Abstract
In gene-centric comparative metagenomics, differences in observed relative gene abundances among samples are often assumed to reflect the biological importance of individual genes in different habitats. Statistical tests and data mining for genes that represent habitat-specific adaptations are frequently based on this measure. We demonstrate that this measure is biased by the average genome size of the communities sampled. Average genome sizes can be estimated from the metagenomic data themselves, and taken into account in comparative analyses. We suggest that this would enable ecologically more meaningful comparisons, especially when the average genome sizes of compared communities differ substantially. We illustrate the influence of average genome-size differences on comparative analyses, with an example to highlight the need for further exploration of this bias.
Similar content being viewed by others
Main
Shotgun sequencing of total community DNA is becoming an established part of the methodological arsenal of microbial ecology. Owing to its relative novelty and the high complexity of the data it produces, metagenomics is still in an explorative phase in many ways, and several aspects of data acquisition and analysis are just gaining proper attention and treatment (Gomez-Alvarez et al., 2009; Kunin et al., 2009). Here we draw attention to a currently underappreciated aspect of gene-centric comparative metagenomic data analyses: the effect of average genome sizes on sampling probabilities of gene fragments.
In gene-centric comparative metagenomic analyses, sequences are classified into functional categories, and the relative abundances of reads belonging to different categories are compared across samples. The statistical approaches proposed for analyzing these data usually test the null hypothesis that sampling probabilities of genes are equal across samples (Rodriguez-Brito et al., 2006; Kristiansson et al., 2009; Li, 2009). Such analyses often account for differences in sampling effort between samples, and sometimes for average gene length, although the latter property is usually assumed to be relatively constant across samples. Using a simple mathematical model of metagenomic sampling, we demonstrate that sampling probabilities of individual genes (pm), pooled across all taxa present in a community, are expected to depend on the average genome sizes of the samples compared (Ḡ), and on the gene length (lm) and average copy number of the gene concerned (C̄m):

Detailed arguments supporting this equation can be found in the Supplementary Information. Previously, Raes et al. (2007a) noted that this issue was likely to be important, without attempting to analyze its impact on comparative sequence analysis.
In practical terms, this means that even universally distributed single-copy genes will show apparent differences in relative abundances among samples if average genome sizes differ substantially across samples. Thus, in principle, statistically significant, but biologically uninteresting, variation can emerge from analyses that fail to account for differences in the average genome sizes of communities. A number of recent publications have described methods for estimating the average genome size of communities and have demonstrated substantial genome-size variation among communities, for example at different geographical locations or depths in marine environments (Raes et al., 2007b; Angly et al., 2009; Konstantinidis et al., 2009).
To illustrate, we considered a metagenomic data set from the Northern Pacific (DeLong et al., 2006). The number of reads sequenced from each sample varied between 6812 and 11 479 (Supplementary Figure 1a). The estimated genome sizes of these samples ranged from 3.36 to 5.56 million base pairs (Supplementary Figure 1b), somewhat higher than the values previously estimated for other oceanic metagenomic samples (Raes et al., 2007a). Notably, the sample from 10 m depth had a markedly lower estimated average genome size than the rest of the samples, increasing the probability of sampling any gene category in this sample compared with the others. We combined the effects of total numbers of sequences and estimated average genome sizes into a composite measure, which we termed as the effective sequence count (ESC, see Supplementary Information for mathematical details). ESCs ranged from 6305 to 11 393 ‘pseudo-reads’ (see Supplementary Figure 1c), suggesting that an approximately 1.8-fold difference is expected in the sampling probabilities of universal single-copy genes between the 70- and 200-m samples, representing the two extremes. Although the data set concerned is too small for confidently detecting sampling probability differences of this order of magnitude, when testing the above 35 universal single-copy cluster of orthologous groups (COGs) individually, six of them even showed significant trends with depth at an (uncorrected) α-level of 5% when average genome sizes were ignored.
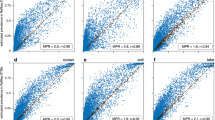
To illustrate the potential effect of correcting for average genome-size differences on apparent trends in gene abundance, we used COG0415 (deoxyribopyrimidine photolyase), which was featured previously to illustrate a significant trend with depth in the same data set (DeLong et al., 2006; Kristiansson et al., 2009). Notably, the equal total counts of fragments annotated as COG0415 at the surface and at 70 m depth (Figure 1a) turn into a decrease at 70 m when a normalization by sequencing effort is applied (Figure 1b), and into an increase when both sequencing effort and community-averaged genome size are taken into account for normalization (Figure 1c). Based on the mathematical arguments detailed in the Supplementary Information, we suggest that the trend observed in Figure 1b confounds differences among communities in average genome size (that is, differences that are not specific to the gene of interest) with differences in the average copy number of COG0415 per cell. Figure 1c gives a biologically more meaningful picture about the differential presence of this gene in genomes of organisms occurring at different depths by accounting for the former, not gene-specific, factor. We further explore the possible effects of community average genome-size differences on comparative analyses in the Supplementary Information.
Apparent relative abundances of COG0415 in the HOTS data set against depth using different standardizations. (a) Total counts of fragments annotated as COG0415; (b) the same, after correction for different sampling efforts, as quantified using number of reads sequenced per sample (this pattern changes little when using the total number of base pairs or the number of annotated reads per sample for normalization; the latter are not shown); (c) arbitrarily scaled relative average copy numbers of COG0415 against depth: ratios of gene counts and effective sequence counts (see Supplementary Information for the definition of the latter). Note in particular the different conclusions suggested by these different standardizations about COG0415 abundance at the surface vs at 70 m depth.
In conclusion, we propose to add community genome size to the list of potential biasing factors for gene-centric comparative metagenomics (ranging from sampling and sample preparation issues, and sequencing technologies, to annotation protocols, methods for measuring sampling effort, and gene length effects; see, for example, Raes et al., 2007a). Based on the simple mathematical considerations represented in Equation (1) and detailed in the Supplementary Information, we propose that decomposing relative gene abundance differences among metagenomic samples into ‘metagenome-wide’ and gene specific components—that is, differences in average genome sizes vs differences in relative gene-copy numbers—is expected to improve the biological relevance of inferences.
References
Angly FE, Willner D, Prieto-Davo A, Edwards RA, Schmieder R, Vega-Thurber R et al. (2009). The GAAS metagenomic tool and its estimations of viral and microbial average genome size in four major biomes. PLOS Comp Biol 5: e1000593.
DeLong EF, Preston CM, Mincer T, Rich V, Hallam SJ, Frigaard N-U et al. (2006). Community genomics among stratified microbial assemblages in the ocean's interior. Science 311: 496–503.
Gomez-Alvarez V, Teal TK, Schmidt TM . (2009). Systematic artifacts in metagenomes from complex microbial communities. ISME J 3: 1314–1317.
Konstantinidis KT, Braff J, Karl DM, DeLong EF . (2009). Comparative metagenomic analysis of a microbial community residing at a depth of 4000 meters at station ALOHA in the North Pacific subtropical gyre. Appl Environ Microbiol 75: 5345–5355.
Kristiansson E, Hugenholtz P, Dalevi D . (2009). ShotgunFunctionalizeR: an R-package for functional comparison of metagenomes. Bioinformatics 25: 2737–2738.
Kunin V, Engelbrektson A, Ochman H, Hugenholtz P . (2009). Wrinkles in the rare biosphere: pyrosequencing errors lead to artificial inflation of diversity estimates. Environ Microbiol. doi: 10 1111/j.1462-2920.2009.02051.x.
Li W . (2009). Analysis and comparison of very large metagenomes with fast clustering and functional annotation. BMC Bioinformatics 10: 359.
Raes J, Foerstner KU, Bork P . (2007a). Get the most out of your metagenome: computational analysis of environmental sequence data. Curr Opin Microbiol 10: 490–498.
Raes J, Korbel JO, Lercher MJ, von Mering C, Bork P . (2007b). Prediction of effective genome size in metagenomic samples. Genome Biol 8: R10.
Rodriguez-Brito B, Rohwer F, Edwards RA . (2006). An application of statistics to comparative metagenomics. BMC Bioinformatics 7: 162.
Acknowledgements
This work was supported by a Marine Microbiology Initiative Investigator award from the Gordon and Betty Moore Foundation and the Society of General Microbiology President's Fund.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Beszteri, B., Temperton, B., Frickenhaus, S. et al. Average genome size: a potential source of bias in comparative metagenomics. ISME J 4, 1075–1077 (2010). https://doi.org/10.1038/ismej.2010.29
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2010.29