructing a realistically complex yet defined mixture of RNA, the risk of false positive gene identification during RNA hybridization was not assessed as it was for DNA. However, as even with a 2 × cutoff multiplier the microbial mat RNA hybridization yielded only a two-gene subset of the 31 genes identified in the microbial mat DNA hybridization, we saw no reason to believe that the RNA hybridization method would be more prone to false positives than the DNA method.
One finding of this study was that as community diversity increases, either specificity or sensitivity of gene detection must be traded-off due to the increased frequency of non-target cross hybridization. More cross hybridization from the complex microbial mat sample generated a higher median probe intensity for all probes on the array. This led in turn to a higher cutoff probe intensity for defining bright probes in the microbial mat arrays than that used for the reductive dehalogenation samples. It appears that the microbial mat analyzed in this study is above the upper limit of community diversity at which this technique can accurately characterize gene presence and expression. Microbial mats are some of the most diverse microbial ecosystems ever characterized (Ley et al., 2006), and this shortcoming should not be seen as necessarily applying in less diverse microbial ecosystems, like soil or water (Lozupone and Knight, 2007). For most other functional gene microarrays that do not involve a PCR gene enrichment step, array specificity and sensitivity is determined using very simple defined mixtures of RNA fragments or genomic DNA (He et al., 2010; Dugat-Bony et al., 2011). These results are then extrapolated to complex natural microbial ecosystems. We have shown here that for one of the most complex known microbial ecosystems, the phototrophic microbial mat, an unforgiving trade-off between specificity and sensitivity must be made. This effect is important to keep in mind when analyzing results from all DNA microarray platforms in molecular microbial ecology.
False negative results
Several cyanobacterial hoxH (NiFe) hydrogenase gene clone libraries were sequenced from the microbial mats. The hydrogenase genes (GenBank accessions JF816258–JF816271) identified in these clone libraries mostly possessed maximum nucleotide identity with array genes of 68.8–78.9%. Only two genes sequences from this library showed >80% identity with any genes on the array, these were two Microcoleus-related clones and were 81.8 and 93.3% identical to the Microcoleus chthonoplastes PCC 7420 (NiFe) hydrogenase identified by the microarray (Table 5). Consistent with our earlier results, it is expected that these low sequence identity genes identified in the clone library would not be detected by the Hydrogenase Chip, as they were well below the necessary sequence identity threshold to produce a BPF >90%.
A notable absence was that of any hydrogenase gene from a fermenting microorganism in both the reductive dechlorinating chemostat and the soil column. Although the Hydrogenase Chip successfully identified other important physiological groups in these ecosystems, we cannot rule out that hydrogenases, whose genes were not on the array due to the absence of their sequences in the databases, and thus were not detected, may have an important role in a given environment. An example is the non-detection of the Desulfovibrio hynA-1 in the sulfate-reducing batch cultures. This is also the case for the non-detection of any potential formate-oxidizing syntrophic hydrogen-producing microorganisms (Dolfing et al., 2008) in the formate-amended reductive dehalogenating column and chemostat.
These false negative results illustrate an inevitable consequence of the tiling microarray design, in that only genes in the environment with nucleotide sequences highly identical to genes printed on the array will be detected at a significant level of confidence. This drawback will become less pronounced as genome and metagenome sequencing continues and as higher density oligonucleotide microarrays are used to query a broader swathe of nucleotide sequence space.
Semiquantitative gene and gene transcript abundance measurement
The fluorescence intensity signals of the probes provided information about the major differences in gene abundance for both H2-consuming microbes in the reductively dechlorinating microcosms and H2-producing microbes in the phototrophic microbial mat. The trends revealed by these differences are consistent with quantification performed by quantitative PCR (qPCR). Relative abundances of hydrogenase genes amongst the different amendments matched trends observed by qPCR examination of the same DNA samples (Azizian et al., 2010) and RNA samples (this study). We were initially concerned that MDA amplification of DNA or whole-community RNA amplification for microarray analysis would disrupt the measurable gene abundance trends, but as the qPCR was performed on unamplified DNA and cDNA it appears that this concern was not realized. One quantification challenge that all users of DNA microarrays have faced is that different regions of a gene will produce different probe intensities for a given gene concentration, and that this variability is difficult to predict computationally (Bruun et al., 2007; Dugat-Bony et al., 2011). In this study, we circumvented this problem by tiling all regions of all the genes on the array and then assessing the hybridization of each gene as a whole while estimating the variability with the median absolute deviation.
References
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Azizian MF, Marshall IPG, Behrens S, Spormann AM, Semprini L . (2010). Comparison of lactate, formate, and propionate as hydrogen donors for the reductive dehalogenation of trichloroethene in a continuous-flow column. J Contam Hydrol 113: 77–92.
Bebout BM, Garcia-Pichel F . (1995). UV B-induced vertical migrations of cyanobacteria in a microbial mat. Appl Environ Microbiol 61: 4215–4222.
Bodrossy L, Stralis-Pavese N, Murrell JC, Radajewski S, Weilharter A, Sessitsch A . (2003). Development and validation of a diagnostic microbial microarray for methanotrophs. Environ Microbiol 5: 566–582.
Bouchard B, Beaudet R, Villemur R, McSween G, Lépine F, Bisaillon JG . (1996). Isolation and characterization of Desulfitobacterium frappieri sp. nov., an anaerobic bacterium which reductively dechlorinates pentachlorophenol to 3-chlorophenol. Int J Syst Bacteriol 46: 1010–1015.
Boyd E, Spear J, Peters J . (2009). [FeFe]-hydrogenase genetic diversity provides insight into molecular adaptation in a saline microbial mat community. Appl Environ Microbiol 75: 4620–4623.
Bruun GM, Wernersson R, Juncker AS, Willenbrock H, Nielsen HB . (2007). Improving comparability between microarray probe signals by thermodynamic intensity correction. Nucleic Acids Res 35: e48.
Bryant MP, Wolin EA, Wolin MJ, Wolfe RS . (1967). Methanobacillus omelianskii, a symbiotic association of two species of bacteria. Archiv für Mikrobiologie 59: 20–31.
Christiansen N, Ahring BK . (1996). Desulfitobacterium hafniense sp. nov., an anaerobic, reductively dechlorinating bacterium. Int J Syst Evol Microbiol 46: 442.
Cord-Ruwisch R, Lovley D, Schink B . (1998). Growth of geobacter sulfurreducens with acetate in syntrophic cooperation with hydrogen-oxidizing anaerobic partners. Appl Environ Microbiol 64: 2232–2236.
DeSantis TZ, Brodie EL, Moberg JP, Zubieta IX, Piceno YM, Andersen GL . (2007). High-density universal 16S rRNA microarray analysis reveals broader diversity than typical clone library when sampling the environment. Microb Ecol 53: 371–383.
Dillon JG, Miller S, Bebout B, Hullar M, Pinel N, Stahl DA . (2009). Spatial and temporal variability in a stratified hypersaline microbial mat community. FEMS Microbiol Ecol 68: 46–58.
Dolfing J, Jiang B, Henstra AM, Stams AJM, Plugge CM . (2008). Syntrophic growth on formate: a new microbial niche in anoxic environments. Appl Environ Microbiol 74: 6126–6131.
Dugat-Bony E, Missaoui M, Peyretaillade E, Biderre-Petit C, Bouzid O, Gouinaud C et al. (2011). HiSpOD: probe design for functional DNA microarrays. Bioinformatics 27: 641–648.
Edgar R . (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797.
Finneran KT, Forbush HM, VanPraagh CVG, Lovley DR . (2002). Desulfitobacterium metallireducens sp. nov., an anaerobic bacterium that couples growth to the reduction of metals and humic acids as well as chlorinated compounds. Int J Syst Evol Microbiol 52: 1929–1935.
Fourçans A, Solé A, Diestra E, Ranchou-Peyruse A, Esteve I, Caumette P et al. (2006). Vertical migration of phototrophic bacterial populations in a hypersaline microbial mat from Salins-de-Giraud (Camargue, France). FEMS Microbiol Ecol 57: 367–377.
Gao H, Yang ZK, Gentry TJ, Wu L, Schadt CW, Zhou J . (2007). Microarray-based analysis of microbial community RNAs by whole-community RNA amplification. Appl Environ Microbiol 73: 563–571.
Gattiker A, Gasteiger E, Bairoch A . (2002). ScanProsite: a reference implementation of a PROSITE scanning tool. Appl Bioinformatics 1: 107–108.
Guindon S, Gascuel O . (2003). A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52: 696–704.
He Z, Deng Y, Van Nostrand JD, Tu Q, Xu M, Hemme CL et al. (2010). GeoChip 3.0 as a high-throughput tool for analyzing microbial community composition, structure and functional activity. ISME J 4: 1167–1179.
He Z, Gentry TJ, Schadt CW, Wu L, Liebich J, Chong SC et al. (2007). GeoChip: a comprehensive microarray for investigating biogeochemical, ecological and environmental processes. ISME J 1: 67–77.
Hoehler TM . (2005). Biogeochemistry of dihydrogen (H2). Met Ions Biol Syst 43: 9–48.
Hoehler TM, Albert DB, Alperin MJ, Bebout BM, Martens CS, Des Marais DJ . (2002). Comparative ecology of H2 cycling in sedimentary and phototrophic ecosystems. Antonie Van Leeuwenhoek 81: 575–585.
Hoehler TM, Bebout BM, Des Marais DJ . (2001). The role of microbial mats in the production of reduced gases on the early Earth. Nature 412: 324–327.
Hughes TR, Mao M, Jones AR, Burchard J, Marton MJ, Shannon KW et al. (2001). Expression profiling using microarrays fabricated by an ink-jet oligonucleotide synthesizer. Nat Biotechnol 19: 342–347.
Karkhoff-Schweizer RR, Huber DP, Voordouw G . (1995). Conservation of the genes for dissimilatory sulfite reductase from Desulfovibrio vulgaris and Archaeoglobus fulgidus allows their detection by PCR. Appl Environ Microbiol 61: 290–296.
Kim H, Zhao B, Snesrud EC, Haas BJ, Town CD, Quackenbush J . (2002). Use of RNA and genomic DNA references for inferred comparisons in DNA microarray analyses. Biotechniques 33: 924–930.
Leloup J, Fossing H, Kohls K, Holmkvist L, Borowski C, Jørgensen BB . (2009). Sulfate-reducing bacteria in marine sediment (Aarhus Bay, Denmark): abundance and diversity related to geochemical zonation. Environ Microbiol 11: 1278–1291.
Ley RE, Harris JK, Wilcox J, Spear JR, Miller SR, Bebout BM et al. (2006). Unexpected diversity and complexity of the Guerrero Negro hypersaline microbial mat. Appl Environ Microbiol 72: 3685–3695.
Li W, Godzik A . (2006). Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22: 1658–1659.
Lozupone CA, Knight R . (2007). Global patterns in bacterial diversity. Proc Natl Acad Sci USA 104: 11436–11440.
Markowitz VM, Ivanova NN, Szeto E, Palaniappan K, Chu K, Dalevi D et al. (2008). IMG/M: a data management and analysis system for metagenomes. Nucleic Acids Res 36: D534–D538.
Miller SM, Tourlousse DM, Stedtfeld RD, Baushke SW, Herzog AB, Wick LM et al. (2008). An in-situ synthesized virulence and marker gene (VMG) biochip for the detection of bacterial pathogens in water. Appl Environ Microbiol 74: 2200–2209.
Ohkuma M, Noda S, Horikoshi K, Kudo T . (1995). Phylogeny of symbiotic methanogens in the gut of the termite Reticulitermes speratus. FEMS Microbiol Lett 134: 45–50.
Palmer C, Bik EM, Eisen MB, Eckburg PB, Sana TR, Wolber PK et al. (2006). Rapid quantitative profiling of complex microbial populations. Nucleic Acids Res 34: e5.
Pozhitkov AE, Nies G, Kleinhenz B, Tautz D, Noble PA . (2008). Simultaneous quantification of multiple nucleic acid targets in complex rRNA mixtures using high density microarrays and nonspecific hybridization as a source of information. J Microbiol Methods 75: 92–102.
R Foundation for Statistical Computing (2008). R Foundation for Statistical Computing: Vienna, Austria.
Rahm BG, Richardson RE . (2008). Correlation of respiratory gene expression levels and pseudo-steady-state PCE respiration rates in Dehalococcoides ethenogenes. Environ Sci Technol 42: 416–421.
Rich VI, Konstantinidis K, DeLong EF . (2008). Design and testing of ‘genome-proxy’ microarrays to profile marine microbial communities. Environ Microbiol 10: 506–521.
Roeselers G, Huisjes EH, van Loosdrecht MCM, Muyzer G . (2008). Diversity and expression of cyanobacterial hupS genes in pure cultures and in a nitrogen-limited phototrophic biofilm. FEMS Microbiol Ecol 63: 292–300.
Sanford R, Cole J, Loffler F, Tiedje J . (1996). Appl Environ Microbiol 62: 3800–3808.
Smidt H, de Vos WM . (2004). Anaerobic microbial dehalogenation. Annu Rev Microbiol 58: 43–73.
Smyth GK, Speed T . (2003). Normalization of cDNA microarray data. Methods 31: 265–273.
Stams AJM, Plugge CM . (2009). Electron transfer in syntrophic communities of anaerobic bacteria and archaea. Nat Rev Microbiol 7: 568–577.
Taroncher-Oldenburg G, Griner EM, Francis CA, Ward BB . (2003). Oligonucleotide microarray for the study of functional gene diversity in the nitrogen cycle in the environment. Appl Environ Microbiol 69: 1159–1171.
Thauer RK, Kaster A-K, Goenrich M, Schick M, Hiromoto T, Shima S . (2010). Hydrogenases from methanogenic archaea, nickel, a novel cofactor, and H2 storage. Annu Rev Biochem 79: 507–536.
Utkin I, Woese C, Wiegel J . (1994). Isolation and characterization of Desulfitobacterium dehalogenans gen. nov., sp. nov., an anaerobic bacterium which reductively dechlorinates chlorophenolic compounds. Int J Syst Bacteriol 44: 612–619.
Vignais PM . (2008). Hydrogenases and H(+)-reduction in primary energy conservation. Results Probl Cell Differ 45: 223–252.
Vignais PM, Billoud B . (2007). Occurrence, classification, and biological function of hydrogenases: an overview. Chem Rev 107: 4206–4272.
Vignais PM, Billoud B, Meyer J . (2001). Classification and phylogeny of hydrogenases. FEMS Microbiol Rev 25: 455–501.
Wolber PK, Collins PJ, Lucas AB, De Witte A, Shannon KW . (2006). The Agilent in situ-synthesized microarray platform. Meth Enzymol 410: 28–57.
Wolin M, Miller T . (1982). Interspecies hydrogen transfer: 15 years later. ASM American Society for Microbiology News 48: 561–565.
Xing D, Ren N, Rittmann BE . (2008). Genetic diversity of hydrogen-producing bacteria in an acidophilic ethanol-H2-coproducing system, analyzed using the [Fe]-hydrogenase gene. Appl Environ Microbiol 74: 1232–1239.
Yang Y, McCarty PL . (1998). Competition for hydrogen within a chlorinated solvent dehalogenating anaerobic mixed culture. Environ Sci Technol 32: 3591–3597.
Yu S, Dolan M, Semprini L . (2005). Kinetics and inhibition of reductive dechlorination of chlorinated ethylenes by two different mixed cultures. Environ Sci Technol 39: 195–205.
Acknowledgements
This work was supported by the Strategic Environmental Research Defense Project (SERDP) to AMS by Grant ER-1588. IPGM was supported by a Bio-X Graduate Research Fellowship through the Stanford University. We gratefully acknowledge useful comments by the editor and anonymous reviewers, useful discussions with Dagmar Woebken and a useful discussion of statistics with Gerald Teng and Jim Mussared. We also thank Shirley Galbiati for assistance in performing the sensitivity analysis hybridization.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Marshall, I., Berggren, D., Azizian, M. et al. The Hydrogenase Chip: a tiling oligonucleotide DNA microarray technique for characterizing hydrogen-producing and -consuming microbes in microbial communities. ISME J 6, 814–826 (2012). https://doi.org/10.1038/ismej.2011.136
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.136
Keywords
This article is cited by
-
Proposal of Thermoactinomyces mirandus sp. nov., a filamentous, anaerobic bacterium isolated from a biogas plant
Antonie van Leeuwenhoek (2021)
-
Review of biological diagnostic tools and their applications in geoenvironmental engineering
Reviews in Environmental Science and Bio/Technology (2015)
-
Anoxic carbon flux in photosynthetic microbial mats as revealed by metatranscriptomics
The ISME Journal (2013)