Abstract
It was hypothesized that seasonality and resource availability altered through tree girdling were major determinants of the phylogenetic composition of the archaeal and bacterial community in a temperate beech forest soil. During a 2-year field experiment, involving girdling of beech trees to intercept the transfer of easily available carbon (C) from the canopy to roots, members of the dominant phylogenetic microbial phyla residing in top soils under girdled versus untreated control trees were monitored at bimonthly intervals through 16S rRNA gene-based terminal restriction fragment length polymorphism profiling and quantitative PCR analysis. Effects on nitrifying and denitrifying groups were assessed by measuring the abundances of nirS and nosZ genes as well as bacterial and archaeal amoA genes. Seasonal dynamics displayed by key phylogenetic and nitrogen (N) cycling functional groups were found to be tightly coupled with seasonal alterations in labile C and N pools as well as with variation in soil temperature and soil moisture. In particular, archaea and acidobacteria were highly responsive to soil nutritional and soil climatic changes associated with seasonality, indicating their high metabolic versatility and capability to adapt to environmental changes. For these phyla, significant interrelations with soil chemical and microbial process data were found suggesting their potential, but poorly described contribution to nitrification or denitrification in temperate forest soils. In conclusion, our extensive approach allowed us to get novel insights into effects of seasonality and resource availability on the microbial community, in particular on hitherto poorly studied bacterial phyla and functional groups.
Similar content being viewed by others
Introduction
Trees release large proportions of their accumulated carbon (C) and nitrogen (N) in the form of tree residues (for example, litter, dead roots) and root exudates to the soil organic matter (Yarwood et al., 2009; Fontaine et al., 2004). The soil organic matter pool provides an important energy source for soil microorganisms, which are the major performing agents in decomposition and soil organic matter transformation, the key processes in terrestrial C and N cycling (Buckley and Schmidt, 2002). Quantity and quality of available C and N control soil microbial population dynamics and microbial processes including nitrification or denitrification (Schimel and Weintraub, 2003; Magill and Aber, 2000). In particular, it was shown that microbial community structures were shaped by N cycle dynamics in forest soils (Högberg et al., 2007; Lejon et al., 2005; Grayston and Prescott, 2005).
Belowground C and N transfer, shaped by trees and through microbial processes, is influenced by external factors such as seasonality. Seasonally alternating climatic conditions take a decisive control on tree physiology, photosynthesis and discharge of C and N into soil (Cannell and Dewar, 1994; Waring and Running, 1998). It can be concluded that cyclic changes in tree physiology have a significant influence on soil microbial communities. In addition, temporal variability in the soil microbial community composition was shown in response to seasonal variation in temperature, moisture and plant activity (Koch et al., 2007; Waldrop and Firestone, 2006; Horz et al., 2004; Buckley and Schmidt, 2002).
Seasonal and other temporal alternations in climatic conditions are determinants of soil N cycle dynamics (Cookson et al., 2006; Wolsing and Priemé, 2004; Horz et al., 2004). Soil N cycling includes both reductive and oxidative processes, in which soil microbes have a predominant role (Cabello et al., 2004). Key microbial processes within the soil N cycle are catalyzed by key enzymes, including amoA gene encoding a subunit of ammonia monooxygenase in nitrification, as well as nirS and nirK gene (nitrite reductases) and nosZ gene (nitrous oxide (N2O) reductase) involved in denitrification. The diversity and abundance of microorganisms carrying these genes and the actual link to N2O emission have been extensively studied in diverse soil ecosystems (for example, Henderson et al., 2010, Leininger et al., 2006; Philippot et al., 2006).
Tree girdling is a procedure to remove the bark and phloem from a tree down to the youngest xylem effectively excluding rhizodeposition into soil and thus restricting resource availability without disturbing the soil-root-microbe ecosystem (Högberg et al., 2001). It was shown that manipulation of C and N availability by tree girdling leads to significant modifications in soil nutrient stoichiometry. Weintraub et al. (2007) measured lower dissolved organic C and N as well as an increase over time in nitrate and ammonium in girdled plots of a subalpine forest. Högberg et al. (2007) observed a tendency towards higher inorganic N levels in girdled plots of a boreal forest, whereas Ekberg et al. (2007) detected a decrease in total organic C in girdled plots of temperate spruce stand. It is thus likely that girdling-related changes in soil chemistry, in particular labile C and N pools have considerable effects on the soil microbial community structure (Dannenmann et al., 2009; Weintraub et al., 2007).
Although these examples show that soil microbial communities are influenced by C and N availability as well as by seasonality, the effects on different bacterial phyla or functional groups in temperate forest soils are still poorly understood. The objective of this study therefore was to assess in depth the microbial community response to C and N availability and to seasonal changes. A 2-year field experiment was carried out in a temperate beech forest (Klausenleopoldsdorf, Lower Austria). The major hypothesis was that seasonality along with altered C and N availability achieved through tree girdling control the community structure of the affected bacterial and archaeal population. Abundance and community structure of the total soil bacterial and archaeal population and specific phyla, that is, acidobacteria, alpha- and beta-proteobacteria and verrucomicrobia, as well as nitrifying and denitrifying microbial communities were investigated. Detected community changes were related to seasonality and tree girdling induced alterations in labile C and N pools as well as to soil moisture and soil temperature variations.
Materials and methods
Experimental site, samplings and geochemical data
The experimental study site was situated in a 65-year old beech forest (forest community Hordelymo-Fagetum with main tree species Fagus sylvatica L.) in Klausenleopoldsdorf (geographical location: 48°07′N, 16°03′E, 510 m above see level), Lower Austria, approximately 40 km southwest of Vienna. At the study site, representing an extensively managed forest-monitoring site according to the International Co-operative Programme on Assessment and Monitoring of Air Pollution Effects on Forests, a mean annual temperature of 7.6 °C and a mean annual precipitation of 768 mm were determined. The soil defined as a dystric cambisol had developed from the Laab formation (Eocene) and major geochemical properties were determined previously (pH value: 4.0; total organic C: 4.36%; total N: 0.33%, C-to-N ratio: 13.1) (Hackl et al., 2004).
The field experiment was started with girdling of trees in May 2006. Tree bark was removed at 10 cm length around the trunk at about 1.5 m above ground. Three girdling plots of 20 × 20 m were installed, of which only the inner 10 × 10 m were used for soil samplings. Six replicate control plots without tree girdling measuring 5 × 5 m were installed. Understory vegetation was removed from all plots and was repeated in the second spring season wherever necessary. During the whole experimental phase, soil temperature and soil moisture were measured continuously (Kaiser et al., 2010). Further details about tree vitality, leaf litter amount and quality as well as fine root biomass in girdled and control plots have been reported by Kaiser et al. (2010).
Sampling was performed bimonthly from May 2006 until May 2008. Two replicate samples were taken from each plot (six replicates in total), with four subsamples (soil cores of 10 × 10 × 5–10 cm, depending on the depth of A horizon) collected in each subplot. Generally, sampling was based on a predetermined sampling scheme to avoid sampling of already disturbed soil and to warrant independent soil samples throughout the experimental period. The four sub-samples of each plot were pooled, sieved through 2 mm mesh (5 mm mesh in case of wet soils) and stored at −20 °C. C and N pool data used for statistical purposes in this study were taken from Kaiser et al. (2010) and Kitzler et al. (unpublished data).
Microbial community structure (terminal restriction fragment length polymorphism (T-RFLP) analysis)
Bulk soil DNA was isolated (FastDNA Spin for Soil Kit, MP Biomedicals, Solon, OH, USA) and extracts were quantified photometrically (Nanodrop ND-1000, Nanodrop Technologies, Wilmington, DE, USA). Bacterial and archaeal 16S rRNA genes were PCR-amplified using primers sets targeting total bacteria and archaea as well as four selected bacterial phyla. Based on a soil 16S rRNA gene library of one hundred clones generated before start of the field experiment, acidobacteria (28%), alpha- and beta-proteobacteria (18% and 14%, respectively), as well as verrucomicrobia (16%) have been selected as the four most dominant bacterial community members in the soils of the studied experimental site (Supplementary Table S1; NCBI accession numbers HM364804 to HM364903). For T-RFLP analysis, all forward primers were labeled with 6-carboxyfluorescein at the 5′ ends. From each DNA sample, two replicate PCRs were done. Composition of PCR cocktails and amplification details are provided in Table 1. Amplicons (5 μl) were checked on ethidium bromide-stained 1% (w/v) agarose gels. Replicate amplicons were pooled, purified (Sephadex G-50, GE Healthcare Biosciences, Waukesha, WI, USA), and approximately 200 ng of each amplicon were digested with 5 U AluI (Invitrogen, Carlsbad, CA, USA) at 37 °C for 4 hrs. Before the T-RFLP analysis, digests were purified (Sephadex G-50) and an aliquot of 5 μl was mixed with 15 μl HiDi formamide (Applied Biosystems, Foster City, CA, USA) and 0.3 μl internal size standard (500 ROX Size Standard, Applied Biosystems). Labeled terminal-restriction fragments were denatured at 92 °C for 2 min, chilled on ice and detected on an ABI 3100 automatic DNA sequencer (Applied Biosystems) in the GeneScan mode. Gelquest software package (version 2.2.1, SequentiX, Klein Raden, Germany) was used to compare relative lengths of terminal-restriction fragments with the 500 ROX size standard and to compile electropherograms into a numeric data set, in which fragment length and peak height >50 fluorescence units were used for profile comparison. T-RFLP profiles used for statistical analyses were normalized according to Dunbar et al. (2000).
Microbial abundance (quantitative PCR (qPCR) analysis)
For standard preparation, amplicons from each investigated taxonomic group and functional genes (Table 2) were purified (Invisorb Spin PCRapid kit, Invitek, Berlin, Germany), ligated into the StrataClone PCR cloning vector pSC-A (Stratagene, La Jolla, CA, USA), and ligation products were transformed with StrataClone SoloPack competent cells (Stratagene). Specificity of clones used as qPCR standards were checked with Basic Local Alignment Search Tool. Plasmid DNA was isolated (Plasmid Miniprep Kit, Bio-Rad Laboratories, Hercules, CA, USA) and quantified as described above. For qPCR, each 25 μl PCR cocktail contained 12.5 μl iQ SybrGreen Supermix (Bio-Rad Laboratories), 0.4 μM of each oligonucleotide (Table 2), 1.0 mg ml−1 bovine serum albumin and 10 and 50 ng template DNA for taxonomic groups and functional genes, respectively. Apart from the bacterial amoA gene assay, all functional gene qPCRs were supplemented with 0.625 μl dimethyl sulfoxide. PCR reactions were run on an iCycler iQ Multicolor Real-time PCR Detection System (Bio-Rad Laboratories) and were started with 3 min at 95 °C, followed by amplification cycles specific for each phylum and functional gene (Table 2). Melting curve analysis of amplicons was conducted to confirm that fluorescence signals originated from specific amplicons and not from primer-dimers or other artifacts. Each DNA sample was processed in triplicate reactions, whereas standard curves were generated using duplicate 10-fold dilutions of isolated plasmid DNA. Automated analysis of PCR amplicon quality (for example, PCR baseline subtraction, Ct-threshold setting to the linear amplification phase) and quantity was performed with iCycler Optical System Software Version 3.1 (Bio-Rad Laboratories).
Statistical analyses
Analysis of variance combined with post hoc Tukey-B tests (SPSS for Windows, version 11.7, SPSS Inc., Chicago, IL, USA) was performed according to Rasche et al. (2006) to determine significant treatment effects (tree girdling, seasonality) on abundance and community structure of the investigated microbial groups and functional genes. Pearson's linear correlation coefficients were calculated for assessing significant relations between microbial abundance and geochemical parameters (SPSS for Windows). Effect of tree girdling on T-RFLP data sets obtained from each target group was further assayed based on Bray–Curtis similarity coefficients (Legendre and Legendre, 1998). Therefore, a similarity matrix was generated for all possible pairs of samples of each target group. This similarity matrix was used for analysis of similarity (ANOSIM) statistics (Clarke and Green, 1988) to test the hypothesis that bacterial, archaeal and taxonomic communities were altered by tree girdling over the investigation period. ANOSIM generates a test statistics, R. The magnitude of R indicates the degree of separation between two communities, with a score of 1 indicating complete separation and 0 indicating no separation. Calculation of similarity coefficients and ANOSIM were carried out using Primer six for Windows (version 6.1.5, Primer-E Ltd., Plymouth, UK). To test the influence of environmental variables on the microbial community structure, canonical correspondence analyses were carried out in Canoco (version 4.5 for Windows, PRI Wageningen, the Netherlands) (Lepš and Šmilauer, 2003). Presence or absence as well as relative height of terminal-restriction fragments were used as ‘species’ data whereas geochemical data were included in the analysis as ‘environmental’ variables. Resulting ordination biplots approximated the weighted differences between the individual communities (T-RFLP patterns) with respect to each of the geochemical factors, which were represented as arrows. The length of the corresponding arrows indicated the relative importance of the geochemical factor in explaining variation in the six microbial T-RFLP profiles, whereas the angle between arrows indicated the degree to which they were correlated. A Monte Carlo permutation test based on 1000 random permutations was used to calculate the impact of geochemical variables on community patterns.
Results
Microbial community structure (T-RFLP analysis)
Compared with tree girdling, seasonality had the greatest, significant influence on the total bacterial and archaeal community structure as well as on the four selected phyla (P<0.001) (Table 3). The community structure of the total archaea was changed by tree girdling (P<0.001), whereas total bacterial was not (P>0.05). Alpha-proteobacteria and acidobacteria showed a detectable community change due to tree girdling (P<0.01), whereas beta-proteobacteria and verrucomicrobia appeared not altered (P>0.05). A clear interaction between the two factors ‘seasonality’ and ‘tree girdling’ was determined indicating an interrelated influence of both factors on the community dynamics of the bacterial and archaeal community (P<0.05) (Table 3).
ANOSIM detected a distinct tree girdling-induced community change among archaea, which was confirmed by several significant R values ranging between 0.124 and 0.913 indicating distinct structural differences between two individual archaeal communities (control versus tree girdling) (P<0.05) (Table 4). Based on R values, greatest community differentiations became measurable during early fall and winter months. A comparable trend was determined for total bacteria, although the differences were less pronounced as compared with the total archaea explained by a smaller number of significant R values. For alpha-proteobacteria and acidobacteria, ANOSIM calculated several significant R values, whereas for beta-proteobacteria and verrucomicrobia only at three sampling dates significant tree girdling effects (P<0.05) were found.
Canonical correspondence analysis was used to test the significant dependence of the community differentiations on tree girdling and seasonality-related changes in geochemical parameters (Table 5, Figure 1). Multivariate testing, based on 1000 Monte Carlo permutations, confirmed the significance of the two canonical axes (P<0.05). The total percentage variance of the microbiota-environment relation ranged between 47.6% (verrucomicrobia) and 70.8% (total bacteria) (Table 5). The first canonical axis attributed the greatest influence explaining the microbiota-environment relation. The strong relation between the T-RFLP data sets and geochemical data was substantiated by high correlation coefficients of at least 0.390 (Table 5). Figure 1 illustrates the relationship between the community changes and geochemical data and shows further the clear seasonality and tree girdling related community differentiations overall the 2-year experimental period. Generally, higher soil moisture was determined in the second experimental year, whereas higher soil temperatures were measured in the first year of the field experiment (Kaiser et al., 2010). These soil climatic differentiations were clearly reflected in all six T-RFLP community patterns, in which distinct community shifts were observed between the first and the second year (Figure 1). The first year samples tended to cluster if higher soil temperatures and lower soil moisture were observed, whereas the opposite effect was determined for the second year samples. When explaining the effect of determined chemical parameters on assayed microbial communities, nitrate contents were in average highest in the first year of investigation and peaked during summer months (Kaiser et al., 2010). Dissolved organic nitrogen (DON) showed a clear decrease in the second year as compared to the first year, whereas no clear trend was observable for dissolved organic carbon (DOC) and ammonia (Kaiser et al., 2010). These changes in chemical parameters were clearly reflected in alterations within the soil microbial communities (Figure 1). In particular, the first year samples tended to cluster with high nitrate values, whereas the second year samples indicated a clear correlation with high DON values. In general, seasonality effect was overwhelming the effect of tree girdling.
Biplots of canonical correspondence analysis of the six individual 16S rRNA gene-based T-RFLP data sets obtained from (a) total bacteria, (b) total archaea, (c) alpha-proteobacteria, (d) beta-proteobacteria, (e) acidobacteria and (f) verrucomicrobia. Sampling years (Seasonality) are indicated with colors (2006, green; 2007, red; 2008, black), whereas control and tree girdling samples are shown with open and bold symbols, respectively. Geochemical data (soil moisture (Moist), soil temperature (Temp), soil nitrate (NO3), soil ammonia (NH4), dissolved soil organic carbon (DOC) and dissolved soil organic nitrogen (DON) are presented with black arrows.
Microbial abundance (qPCR analysis)
Analysis of variance determined significant effects of seasonality on bacterial and archaeal abundance as measured by qPCR of 16S rRNA genes and functional genes (P<0.001) (Table 3, Figure 2). Seasonality effect was most pronounced for beta-proteobacteria, which showed abundance shifts of 236 and 242% for the control (average 1.14 × 1010 16S rRNA gene copies) and girdled (average 1.18 × 1010) plots, respectively, over the whole experimental period. The smallest seasonality effect was determined for total archaea (average 6.13 × 107) with 63% variation in the control plots over the whole experiment. However, abundance of total archaea measured in the girdled plots (average 1.09 × 108) revealed a seasonality related fluctuation of 112%. The other microbial groups assayed in the control and girdled plots took intermediate positions with seasonal changes ranging between 65% (verrucomicrobia in control plots) and 97% (alpha-proteobacteria in girdled plots). In contrast to seasonality, tree girdling had a minor effect and was only significant for total archaea, alpha-proteobacteria and acidobacteria (P<0.05). For these three groups, significantly higher 16S RNA gene copies were determined in the girdled plots as compared with the control plots (44%, total archaea; 18%, alpha-proteobacteria and acidobacteria). No significant differences were found for beta-proteobacteria (3% higher in girdled plots) and verrucomicrobia (2% higher in girdled plots). Contrastingly, 16S rRNA gene copies tended to be 7% greater in control plots in comparison with girdled plots when analyzing total bacterial abundance (P>0.05). Except for beta-proteobacteria, a clear interaction between both factors could be calculated indicating an interrelated influence of ‘seasonality’ and ‘tree girdling’ on the abundance of bacterial and archaeal communities (P<0.05) (Table 3).
Quantitative PCR data of the six assayed microbial groups (a) total bacteria, (b) total archaea, (c) alpha-proteobacteria, (d) beta-proteobacteria, (e) acidobacteria and (f) verrucomicrobia as determined over a 2-year period starting in May 2006. Presented data (16S rRNA gene copies per gram dry soil) are average values calculated from six individual soil samples per treatment along with standard errors. White columns show the control plots, whereas grey bars represent the tree girdling plots.
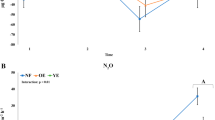
A highly significant effect of seasonality was detected for archaeal and bacterial amoA gene, nirS gene and nosZ gene abundance (P<0.001) (Table 3, Figure 3). All assayed functional genes revealed higher gene copies in the second experimental year (June 2007 to May 2008) as compared with the first investigation period (May 2006 to May 2007). Abundance of bacterial and archaeal amoA genes showed greater seasonal fluctuations as compared with nirS and nosZ genes. Over the whole project period, greatest seasonal changes on archaeal amoA gene abundance were found in control plots (average 1.41 × 107, 187% variation), whereas the effect was smallest for archaeal amoA gene copies in girdled plots (average 3.14 × 107, 145%). Bacterial amoA gene copies took an intermediate position and their seasonal responses were similar to those of archaeal amoA genes. Although seasonality related shifts of the quantities of nirS and nosZ genes were highly significant (P<0.001), their overall seasonal alternation was less intense as compared with the assayed amoA genes. Greatest seasonal differentiations were determined for nosZ gene in the control plots (107%), whereas smallest differences were determined for nirS gene copies in the control plots (91%). Copy numbers of the nirS gene behaved similarly to those of the nosZ gene, but with slightly greater seasonal changes. Distinct differences between copy numbers of assayed functional genes were determined between control and tree girdling plots. Archaeal amoA gene revealed 55% higher numbers in girdled plots (6.83 × 107) as compared with control plots (1.41 × 107) (P<0.01), whereas 71% higher copies of bacterial amoA gene were measured in girdled plots (average 6.83 × 107) as compared with the corresponding controls (average 1.99 × 107) (P<0.001). Effect of tree girdling was not significant for nirS and nosZ genes. Copy numbers of nosZ gene showed 14% lower values in girdled plots as compared with controls (2.04 × 108 versus 2.33 × 108), whereas nirS gene abundance was not significantly changed by tree girdling (2.43 × 108 (controls) versus 2.35 × 108 (tree girdling)) (P>0.05). Significant interactions between seasonality and tree girdling were determined for both amoA genes (P<0.05). Generally, it needs to be pointed out that very small abundance differences may have occurred due to variations in the used qPCR assays. In the latter discussion only those abundance differences were considered for which statistical significances were obtained.
Abundance of the six assayed functional genes for (a) bacterial ammonia monooxygenase (amoA gene), (b) archaeal ammonia monooxygenase (amoA gene), (c) bacterial nitrite reductase (nirS gene) and (d) bacterial nitrous oxide reductase (nosZ gene), as determined by quantitative PCR data over a 2-year period starting in May 2006. The numbers of gene copies are presented per gram dry soil and are average values calculated from six individual soil samples per treatment along with standard errors. White columns show the control plots, whereas grey bars represent the tree girdling plots.
Linear correlation coefficients between microbial abundance and geochemical data
Positive correlations, except for total archaea, verrucomicrobia and functional genes, were calculated for DOC (range from r=0.164 (P<0.05, acidobacteria) to r=0.239 (P<0.01, alpha-proteobacteria)), showing increased gene abundance when high DOC contents were determined. DON revealed a positive correlation with total archaea (r=0.299, P<0.01), acidobacteria (r=0.166, P<0.05) both amoA genes (bacterial amoA gene, r=0.228; archaeal amoA gene, r=0.226) and the nosZ gene (r=0.352) (P<0.01) (Table 6). For soil nitrate, positive correlations were determined between total bacteria (r=0.254, P<0.01), and soil ammonia was positively correlated with beta-proteobacteria (r=0.187, P<0.05) and acidobacteria (r=0.301, P<0.01). Negative correlations were calculated between soil ammonia and total archaea (r= −0.278, P<0.01) as well as verrucomicrobia (r= −0.269, P<0.01) reflecting decreased abundance with increasing soil ammonia levels. Although soil nitrate was only correlated with nosZ gene (r= −0.256), soil ammonia was negatively correlated with all functional genes (at least r= −0.235) (P<0.01). Total N2O emission was positively correlated with total archaeal abundance (r=0.476), acidobacteria (r=0.527), verrucomicrobia (r=0.393), archaeal amoA (r=0.283), nirS (r=0.445) and nosZ (r=0.424) (P<0.01) genes. Soil temperature was negatively correlated with all assayed groups (range from r= −0.163 (P<0.05, acidobacteria) to r= −0.421 (P<0.01, verrucomicrobia)), except with beta-proteobacteria (r= 0.475, P<0.01), indicating an abundance decrease with higher soil temperatures. Similar trends were observed for soil moisture (range from r= −0.233 (P<0.01, total bacteria) to r= −0.398 (P<0.01, beta-proteobacteria)); however, for total archaea and acidobacteria an increase in abundance was determined when higher soil moisture occurred (r=0.327 (P<0.01, total archaea) and r=0.289 (P<0.01, acidobacteria)). Functional genes were negatively correlated with soil temperature (range from r= −0.259 (bacterial amoA gene) to r= −0.456 (nosZ gene)), whereas positive correlations were determined between functional genes and soil moisture (range from r=0.247 (archaeal amoA gene) to r=0.418 (nosZ gene)) (P<0.01).
Discussion
Previous studies on the effects of seasonality and resource availability on dynamics of soil microbial communities under field conditions have been restricted in resolution by the use of wide sampling intervals and short investigation periods, and have focused on broad microbial domains rather than specific phyla. To overcome these limitations, we used a bi-monthly sampling scheme during a 2-year tree girdling field experiment period to study in detail the effect of seasonality and resource availability on the soil microbial community. Structure (T-RFLP analysis) and abundance (qPCR) of microbial communities were analyzed at different taxonomic scales including bacterial and archaeal domains and specific bacterial phyla, which occur prominently in the assayed soil, that is, acidobacteria, alpha- and beta-proteobacteria and verrucomicrobia, as well as nitrifying and denitrifying bacteria and archaea. Our results showed that resource availability due to seasonal variation, but also due to tree girdling resulted in specific short- and medium-term changes in community structure and abundance of archaea and bacteria as well as representatives of selected phyla. Generally, microbial communities were altered by seasonal effects to a larger extent than by altered root exudation. These community alterations were partly ascribed to the influence of seasonality and tree girdling on physicochemical parameters such as DOC) and DON, nitrate, ammonia, as well as soil temperature and soil moisture.
Seasonality in temperate forest soils is reflected by alterations in soil moisture and soil temperature, being acknowledged control factors of soil microbial communities (Stres et al., 2008; Tabuchi et al., 2008; Cleveland et al., 2007). Both parameters were responsible for compositional shifts in soil bacterial communities determined in this study. Similarly, changes in soil temperature and moisture appeared to be determinants of the archaeal community in the present field study. Also Shen et al. (2008) and Tourna et al. (2008) have found temporal shifts in archaeal abundance, and Stres et al. (2008) and Tourna et al. (2008) evidenced responsiveness of soil archaea to variations in soil temperature and soil moisture. Seasonal changes in soil climate were further closely linked to short- and medium-term variations in resource availability, which further correlated with the quantity and quality of organic matter entering the soil, as it was also previously suggested (Bell et al., 2009; Cookson et al., 2006; Krave et al., 2002). Consequently, total bacterial communities and individual phyla studied were clearly shaped by supply of DOC, DON and mineral N (that is, ammonia, nitrate), which is in agreement with previously published data (Drenovsky et al., 2004; Zak et al., 2003; Alden et al., 2001).
Abundance of acidobacteria was positively correlated with soil DOC and mineral N contents, which is in agreement with previous reports (Tabuchi et al., 2008; Hayatsu et al., 2008; Ruppel et al., 2007). Several members of this phylum have been evidenced to be facultative or obligatory anaerobic organotrophs (Jones et al., 2009; Fierer et al., 2007). Hence, their abundance may be particularly favored by high soil moisture together with effects on community structure, as we found in the present field study and was proven by other reports (for example, Janssen, 2006). Throughout the field experiment, acidobacterial communities were more abundant and underwent significant structural changes in girdled, C-limited plots. This may signify their high metabolic versatility to be well-adapted to resource limitation and their ability to decompose complex C substrates deriving from the rather recalcitrant soil organic matter pool (Ward et al., 2009; Hansel et al., 2008; Eichorst et al., 2007; Fierer et al., 2007).
Girdling prevents the uptake of available nutrients such as ammonia and nitrate by trees (Högberg et al., 2001), and therefore resulted in relatively higher mineral N concentrations in soils of girdled plots. We found significantly higher bacterial amoA gene copies in girdled plots than in controls substantiating our assumption that N availability is a crucial controlling factor for ammonia oxidizing bacteria (Fierer et al., 2009). No correlation was seen between ammonia oxidizing bacteria and soil nitrate content. However, net changes in the soil nitrate pool do not reflect nitrifying activity, as apart from microbes plants also utilize nitrate as N source (Adair and Schwartz, 2008). Copies of bacterial amoA genes were further positively correlated with DON. DON is the precursor of ammonia (mineralization) and thus is essential for the constant replenishment of the ammonia pool as substrate for nitrification, thus indicating that DON is essential for maintaining ammonia oxidizing bacteria metabolism (You et al., 2009; Brierley et al., 2001). Moist soil conditions pronouncing the diffusion of substrates (for example, nitrate and ammonia) to microbes offered obviously a favourable environment for sustaining and increasing the abundance of ammonia oxidizing bacteria, which is in agreement with previously published information (Fierer et al., 2009; Adair and Schwartz, 2008).
We found that seasonality and varying resource availability changed the abundance of ammonia oxidizing archaea (AOA). Decreasing soil temperature was correlated with increasing AOA abundance, which is in contrast to the results of a soil microcosm study by Tourna et al. (2008). However, Urakawa et al. (2008) and Caffrey et al. (2007) investigated marine ecosystems in which decreased phylogenetic diversity and abundance of AOA were found with increasing temperature, respectively. Based on these contradictory results, we suggest further experiments under field conditions to substantiate that decreasing soil temperature promotes AOA abundance. Our results suggested a potential dependence of AOA on ammonia availability, which was supported by recent studies (He et al., 2007; Santoro et al., 2008). Because of the negative correlation between AOA and ammonia concentrations, we conclude that the ammonia decrease may have been the consequence of pronounced ammonia oxidation activity in the assayed temperate soils, whereas Valentine, (2007) proposed that AOA seem to be better adapted to low ammonia concentrations in soil. However, it remains poorly investigated to which extent soil AOA react to different concentrations of ammonium in soils (Jia and Conrad, 2009; Chen et al., 2008).
N2O emissions were positively correlated with the abundance of total archaea and AOA indicating that archaea may be directly involved in denitrification processes. Although denitrification is often considered a bacterial process, the measured high abundance of archaea suggested that denitrification was probably also widespread among the archaea studied in this field experiment. But further research is required to substantiate this assumption as only limited information is available for archaeal denitrification in soil ecosystems so far (for example, Bartossek et al., 2010; Hayatsu et al., 2008). However, it has been confirmed that several archaeal members perform both assimilatory and dissimilatory reduction processes to produce for example, N2O (Hayatsu et al., 2008; Cabello et al., 2004; Zehr and Ward, 2002), and their actual contribution to denitrification was proven by the presence of denitrification genes (for example, nir and nos genes) in the genomes of several archaeal species (Bartossek et al., 2010; Cabello et al., 2004).
In conclusion, our field study in a temperate ecosystem including tree girdling to induce soil C limitation is the first field survey that has been performed for two consecutive years with a bi-monthly sampling scheme. This approach allowed us to get a detailed insight into short- and medium-term effects of seasonality and resource availability on the soil microbial community, which has been explored at domain level as well as at a smaller taxonomic scale using selected bacterial phyla and functional groups. We showed that community structure and abundance of archaea and acidobacteria appeared to be particularly altered by these two factors, reflecting their potentially high metabolic versatility in the assayed soils. Further, our extensive field survey revealed that belowground C allocation along with seasonal climatic influences changed the abundance of nitrifying and denitrifying bacteria and archaea and showed a sound correlation with treatment-related dynamics of physicochemical parameters in the investigated soils. Based on our proposed assumptions, it will be essential to promote future research to further explore and understand the role of various phylogenetic and functional groups in terrestrial environments as well as their individual response to various environmental parameters and particularly their resilience to climate change (Cruz-Martínez et al., 2009; Youssef and Elshahed, 2009).
References
Adair KL, Schwartz E . (2008). Evidence that ammonia-oxidizing archaea are more abundant than ammonia-oxidizing bacteria in semiarid soils of northern Arizona, USA. Microbial Ecol 56: 420–426.
Alden L, Demoling F, Bååth E . (2001). Rapid method of determining factors limiting bacterial growth in soil. Appl Environ Microbiol 67: 1830–1838.
Bartossek R, Nicol GW, Lanzen A, Klenk H-P, Schleper C . (2010). Homologues of nitrite reductases in ammonia-ozidizing archaea: diversity and genomic context. FEMS Environ Microbiol 12: 1075–1088.
Barns SM, Takala SL, Kuske CR . (1999). Wide distribution and diversity of members of the bacterial kingdom Acidobacterium in the environment. Appl Environ Microbiol 65: 1731–1737.
Bell CW, Acosta-Martinez V, McIntyre NE, Cox S, Tissue DT, Zak JC . (2009). Linking microbial community structure function to seasonal differences in soil moisture temperature in a Chihuahuan Desert grassland. Microbial Ecol (in press; doi:10.1007/s00248-009-9529-5).
Brierley EDR, Wood M, Shaw PJA . (2001). Nitrogen cycling and proton fluxes in an acid forest soil. Plant Soil 229: 83–96.
Buckley DH, Schmidt TM . (2002). Exploring the biodiversity of soil: a microbial rainforest. Biodiversity of Microbial Life, In: Staley, JT and Reysenbach, AL (eds). Wiley-Liss: New York, NY, pp 183–208.
Cabello P, Roldán MD, Moreno-Vivían C . (2004). Nitrate reduction and the nitrogen cycle in archaea. Microbiology 150: 3527–3546.
Caffrey JM, Bano N, Kalanetra K, Hollibaugh JT . (2007). Ammonia oxidation and ammonia-oxidizing bacteria and archaea from estuaries with differing histories of hypoxia. The ISME J 1: 660–662.
Cannell MGR, Dewar RC . (1994). Carbon allocation in trees: a review of concepts for modelling. Adv Ecol Res 25: 59–104.
Chen XP, Zhu YG, Xia Y, Shen JP, He JZ . (2008). Ammonia-oxidizing archaea: important players in paddy rhizosphere soil? Environ Microbiol 10: 1978–1987.
Clarke KR, Green RH . (1988). Statistical design and analysis for a ‘biological effects’ study. Mar Ecol Prog Ser 46: 213–226.
Cleveland CC, Nemergut DR, Schmidt SK, Townsend AR . (2007). Increases in soil respiration following labile carbon additions linked to rapid shifts in soil microbial community composition. Biogeochemistry 82: 229–240.
Cookson WR, Marschner P, Clark IM, Milton N, Smirk MN, Murphy DV et al. (2006). The influence of season, agricultural management, and soil properties on gross nitrogen transformations and bacterial community structure. Aust J Soil Res 44: 453–465.
Cruz-Martínez K, Suttle KB, Brodie EL, Power ME, Andersen GL, Banfield JF . (2009). Despite strong seasonal responses, soil microbial consortia are more resilient to long-term changes in rainfall than overlying grassland. ISME J 3: 738–744.
Dannenmann M, Simon J, Gasche R, Holst J, Naumann PS, Koegel-Knabner I et al. (2009). Tree girdling provides insight on the role of labile carbon in nitrogen partitioning between soil microorganisms and adult European beech. Soil Biol Biochem 41: 1622–1631.
Drenovsky RE, Vo D, Graham KJ, Scow KM . (2004). Soil water content and organic carbon availability are major determinants of soil microbial community composition. Microbial Ecol 48: 424–430.
Dunbar J, Ticknor LO, Kuske CR . (2000). Assessment of microbial diversity in four Southwestern United States soils by 16S rRNA gene terminal restriction fragment analysis. Appl Environ Microbiol 66: 2943–2950.
Eichorst SA, Breznak JA, Schmidt TM . (2007). Isolation and characterization of soil bacteria that define Terriglobus gen. nov., in the phylum Acidobacteria. Appl Environ Microbiol 73: 2708–2717.
Ekberg A, Buchmann N, Gleixner G . (2007). Rhizospheric influence on soil respiration and decomposition in a temperate Norway spruce stand. Soil Biol Biochem 39: 2103–2110.
Fierer N, Bradford MA, Jackson RB . (2007). Toward an ecological classification of soil bacteria. Ecology 88: 1354–1364.
Fierer N, Carney KM, Horner-Devine MC, Megonigal JP . (2009). The biogeography of ammonia-oxidizing bacterial communities in soil. Microbial Ecol 58: 435–445.
Fontaine S, Bardoux G, Benest D, Verdier B, Mariotti A, Abbadie L . (2004). Mechanisms of the priming effect in a savannah soil amended with cellulose. Soil Sci Soc Am J 68: 125–131.
Francis CA, Roberts KJ, Beman JM, Santoro AE, Oakley BB . (2005). Ubiquity and diversity of ammonia-oxidizing archaea in water columns and sediments of the ocean. P Natl Acad Sci USA 102: 14683–14688.
Grayston SJ, Prescott CE . (2005). Microbial communities in forest floors under four tree species in coastal British Columbia. Soil Biol Biochem 37: 1157–1167.
Hackl E, Zechmeister-Boltenstern S, Bodrossy L, Sessitsch A . (2004). Comparison of diversities and compositions of bacterial populations inhabiting natural forest soils. Appl Environ Microbiol 70: 5057–5065.
Hansel CM, Fendorf S, Jardine PM, Francis CA . (2008). Changes in bacterial and archaeal community structure and functional diversity along a geochemically variable soil profile. Appl Environ Microbiol 74: 1620–1633.
Hayatsu M, Tago K, Saito M . (2008). Various players in the nitrogen cycle: diversity and functions of the microorganisms involved in nitrification and denitrification. Soil Sci Plant Nutr 54: 33–45.
He J, Shen J, Zhang L, Zhu Y, Zheng Y, Xu M et al. (2007). Quantitative analyses of the abundance and composition of ammonia-oxidizing bacteria and ammonia-oxidizing archaea of a Chinese upland red soil under long-term fertilization. Environ Microbiol 9: 2364–2374.
Henderson SL, Dandie CE, Patten CL, Zebarth BJ, Burton DL, Trevors JT et al. (2010). Changes in denitrifier abundance, denitrification gene mRNA levels, nitrous oxide emissions, and denitrification in anoxic soil microcosms amended with glucose and plant pesidues. Appl Environ Microbiol 76: 2155–2164.
Henry S, Bru D, Stres B, Hallet S, Philippot L . (2006). Quantitative detection of the nosZ gene, encoding nitrous oxide reductase, and comparison of the abundances of 16S rRNA, narG, nirK, and nosZ genes in soils. Appl Environ Microbiol 72: 5181–5189.
Högberg P, Nordgren A, Buchmannn N, Taylor AFS, Ekblad A, Högberg MN et al. (2001). Large-scale forest girdling shows that current photosynthesis drives soil respiration. Nature 411: 789–792.
Högberg MN, Chen Y, Högberg P . (2007). Gross nitrogen mineralisation and fungi-to-bacteria ratios are negatively correlated in boreal forests. Biol Fert Soils 44: 363–366.
Horz H-P, Barbook A, Field CB, Bohannan BJM . (2004). Ammonia-oxidizing bacteria respond to multifactorial global change. P Natl Acad Sci USA 101: 15136–15141.
Janssen PH . (2006). Identifying the dominant soil bacterial taxa in libraries of 16S rRNA and 16S rRNA genes. Appl Environ Microbiol 72: 1719–1728.
Jia Z, Conrad R . (2009). Bacteria rather than Archaea dominate microbial ammonia oxidation in an agricultural soil. Environ Microbiol 11: 1658–1671.
Jones R, Robeson MS, Lauber CL, Hamady M, Knight R, Fierer N . (2009). A comprehensive survey of soil acidobacterial diversity using pyrosequencing and clone library analyses. ISME J 3: 442–453.
Kaiser C, Koranda M, Kitzler B, Fuchslueger L, Schnecker J, Schweiger P et al. (2010). Belowground carbon allocation by trees drive seasonal pattern of extracellular enzyme activities by altering microbial community composition in a beech forest soil. New Phytologist 187: 843–858.
Koch O, Tscherko D, Kandeler E . (2007). Temperature sensitivity of microbial respiration, nitrogen mineralization, and potential soil enzyme activities in organic alpine soils. Global Biogeochem Cycles 21: GB4017.
Krave AS, Lin B, Braster M, Laverman AM, van Stralen NM, Roling WF et al. (2002). Stratification and seasonal stability of diverse bacterial communities in a Pinus merkusii (pine) forest soil in central Java, Indonesia. Environ Microbiol 4: 361–373.
Lane D . (1991). 16S/23S rRNA sequencing, In: Stackebrandt, A and Goodfellow, M (eds). Nucleic Acid Techniques Systematics. John Wiley: West Sussex, UK, pp 115–175.
Legendre P, Legendre L . (1998). Numerical Ecology 2nd edn. Elsevier: Amsterdam, The Netherlands.
Leininger S, Urich T, Schloter M, Schwark L, Qi J, Nicol GW et al. (2006). Archaea predominate among ammonia-oxidizing prokaryotes in soils. Nature 442: 806–809.
Lejon DPH, Chaussod R, Ranger J, Ranjard L . (2005). Microbial community structure and density under different tree species in an acid forest soil (Morvan, France). Microbial Ecol 50: 614–625.
Lepš J, Šmilauer P . (2003). Multivariate Analysis of Ecological Data using CANOCO. Cambridge University Press: Oxford, UK, pp 282.
Liu W-T, Marsh TL, Cheng H, Forney LJ . (1997). Characterization of microbial diversity by determining terminal restriction length polymorphisms of genes encoding 16S rRNA. Appl Environ Microbiol 63: 4516–4522.
Lueders T, Friedrich M . (2000). Archaeal population dynamics during sequential reduction processes in rice field soil. Appl Environ Microbiol 66: 2732–2742.
Magill AH, Aber JD . (2000). Dissolved organic carbon and nitrogen relationships in forest litter as affected by nitrogen deposition. Soil Biol Biochem 32: 603–613.
Michotey V, Méjean V, Bonin P . (2000). Comparison of methods for quantification of cytochrome cd1-denitrifying bacteria in environmental marine samples. Applied and Environmental Microbiology 66: 1564–1571.
Muyzer G, Dewaal EC, Uitterlinden AG . (1993). Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59: 695–700.
O′Farrell KA, Janssen PH . (1999). Detection of verrucomicrobia in a pasture soil by PCR-mediated amplification of 16S rRNA genes. Appl Environ Microbiol 65: 4280–4284.
Overmann J, Coolen MJL, Tuschak C . (1999). Specific detection of different phylogenetic groups of chemocline bacteria based on PCR and denaturing gradient gel electrophoresis of 16S rRNA gene fragments. Arch Microbiol 172: 83–94.
Philippot L, Kuffner M, Chèneby D, Depret G, Laguerre G, Martin-Laurent F . (2006). Genetic structure and activity of the nitrate-reducers community in the rhizosphere of different cultivars of maize. Plant Soil 287: 177–186.
Rasche F, Hödl V, Poll C, Kandeler E, Gerzabek MH, van Elsas JD et al. (2006). Rhizosphere bacteria affected by transgenic potatoes with antibacterial activities compared with the effects of soil, wild-type potatoes, vegetation stage and pathogen exposure. FEMS Microbiol Ecol 56: 219–235.
Rotthauwe J-H, Witzel K-P, Liesack W . (1997). The ammonia monooxygenase structural gene amoA as a functional marker: molecular fine-scale analysis of natural ammonia-oxidizing populations. Appl Environ Microbiol 63: 4704–4712.
Ruppel S, Torsvik V, Daae FL, vreås L, Rühlmann J . (2007). Nitrogen availability decreases prokaryotic diversity in sandy soils. Biol Fert Soils 43: 449–459.
Santoro AE, Francis CA, de Sieyes NR, Boehm AB . (2008). Shifts in the relative abundance of ammonia-oxidizing bacteria and archaea across physicochemical gradients in a subterranean estuary. Environ Microbiol 10: 1068–1079.
Schimel JP, Weintraub MN . (2003). The implications of exoenzyme activity on microbial carbon and nitrogen limitation in soil: a theoretical model. Soil Biol Biochem 35: 549–563.
Shen JP, Zhang LM, Zhou YB, Zhang JB, He JZ . (2008). Abundance and composition of ammonia-oxidizing bacteria and ammonia-oxidizing archaea communities of an alkaline sandy loam. Environ Microbiol 10: 1601–1611.
Stres B, Danevèiè T, Pal L, Mrkonjiæ M, Resman L, Leskovec S et al. (2008). Influence of temperature and soil water content on bacterial, archaeal and denitrifying microbial communities in drained fen grassland soil microcosms. FEMS Microbiol Ecol 66: 110–122.
Tabuchi H, Kato K, Nioh I . (2008). Season and soil management affect soil microbial communities estimated using phospholipid fatty acid analysis in a continuous cabbage (Brassica oleracea var. capitata) cropping system. Soil Sci Plant Nutr 54: 369–378.
Throbäck IN, Enwall K, Jarvis A, Hallin S . (2004). Reassessing PCR primers targeting nirS, nirK and nosZ genes for community surveys of denitrifying bacteria with DGGE. FEMS Microbiol Ecol 49: 401–417.
Tourna M, Freitag TE Nicol GW, Prosser JI . (2008). Growth, activity and temperature responses of ammonia-oxidizing archaea and bacteria in soil microcosms. Environ Microbiol 10: 1357–1364.
Urakawa H, Tajima Y, Numata Y, Tsuneda S . (2008). Low temperature decreases the phylogenetic diversity of ammonia-oxidizing archaea and bacteria in aquarium biofiltration systems. Appl Environ Microbiol 74: 894–900.
Valentine DL . (2007). Adaptations to energy stress dictate the ecology and evolution of the archaea. Nat Rev Microbiol 5: 316–323.
Waldrop MP, Firestone MK . (2006). Altered utilization patterns of young and old soil C by microorganisms caused by temperature shifts and N additions. Biogeochemistry 67: 235–248.
Ward NL, Challacombe JF, Janssen PH, Henrissat B, Coutinho PM, Wu M et al. (2009). Three genomes from the phylum Acidobacteria provide insight into the lifestyles of these microorganisms in soils. Appl Environ Microbiol 75: 2046–2056.
Waring RH, Running SW . (1998). Forest ecosystems: analysis at multiple scales, 2nd edn. Academic Press: San Diego, CA.
Weintraub MN, Scott-Denton LE, Schmidt SK, Monson RK . (2007). The effects of tree rhizodeposition on soil exoenzyme activity, dissolved organic carbon, and nutrient availability in a subalpine forest ecosystem. Oecologia 154: 327–338.
Weisburg WG, Barns SM, Pelletier DA, Lane DJ . (1991). 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173: 697–703.
Wolsing M, Priemé A . (2004). Observation of high seasonal variation in community structure of denitrifying bacteria in arable soil receiving artificial fertilizer and cattle manure by determining T-RFLP of nir gene fragments. FEMS Microbiol Ecol 48: 261–271.
Yarwood SA, Myrold DD, Högberg MN . (2009). Termination of belowground C allocation by trees alters soil fungal and bacterial communities in a boreal forest. FEMS Microbiol Ecol 70: 151–162.
Youssef NH, Elshahed MS . (2009). Diversity rankings among bacterial lineages in soil. ISME J 3: 305–313.
You J, Das A, Dolan EM, Hu Z . (2009). Ammonia-oxidizing archaea involved in nitrogen removal. Water Res 43: 1801–1809.
Zak DR, Holmes WE, White DC, Peacock AD, Tilman D . (2003). Plant diversity, soil microbial communities, and ecosystem function: are there any links? Ecology 84: 2042–2050.
Zehr JP, Ward BB . (2002). Nitrogen cycling in the ocean: new perspectives on processes and paradigms. Appl Environ Microbiol 68: 1015–1024.
Acknowledgements
This study was financed by the Austrian Science Fund (FWF, Project number: P18495-B03). We thank Dr. Evelyn Hackl (AIT) for her valuable comments and suggestions on the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Rasche, F., Knapp, D., Kaiser, C. et al. Seasonality and resource availability control bacterial and archaeal communities in soils of a temperate beech forest. ISME J 5, 389–402 (2011). https://doi.org/10.1038/ismej.2010.138
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2010.138
Keywords
This article is cited by
-
Nitrogen Addition Enhances Soil Nitrogen Mineralization Through an Increase in Mineralizable Organic Nitrogen and the Abundance of Functional Genes
Journal of Soil Science and Plant Nutrition (2024)
-
Drivers of organic carbon stocks in eutrophic lake sediments after reestablishment of submerged aquatic vegetation
Plant and Soil (2024)
-
Erosion and deposition significantly affect the microbial diversity, co-occurrence network, and multifunctionality in agricultural soils of Northeast China
Journal of Soils and Sediments (2024)
-
Giant African snail invasion homogenizes seasonal soil biodiversity in tropical coral islands
Plant and Soil (2024)
-
Changes in composition and function of soil microbial communities during secondary succession in oldfields on the Tibetan Plateau
Plant and Soil (2024)