Abstract
Marine cyanobacteria from the genus Synechococcus are found throughout the world's oceans and are important contributors to global primary productivity and carbon cycling. Cultured isolates and environmental DNA clone libraries of Synechococcus have demonstrated the diversity of these microbes. However, the natural distribution of this diversity through space and time and the ecological significance of their distribution are still poorly understood. To understand the seasonal dynamics of Synechococcus diversity, we have developed a quantitative PCR strategy using the gene encoding as a subunit of DNA-dependent RNA polymerase (rpoC1) and applied it to a 3-year time series of surface samples from the Scripps Institution of Oceanography pier (La Jolla, CA, USA), a coastal site in the northeastern Pacific Ocean. Synechococcus from clades I and IV were dominant throughout the time series and correlated with total Synechococcus abundance. The relative abundance of these two dominant clades showed evidence of a seasonal cycle. Synechococcus from clade IV were typically more abundant, but those from clade I dominated during periods just before the annual spring bloom of Synechococcus. Synechococcus from clades II and III were absent during spring and early summer, but appeared at low abundances in late summer and winter possibly due to changes in circulation in the Southern California Bight. As the first long-term time series describing Synechococcus population diversity, these temporal dynamics were used to interpret the genetic/genomic diversity observed in the environment and the potential factors regulating their distribution.
Similar content being viewed by others
Introduction
Marine cyanobacteria from the genus Synechococcus have a fundamental biogeochemical role in the world's oceans. Due to their photosynthetic lifestyle and sheer abundance, they are significant participants in oceanic primary production and the global carbon cycle (Johnson and Sieburth, 1979; Waterbury et al., 1979; Glover, 1985; Li, 1994; Smith and Kemp, 2001; Richardson and Jackson, 2007). As a component of the picophytoplankton, this group accounts for 40% of global ocean primary productivity (Agawin et al., 2000). In some coastal environments, Synechococcus accounts for 20% of the primary productivity (Li, 1994). But to properly understand their role in the environment, we need a better understanding of the ecological niches they occupy, how they adapt to changing environments and how these adaptations are linked to their evolution and diversification.
There is a strong foundation of genetic information describing the diversity of marine Synechococcus. Most marine Synechococcus form a monophyletic group based on phylogenetic trees constructed with 16S rRNA, rpoC1 and other conserved gene sequences and are closely related with the other major marine cyanobacterial genus, Prochlorococcus (Palenik, 1994; Toledo and Palenik, 1997; Rocap et al., 2002; Fuller et al., 2003; Zeidner et al., 2003; Chen et al., 2004; Gibson et al., 2006; Jenkins et al., 2006; Penno et al., 2006).
Within the marine Synechococcus group, distinct clades have been recognized based on rpoC1 gene phylogenies (Palenik, 1994; Ferris and Palenik, 1998). These clades are supported by other gene phylogenies and have been named numerically with approximately 10 clades currently identified (Rocap et al., 2002; Fuller et al., 2003; Ahlgren and Rocap, 2006; Penno et al., 2006). However, as these clades have been delineated based on gene sequence phylogenies, phenotypic traits that define each Synechococcus clade have not been identified except for motility, which is found only in clade III strains (Toledo et al., 1999). In addition, little is known about biogeographical patterns in Synechococcus clades and their causes.
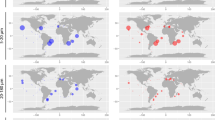
The spatial biogeography of Synechococcus diversity has been broadly described. Synechococcus from clades I, II and IV are the most common throughout the world's oceans (Zwirglmaier et al., 2008). Clades I and IV appear to be the dominant type in temperate, coastal environments (Brown and Fuhrman, 2005; Brown et al., 2005; Zwirglmaier et al., 2007; Zwirglmaier et al., 2008) and clade II is distributed in subtropical/tropical environments (Fuller et al., 2006; Zwirglmaier et al., 2008) and in off-shore, oligotrophic environments (Ferris and Palenik, 1998; Toledo and Palenik, 2003; Ahlgren and Rocap, 2006).
Few studies, however, have examined the temporal changes in Synechococcus diversity and abundance. These few studies have all sampled the Gulf of Aqaba in the Red Sea where Synechococcus from clade II are dominant (Fuller et al., 2005; Muhling et al., 2005, 2006; Penno et al., 2006). The genetic composition of Synechococcus from the Gulf of Aqaba has been shown to change over time, possibly due to changes in nutrient concentrations or viral infections (Muhling et al., 2005) but these studies were of short duration, typically 1 year, and used Southern hybridization, clone library or DNA fingerprinting methods which provided relative quantification only.
In this study, we have developed a quantitative PCR (qPCR) method based on rpoC1 phylogeny to assess the abundance of the three major Synechococcus clades, clades I, II and IV and a fourth clade, clade III. The rpoC1 gene sequence has proven to be a robust genetic marker for describing Synechococcus diversity from several marine environments (Palenik, 1994; Toledo and Palenik, 1997; Ma et al., 2004; Muhling et al., 2006). We have applied this qPCR method to a 3-year time series located at the Scripps Institution of Oceanography (SIO) pier, a coastal Pacific site of the Southern California Bight, and the data were analyzed in relation to physical and chemical parameters from the same site. As the first long-term time series describing Synechococcus population diversity, these temporal dynamics provide essential information to understand the interplay between genetic/genomic diversity, physiology and the environment.
Materials and methods
Sample collection
Surface seawater samples were collected from the end of the SIO pier in La Jolla, CA (32°53′N, 117°15′W) weekly in the morning from January 2005 to January 2008. The sampling site is beyond the surf zone and has a bottom depth of 7 m on average with a spring tidal range of 2 m. From the seawater samples, duplicates of 1 ml seawater were preserved with 0.25% gluteraldehyde (Sigma-Aldrich, St Louis, MO, USA), incubated at room temperature for 15 min, and stored at −80 °C until analyzed by flow cytometry. For DNA analysis, either 500 or 1000 ml of the sample was filtered through a 0.2 μm Supor filter disc (Pall Life Sciences, Ann Arbor, MI, USA) to collect microorganisms and the filter disc was stored at –80 °C.
Sample collection was timed to coincide with chlorophyll, temperature, salinity and nutrient (nitrate (NO3−), ammonia (NH4+), phosphate (PO43−), nitrite (NO2−) and reactive silicate) measurements collected biweekly through the Southern California Coastal Ocean Observing System (SCCOOS) program.
Flow cytometry
Flow cytometry was used to assess the abundance of total Synechococcus cells. When using flow cytometry, Synechococcus cells are distinguished from other picoplankton by the fluorescence of their phycoerythrin pigments. The samples were analyzed at the highest flow rate using a FACSort flow cytometer (Becton-Dickinson, Franklin Lakes, NJ, USA) as described by Collier and Palenik (2003) but without applying a compensation matrix. Duplicates were counted for samples also used for DNA extraction and qPCR analysis.
DNA extraction
DNA was extracted from microorganisms collected on 0.2 μm Supor filter discs (Pall) from monthly to bimonthly samples. The filter discs were first incubated in 12.5 mg/ml lysozyme (Roche Applied Science, Indianapolis, IN, USA), 20 mM EDTA (Sigma), and 50 mM Tris (Sigma), pH 8.0 for 30 min at 37 °C. SDS (Bio-Rad, Hercules, CA, USA) and proteinase K (Sigma) were added to final concentrations of 1% and 1 mg ml−1, respectively, and incubated for 2.5 h at 55 °C. RNA was removed by adding 2 mg ml−1 RNAse (Sigma) for 5 min at room temperature. Proteins were removed by extracting twice with an equal volume of 25:24:1 phenol/chloroform/isoamyl alcohol (Invitrogen, Carlsbad, CA, USA) and once with 24:1 chloroform/isoamyl alcohol (Fisher Scientific, Pittsburg, PA, USA). Using phenol/chloroform dissolved the Supor filter and increased the DNA extraction efficiency. The DNA in the aqueous phase was purified using a silica gel column (DNeasy Blood and Tissue Kit; Qiagen, Valencia, CA, USA) following the manufacturer's protocol and eluted from the column using 400 μl of the manufacturer's buffer. DNA was quantified using PicoGreen (Quant-iT; Qiagen), a DNA-binding fluorescent dye, following the manufacturer's protocol.
Cultured representatives of Synechococcus clades (CC9311 for clade I, CC9605 for clade II, WH8102 for clade III and CC9902 for clade IV) were counted, filtered and their genomic DNA extracted following the same procedure as for environmental samples. On average, 1010 cells were filtered for DNA extraction in a final volume of 400 μl. The DNA extractions from cultured strains were used to test the specificity of clade-specific qPCR primers and as concentration standards.
qPCR
For each clade, two oligonucleotide primers were designed to amplify 145–304 bp fragments of the rpoC1 gene (Table 1). rpoC1 is a single copy gene and thus the problems when using the 16S rRNA gene as the marker locus of normalizing cell number with multiple 16S rRNA gene copies were avoided.
The primer sequences are unique for a given clade based on an alignment of all available Synechococcus rpoC1 gene sequences in the nonredundant (nr) GenBank database from both environmental and cultured sources. A partial alignment of primer regions is provided in Supplementary Table S1. Using DNA from cultured representatives (see above), the primers were tested for clade specificity using a range of annealing temperatures in PCR reactions. An appropriate annealing temperature was chosen (Table 1) so that DNA only from the targeted clade amplified.
qPCR reactions consisted of 1X Brilliant SYBR Green QPCR Master Mix (Stratagene, La Jolla, CA, USA), 30 nM Rox reference dye (Stratagene), 0.8 μM of each primer (Integrated DNA Technologies, Coralville, IA, USA), 0.1 mg/ml BSA (New England BioLabs, Ipswich, MA, USA), and template DNA. Ten-fold serial dilutions of environmental DNA (generally ranging from 5 ng to 0.05 pg DNA) were used as templates in the qPCR reactions. The serial dilutions served as replicates and to dilute any PCR inhibitors occurring in the DNA extractions. DNAs from cultured strains of Synechococcus were serially diluted over 6 orders of magnitude for use as concentration standards. Using a Stratagene 3000P machine to acquire SYBR Green fluorescence data, the reactions were incubated at 95 °C for 10 min followed by 40 cycles of 95 °C for 30 s, the clade-specific annealing temperature for 30 s and 72 °C for 30 s, and ended with 95 °C for 1 min, a ramp down to 55 °C, and a ramp up to 95 °C at 0.2 °C s−1 to assess the dissociation curve of the amplified product.
Template DNAs for the standard curves were obtained from cultured cells that were counted before the DNA extraction, thus the amount of DNA added to each standard qPCR reaction represents the DNA from a known number of cells taking into account any inefficiencies in the DNA extraction. For environmental samples, the threshold cycle (Ct) determined from each qPCR was converted to the number of cells based on the standard curve. The number of cells in each qPCR was converted to cells per ml based on the volume of seawater filtered for each sample and the volume of the DNA extraction used in the qPCRs (no. of cells in the qPCR/(volume of seawater filtered/(volume of DNA extracted × volume of DNA in qPCR)).
rpoC1 clone library
Two cyanobacteria-specific primers (SAN-157F=5′ YTN AAR CCN GAR ATG GAY GG and SAC-1039R=5′ CYT GYT TNC CYT CDA TDA TRT C, the numbers in the primer names denote the first nucleotide where the primers are located along the rpoC1 gene) were designed (by J Collier) to amplify an 841 bp fragment of the rpoC1 gene. These primers generate a longer fragment of the rpoC1 gene and are a better match for marine cyanobacteria than previously published primer pairs (Palenik and Haselkorn, 1992; Muhling et al., 2006). Using the DNA extracted from a sample collected from the SIO pier on 12 May 2005, PCR was used to amplify the rpoC1 gene fragment from the cyanobacterial population. Template DNA (20 ng) was combined with 1.33 μM of each primer and 25 μl of Platinum PCR SuperMix (Invitrogen). This mixture was incubated at 94 °C for 2 min, followed by 30 cycles at 94 °C for 30 s, 52 °C for 1 min and 72 °C for 1 min, and ending with an incubation at 72 °C for 10 min. The resulting PCR product was ligated into pCR4-TOPO plasmid vectors and cloned via transformation into TOP10 chemically competent Escherichia coli cells following the manufacturer's protocol (Invitrogen). Plasmid DNA was prepared from 21 clones using the QIAprep Spin Miniprep Kit (Qiagen) following the manufacturer's protocol. The rpoC1 gene fragment insertion was sequenced (SeqXcel, San Diego, CA, USA) from both ends using the T3 and T7 sequencing primers provided by the manufacturer (Invitrogen).
The rpoC1 gene fragment sequences were trimmed of plasmid and primer sequences and deposited in GenBank (accession numbers FJ790799–FJ790819). The sequences were aligned to known rpoC1 gene sequences from cultured representatives of marine cyanobacteria, that is, Synechococcus and Prochlorococcus, available through GenBank. A phylogenetic tree was calculated using the neighbor-joining algorithm in PAUP (v.4.0b10) with distances calculated using the hky85 model. A 50% majority-rule consensus tree calculated from 1000 bootstrap replicates provided statistical support.
Correlation analyses
Weekly total Synechococcus abundances, temperature, chlorophyll, salinity and nutrient measurements were subjected to pairwise correlation analyses. A separate dataset of bimonthly to monthly clade-specific abundances, total Synechococcus abundances, temperature, chlorophyll, salinity and nutrient concentrations was also analyzed. Except for temperature, the data were log transformed (log (x+1)) before analysis to obtain an approximately normal distribution of data values. The Pearson's product-moment correlation coefficient (r) and the nonparametric Spearman's correlation coefficient (ρ) were calculated using JMP v. 5.0.
Results
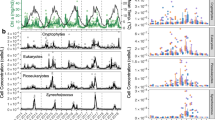
From weekly samples over 3 years, total Synechococcus abundances were determined using flow cytometry. By examining the total abundances, a clear annual pattern emerged (Figure 1). Consistently in mid-May, Synechococcus abundance increased above basal fluctuations indicating that Synechococcus blooms on an annual basis. The timing of the blooms is very consistent, with the annual abundance peaks occurring on 12 May 2005 (6.06 × 105 cells ml−1), 18 May 2006 (2.27 × 105 cells ml−1) and 17 May 2007 (4.84 × 105 cells ml−1). Second abundance peaks occurred in 2006 at the end of June (29 June 2006, 2.73 × 105 cells ml−1) and in 2007 at the end of March (26 March 2007, 3.95 × 105 cells ml−1), but, in general, Synechococcus abundances remained below 2 × 105 cells ml−1 throughout the rest of the year. Abundance minimums occurred throughout the year; the lowest in the time-series was 319 cells ml−1 on 23 March 2006. Total Synechococcus abundances weakly correlated with temperature (Pearson's r=0.30, P=0.0001, Spearman's ρ=0.28, P=0.0002) and negatively with phosphate (r=−0.31, P=0.005, ρ=0.22, P=0.016), but did not correlate significantly with salinity, chlorophyll or any of the other measured nutrients. The lack of correlation with chlorophyll concentration is not surprising as larger phytoplankton, such as diatoms and dinoflagellates, can dominate more than 70% of the chlorophyll (Melissa Carter, personal communication).
Primers were developed for qPCR to assess the genetic composition of the Synechococcus population during this time series. The primers were tested for specificity against genomic DNA from cultured representatives. At the annealing temperature used, the primers were clade specific and did not amplify DNA from any nontarget strain (Figure 2; Supplementary Figure S1). All primers quantified DNA over 6 orders of magnitude equivalent to 102 to 107 cells. Based on the amount of seawater filtered for the DNA extraction, the detection limit for this method is approximately 10 cells ml−1.
Range and specificity of quantitative PCR (qPCR) reactions using dilutions of genomic DNA from cultured Synechococcus strains: (a) clade I qPCR primers targeting CC9311; (b) clade II qPCR primers targeting CC9605; (c) clade III qPCR primers targeting WH8102; (d) clade IV qPCR primers targeting CC9902. The threshold cycle is the PCR cycle at which the fluorescence signal begins to increase. The number of cells in each qPCR reaction, rather than the amount of DNA, is plotted on the x axis. Closed symbols indicate qPCR reactions using DNA from Synechococcus strains targeted by the clade-specific qPCR primers. Open symbols indicate qPCR reactions where DNAs from nontarget strains were assayed.
From bimonthly to monthly samples, qPCR was used to evaluate the diversity of the Synechococcus population from the SIO pier. Synechococcus from clades I and IV consistently dominated this coastal environment throughout the year (Figure 3a and b). Both clade I and clade IV abundances significantly correlated with total Synechococcus concentrations (Figure 4a and b; Supplementary Table S2), and the abundance maxima of these clades coincided with the annual blooms in total Synechococcus (Figure 3a and b). The clade I and clade IV abundances were also strongly correlated to one another (Figure 4c; Supplementary Table S2).
Abundances of the two dominant clades over 3 years from Scripps Institution of Oceanography (SIO) pier surface samples: (a) clade I abundances; (b) clade IV abundances; (c) clade I+clade IV abundances; (d) log2 fold change of clade I to clade IV abundances. Positive values indicate a greater abundance of clade I in the environment. Negative values indicate a greater abundance of clade IV. Zero (0) indicates that each clade was in equal abundance (dashed gray line). In each panel, the dotted gray line indicates total Synechococcus abundances, measured weekly. Clade-specific abundances were measured monthly.
Scatter plots of clade-specific and total Synechococcus abundances from samples assayed by quantitative PCR (qPCR) over the 3-year time series: (a) clade I versus total Synechococcus abundances; (b) clade IV versus total Synechococcus abundances; (c) clade I versus clade IV abundances; (d) clade II versus clade III abundances. In each panel, samples from peaks in Synechococcus abundance are represented by the symbols: + (12 May 2005), × (14 May 2006), ^ (26 March 2007) and * (17 May 2007). In (c), gray symbols represent samples just before the Synechococcus bloom where clade I abundances exceeds clade IV. Results from correlation analyses (Pearson's correlation coefficient (r) and Spearman's correlation coefficient (ρ) and its associated P value) are provided. The best linear fit is shown as a light gray line.
These two clades accounted for an average of 168±54%, of the total Synechococcus abundance (Figure 3c). The values ranged from 83% to 335% with the exception of 12 May 2005 where the two clades accounted for only 34% of the total abundance (see below). The qPCR assay may have overestimated the clade-specific abundances because dividing cells with replicated chromosomes would be counted as single cells by the flow cytometer. This suggests that cells in the environment typically contained replicated chromosomes, but not the cultured cells used for qPCR standards.
The ratio of clade I to clade IV abundances was used to assess changes in the abundance of these two dominant clades relative to one another. For most of the year, clade IV Synechococcus were at least twice as abundant as clade I (log2 fold change <−1.0; Figure 3d). However, each year just before the annual peak in total Synechococcus, clade I increased to twice the abundance of clade IV (log2 fold change >1.0; Figure 3d).
Synechococcus from clades II and III rarely occurred in this environment or occurred in low abundances (Figure 5). Clade II was typically more abundant than clade III, but both clades appeared in the winter when the water is colder and more oligotrophic, and also during the summer when the water is more likely to be stratified. The abundance of these clades was not correlated with total Synechococcus (Supplementary Table S2), but clade II abundances were significantly correlated with clade III abundances suggesting their dynamics may be influenced by similar mechanisms (Figure 4d).
Abundances of the two minor clades over 3 years from Scripps Institution of Oceanography (SIO) pier surface samples: (a) clade II abundances; (b) clade III abundances. In each panel, the dotted gray line indicates total Synechococcus abundances, measured weekly. Clade-specific abundances were measured monthly.
An rpoC1 clone library was generated to determine the composition of the Synechococcus population on 12 May 2005. On this date, a bloom of total Synechococcus was sampled, but qPCR failed to indicate which were the dominant clade types. This sample is clearly an outlier based on the plots of clade I and clade IV abundances versus total Synechococcus (Figures 4a and b). However, phylogenetic analyses of the sequences from the clone library indicated that the majority of the Synechococcus from this sample belonged to clades I and IV (Figure 6).
Neighbor-joining tree calculated from an alignment of rpoC1 sequences from a Scripps Institution of Oceanography (SIO) pier sample on 12 May 2005 and cultured isolates. Sequences from the SIO pier sample are in bold. Values at nodes indicate percentage bootstrap support based on 1000 replicates. Values less than 70% are not shown. The scale bar indicates 10% divergence.
On examining the rpoC1 sequences from the library, the clade I and clade IV qPCR primers do match the respective rpoC1 sequences from this clone library, so it is not known why the qPCR assay failed to recognize the dominance of these two clades on this date. Chemical inhibitors coextracted with the environmental DNA may have inhibited the qPCR reaction. However, dilution of the environmental DNA should have avoided this problem.
Seventeen and three environmental sequences grouped with strong support to cultured clade I and clade IV strains, respectively. For clade I, the environmental sequences grouped as two separate, but strongly supported, clades distinct from cultured strains. This suggests that the genetic diversity of the Synechococcus population in the environment can differ significantly from that of isolated strains as also seen recently in a metagenomic study at our monitoring site (Palenik et al., 2009).
Discussion
We have developed a qPCR strategy to quantify Synechococcus clades, as defined by rpoC1, 16S rRNA and 16S–23S internal transcribed spacer sequences (Ferris and Palenik, 1998; Rocap et al., 2002; Fuller et al., 2003). This approach enabled us to document the changes in Synechococcus diversity from a 3-year time series of a coastal Pacific Ocean site. From this time series, it is clear that the Synechococcus population was dominated by clades I and IV throughout the year (Figure 3a and b). Secondly, the abundances of these two dominant clades were correlated with the total Synechococcus abundance, which showed annual abundance peaks in the month of May (Figures 4a and b). A third intriguing result was that the abundances of Synechococcus from clade I relative to those from clade IV did not remain static, but changed throughout the year with evidence of a seasonal cycle. In particular, although Synechococcus from clade IV were generally the dominant type, Synechococcus from clade I dominated over clade IV just before the annual abundance peaks of total Synechococcus (Figure 3d).
Seasonal pattern of total Synechococcus abundance
The annual pattern of total Synechococcus abundance from this coastal Pacific Ocean site differs from that of a temperate coastal Atlantic site (Woods Hole Harbor) and an oligotrophic site (Sargasso Sea). At the SIO pier, abundance maxima above 2 × 105 cells ml−1 last approximately 1 week and generally occur as the temperature rises above 17 °C. In contrast, from Woods Hole Harbor, Synechococcus blooms when the water reaches 6 °C and the bloom period lasts months, from April until June, with high abundances of Synechococcus (105 per ml) observed until November (Waterbury et al., 1986). In the Sargasso Sea, Synechococcus peaks in March/April with the bloom period lasting several months (Durand et al., 2001).
A rise beyond a threshold temperature appears to be a consistent factor that coincides with the timing of bloom initiation in both coastal Pacific and Atlantic oceans. Higher temperature likely causes an increased growth rate of Synechococcus resulting in greater abundances (Agawin et al., 1998). The duration of the blooms, however, is probably more dependent on other environmental factors such as grazing pressure or nutrient availability. At the SIO pier, the relatively short bloom period suggests that mortality rates (for example, through grazing or viral lysis) also increased causing a rapid decrease in Synechococcus abundance.
Codominance of Synechococcus from clades I and IV
Using qPCR, we have shown that Synechococcus from both clades I and IV dominate the SIO pier coastal environment, yet they have different temporal dynamics. The codominance of the Synechococcus population by clades I and IV has been documented in other coastal environments of the Atlantic Ocean, Mediterranean Sea, eastern South Pacific Ocean, California coast and Arctic Ocean and their dominance is thought to be restricted to higher latitudes above 30°N and below 30°S (Brown and Fuhrman, 2005; Brown et al., 2005; Zwirglmaier et al., 2007, 2008). In the Eastern South Pacific and the Atlantic, clade IV is more abundant than clade I, but in the Arctic they occur equally (Zwirglmaier et al., 2008). However, these biogeographic surveys may have missed the temporal aspects of Synechococcus diversity.
From our 3-year time series, we show that the dominance of one clade over the other is dynamic over time and exhibits seasonal patterns. In particular, Synechococcus from clade I exceeds those from clade IV just before the annual abundance peak suggesting that the environment regularly favors clade I (or disfavors clade IV) to the disadvantage of clade IV. However, the relative importance of genetic compared to environmental factors driving these clade-specific changes in Synechococcus abundance is poorly understood, but it is likely that both play significant roles.
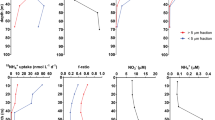
A matching temporal dataset from the SIO pier of temperature, salinity, and nitrate, phosphate, silicate and ammonia concentrations was analyzed to examine potential bottom-up environmental effects driving Synechococcus clade dynamics. Unfortunately none of these significantly correlated with clade-specific abundances (Supplementary Table S2). However, this does not mean that bottom-up effects are insignificant factors in dictating Synechococcus abundance or diversity. Other nutrients that have not been measured could have a significant influence on Synechococcus dynamics. Synechococcus have been shown to use urea as a nitrogen source (Collier et al., 1999) and other organic nutrients are likely important to their survival (Moore et al., 2002, 2005; Zubkov et al., 2003; Bronk et al., 2007). Trace metals may also play a significant role (Hutchins et al., 1999; Leao et al., 2007; Dupont et al., 2008).
The top-down environmental effects on Synechococcus populations are also not well understood. A wide diversity of both protozoan grazers (Caron et al., 1991; Strom, 1991; Christaki et al., 1999, 2002; Guillou et al., 2001; Jeong et al., 2005) and viruses (Suttle and Chan, 1993; Waterbury and Valois, 1993; Wilson et al., 1993) have been identified that prey on Synechococcus. These mortality agents can be the dominating factor regulating total Synechococcus abundances (see for example, Baudoux et al., 2007), but their overall diversity, specificity to Synechococcus types and impact as selective agents remain largely unknown.
Genetic information in association with biogeographic data provides another basis for understanding the factors driving the dominance of clades I and IV in coastal environments and their coexistence. Genome sequences from cultured representatives of clades I and IV are available (Palenik et al., 2006; Dufresne et al., 2008) as are sequences from representatives of several other clades (Palenik et al., 2003) providing a strong backbone of genetic information for this microbial taxon.
CC9311, a representative of clade I isolated from the California Current, has a genome with increased sensory and response regulator genes, fewer phosphate scavenging genes, a greater capacity for metal transport and usage and more genes related to organic nutrient use than an open ocean strain (Palenik et al., 2006). These genes may be key to their survival in a coastal location with higher nutrient availability and a more variable, dynamic environment compared to an oligotrophic one.
The genomes of two clade IV isolates, CC9902 and BL107, have also been sequenced. These strains have fewer sensory histidine kinase and response regulator genes than CC9311 and could represent a ‘specialist’ lifestyle (Dufresne et al., 2008). The genomes of CC9902 and BL107 are approximately 0.38 and 0.33 Mb smaller than CC9311, respectively, and have fewer clade-specific genes. With a larger genetic repertoire, clade I types may be more versatile or ‘opportunists’ (Dufresne et al., 2008). For example, CC9311 has genes capable of transporting iron (II) or halogenating compounds via a vanadium-dependent bromoperoxidase (Palenik et al., 2006), but without more specific physiological investigations, the correlations between genetic composition and niche adaptation remain hypothetical.
The correlations between genetic composition and ecological niche are further complicated by the tremendous genetic diversity that exists within Synechococcus clades. By examining the diversity captured by our rpoC1 clone library, Synechococcus that were dominant in the environment grouped separately from cultured representatives (Figure 6). In addition, as with other microbial taxa (Medini et al., 2005; Tettelin et al., 2005; Kettler et al., 2007), Synechococcus genomes contain mostly conserved core genes, but many highly variable regions exist containing genes with a diversity of functions, likely the result of horizontal gene transfer events, that may be significant to their physiology and ecological niche (Palenik et al., 2009).
Occasional occurrence of Synechococcus from clades II and III
For the two minor clades, clades II and III, their presence in this coastal environment may be more reflective of physical oceanographic patterns. Their occurrence in late summer coincides with periods of strong stratification (Palacios et al., 2004). Stratification has the effect of reducing the input of colder, more saline and nutrient-rich waters from below, therefore surface waters become nutrient depleted. In the fall and winter, stratification is weaker, but there is a consistent northward flow, the Inshore Countercurrent, in the coastal regions of the Southern California Bight in contrast to the southward flow that dominates all other times of the year (Hickey, 1979; Lynn and Simpson, 1987; Di Lorenzo, 2003). The source of the northward flowing waters is the off-shore eastern boundary current of the North Pacific gyre, the California Current, that has turned toward the shore south of the US-Mexico border, then divides and turns north into the Southern California Bight as the Inshore Countercurrent and south alongside Baja California (Hickey, 1979; Lynn and Simpson, 1987). This northward flow brings more oligotrophic, nutrient-poor waters to the coast and may advect Synechococcus from clades II and III that are relatively more abundant off-shore toward the coast (Ferris and Palenik, 1998; Toledo and Palenik, 2003). Although our sampling site is on the coastal shelf, the site is influenced by these major circulation trends, as well as by alongshore currents, cross-shelf tidal currents, and currents generated by local wind stress (Winant and Bratkovich, 1981).
Both stratification and the northward Inshore Countercurrent may have the effect of lowering nutrient concentrations of surface waters in the Southern California Bight and these conditions may explain the occurrence of Synechococcus from clades II and III at the SIO pier. However, we were unable to find a significant relationship between nutrient concentrations and the abundance of these clades.
An alternative explanation may be the advection of Synechococcus from clade II from the south. If clade II is predominantly distributed in subtropical/tropical coastal environments (Zwirglmaier et al., 2008), then northward flow may bring these Synechococcus types to Southern California from the south. Observing a transition from clade I and clade IV dominance in temperate environments to clade II closer to the equator would support this hypothesis. A similar shift in Synechococcus clade dominance has been observed in the Atlantic Ocean (Zwirglmaier et al., 2007).
The observation of Synechococcus from clades II and III in late summer and winter loosely coincides with the presence of Prochlorococcus at this site. Prochlorococcus is usually absent at the SIO pier during the spring months, although in addition to advection to and away from this location, their absence may also be due to increased grazing pressure (Worden et al., 2004).
Conclusion
The biogeography of environmental microbes is a fundamental piece of information that is critical to understanding their evolution, adaptability and function in their respective ecosystems (Ramette and Tiedje, 2007). In addition, as Synechococcus are major contributors to primary production, their biogeography and the genetic and environmental factors influencing the composition of the population has important implications for understanding carbon cycling and other globally important biogeochemical processes. Through the use of qPCR, we have provided an important tool for studying Synechococcus biogeography, ecology and evolution. Although we have used qPCR to assess the temporal dynamics of four globally distributed Synechococcus clades, this method can be applied to any scale of spatial and temporal sampling. With these tools in hand, the key is to discover the scales at which adaptation occurs and how genetic diversity influences and informs us of their ecological niches.
Accession codes
References
Agawin NSR, Duarte CM, Agusti S . (1998). Growth and abundance of Synechococcus sp. in a Mediterranean Bay: seasonality and relationship with temperature. Mar Ecol Prog Ser 170: 45–53.
Agawin NSR, Duarte CM, Agusti S . (2000). Nutrient and temperature control of the contribution of picoplankton to phytoplankton biomass and production. Limnol Oceanogr 45: 591–600.
Ahlgren NA, Rocap G . (2006). Culture isolation and culture-independent clone libraries reveal new marine Synechococcus ecotypes with distinctive light and N physiologies. Appl Environ Microbiol 72: 7193–7204.
Baudoux AC, Veldhuis MJW, Witte HJ, Brussaard CPD . (2007). Viruses as mortality agents of picophytoplankton in the deep chlorophyll maximum layer during IRONAGES III. Limnol Oceanogr 52: 2519–2529.
Bronk DA, See JH, Bradley P, Killberg L . (2007). DON as a source of bioavailable nitrogen for phytoplankton. Biogeosciences 4: 283–296.
Brown MV, Fuhrman JA . (2005). Marine bacterial microdiversity as revealed by internal transcribed spacer analysis. Aquat Microb Ecol 41: 15–23.
Brown MV, Schwalbach MS, Hewson I, Fuhrman JA . (2005). Coupling 16S-ITS rDNA clone libraries and automated ribosomal intergenic spacer analysis to show marine microbial diversity: development and application to a time series. Environ Microbiol 7: 1466–1479.
Caron DA, Lim EL, Miceli G, Waterbury JB, Valois FW . (1991). Grazing and utilization of chroococcoid cyanobacteria and heterotrophic bacteria by protozoa in laboratory cultures and a coastal plankton community. Mar Ecol Prog Ser 76: 205–217.
Chen F, Wang K, Kan JJ, Bachoon DS, Lu JR, Lau S et al. (2004). Phylogenetic diversity of Synechococcus in the Chesapeake Bay revealed by ribulose-1,5-bisphosphate carboxylase-oxygenase (RuBisCO) large subunit gene (rbcL) sequences. Aquat Microb Ecol 36: 153–164.
Christaki U, Courties C, Karayanni H, Giannakourou A, Maravelias C, Kormas KA et al. (2002). Dynamic characteristics of Prochlorococcus and Synechococcus consumption by bacterivorous nanoflagellates. Microb Ecol 43: 341–352.
Christaki U, Jacquet S, Dolan JR, Vaulot D, Rassoulzadegan F . (1999). Growth and grazing on Prochlorococcus and Synechococcus by two marine ciliates. Limnol Oceanogr 44: 52–61.
Collier JL, Brahamsha B, Palenik B . (1999). The marine cyanobacterium Synechococcus sp. WH7805 requires urease (urea amidohydrolase, EC 3.5.1.5) to utilize urea as a nitrogen source: molecular-genetic and biochemical analysis of the enzyme. Microbiology 145: 447–459.
Collier JL, Palenik B . (2003). Phycoerythrin-containing picoplankton in the Southern California Bight. Deep Sea Res Part II 50: 2405–2422.
Di Lorenzo E . (2003). Seasonal dynamics of the surface circulation in the Southern California Current System. Deep Sea Res Part II 50: 2371–2388.
Dufresne A, Ostrowski M, Scanlan DJ, Garczarek L, Mazard S, Palenik BP et al. (2008). Unraveling the genomic mosaic of a ubiquitous genus of marine cyanobacteria. Genome Biol 9: R90.
Dupont CL, Barbeau K, Palenik B . (2008). Ni uptake and limitation in marine Synechococcus strains. Appl Environ Microbiol 74: 23–31.
Durand MD, Olson RJ, Chisholm SW . (2001). Phytoplankton population dynamics at the Bermuda Atlantic Time-series station in the Sargasso Sea. Deep Sea Res Part II 48: 1983–2003.
Ferris MJ, Palenik B . (1998). Niche adaptation in ocean cyanobacteria. Nature 396: 226–228.
Fuller NJ, Marie D, Partensky F, Vaulot D, Post AF, Scanlan DJ . (2003). Clade-specific 16S ribosomal DNA oligonucleotides reveal the predominance of a single marine Synechococcus clade throughout a stratified water column in the Red Sea. Appl Environ Microbiol 69: 2430–2443.
Fuller NJ, Tarran GA, Yallop M, Orcutt KM, Scanlan DJ . (2006). Molecular analysis of picocyanobacterial community structure along an Arabian Sea transect reveals distinct spatial separation of lineages. Limnol Oceanogr 51: 2515–2526.
Fuller NJ, West NJ, Marie D, Yallop M, Rivlin T, Post AF et al. (2005). Dynamics of community structure and phosphate status of picocyanobacterial populations in the Gulf of Aqaba, Red Sea. Limnol Oceanogr 50: 363–375.
Gibson AH, Jenkins BD, Wilkerson FP, Short SM, Zehr JP . (2006). Characterization of cyanobacterial glnA gene diversity and gene expression in marine environments. FEMS Microbiol Ecol 55: 391–402.
Glover HE . (1985). The physiology and ecology of the marine cyanobacterial genus Synechococcus. Adv Aquat Microbiol 3: 49–107.
Guillou L, Jacquet S, Chretiennot-Dinet MJ, Vaulot D . (2001). Grazing impact of two small heterotrophic flagellates on Prochlorococcus and Synechococcus. Aquat Microb Ecol 26: 201–207.
Hickey BM . (1979). The California Current System– hypotheses and facts. Prog Oceanogr 8: 191–279.
Hutchins DA, Witter AE, Butler A, Luther GW . (1999). Competition among marine phytoplankton for different chelated iron species. Nature 400: 858–861.
Jenkins BD, Zehr JP, Gibson A, Campbell L . (2006). Cyanobacterial assimilatory nitrate reductase gene diversity in coastal and oligotrophic marine environments. Environ Microbiol 8: 2083–2095.
Jeong HJ, Park JY, Nho JH, Park MO, Ha JH, Seong KA et al. (2005). Feeding by red-tide dinoflagellates on the cyanobacterium Synechococcus. Aquat Microb Ecol 41: 131–143.
Johnson PW, Sieburth JM . (1979). Chroococcoid cyanobacteria in the sea: a ubiquitous and diverse phototropic biomass. Limnol Oceanogr 24: 928–935.
Kettler GC, Martiny AC, Huang K, Zucker J, Coleman ML, Rodrigue S et al. (2007). Patterns and implications of gene gain and loss in the evolution of Prochlorococcus. PLoS Genet 3: 2515–2528.
Leao PN, Vasconcelos MTSD, Vasconcelos VM . (2007). Role of marine cyanobacteria in trace metal bioavailability in seawater. Microb Ecol 53: 104–109.
Li WKW . (1994). Primary production of prochlorophytes, cyanobacteria, and eukaryotic ultraphytoplankton: measurements from flow cytometric sorting. Limnol Oceanogr 39: 169–175.
Lynn RJ, Simpson JJ . (1987). The California Current System—the seasonal variability of its physical characteristics. J Geophys Res 92: 12947–12966.
Ma Y, Jiao NZ, Zeng YH . (2004). Natural community structure of cyanobacteria in the South China Sea as revealed by rpoC1 gene sequence analysis. Lett Appl Microbiol 39: 353–358.
Medini D, Donati C, Tettelin H, Masignani V, Rappuoli R . (2005). The microbial pan-genome. Curr Opin Genet Dev 15: 589–594.
Moore LR, Ostrowski M, Scanlan DJ, Feren K, Sweetsir T . (2005). Ecotypic variation in phosphorus acquisition mechanisms within marine picocyanobacteria. Aquat Microb Ecol 39: 257–269.
Moore LR, Post AF, Rocap G, Chisholm SW . (2002). Utilization of different nitrogen sources by the marine cyanobacteria Prochlorococcus and Synechococcus. Limnol Oceanogr 47: 989–996.
Muhling M, Fuller NJ, Millard A, Somerfield PJ, Marie D, Wilson WH et al. (2005). Genetic diversity of marine Synechococcus and co-occurring cyanophage communities: evidence for viral control of phytoplankton. Environ Microbiol 7: 499–508.
Muhling M, Fuller NJ, Somerfield PJ, Post AF, Wilson WH, Scanlan DJ et al. (2006). High resolution genetic diversity studies of marine Synechococcus isolates using rpoC1-based restriction fragment length polymorphism. Aquat Microb Ecol 45: 263–275.
Palacios DM, Bograd SJ, Mendelssohn R, Schwing FB . (2004). Long-term and seasonal trends in stratification in the California Current, 1950-1993. J Geophys Res 109: C10016.
Palenik B . (1994). Cyanobacterial community structure as seen from RNA polymerase gene sequence analysis. Appl Environ Microbiol 60: 3212–3219.
Palenik B, Brahamsha B, Larimer FW, Land M, Hauser L, Chain P et al. (2003). The genome of a motile marine Synechococcus. Nature 424: 1037–1042.
Palenik B, Haselkorn R . (1992). Multiple evolutionary origins of prochlorophytes, the chlorophyll B-containing prokaryotes. Nature 355: 265–267.
Palenik B, Ren QH, Dupont CL, Myers GS, Heidelberg JF, Badger JH et al. (2006). Genome sequence of Synechococcus CC9311: insights into adaptation to a coastal environment. Proc Natl Acad Sci USA 103: 13555–13559.
Palenik B, Ren Q, Tai V, Paulsen IT . (2009). Coastal Synechococcus metagenome reveals major roles for horizontal gene transfer and plasmids in population diversity. Environ Microbiol 11: 349–359.
Penno S, Lindell D, Post AF . (2006). Diversity of Synechococcus and Prochlorococcus populations determined from DNA sequences of the N-regulatory gene ntcA. Environ Microbiol 8: 1200–1211.
Ramette A, Tiedje JM . (2007). Biogeography: an emerging cornerstone for understanding prokaryotic diversity, ecology, and evolution. Microb Ecol 53: 197–207.
Richardson TL, Jackson GA . (2007). Small phytoplankton and carbon export from the surface ocean. Science 315: 838–840.
Rocap G, Distel DL, Waterbury JB, Chisholm SW . (2002). Resolution of Prochlorococcus and Synechococcus ecotypes by using 16S-23S ribosomal DNA internal transcribed spacer sequences. Appl Environ Microbiol 68: 1180–1191.
Smith EM, Kemp WM . (2001). Size structure and the production/respiration balance in a coastal plankton community. Limnol Oceanogr 46: 473–485.
Strom SL . (1991). Growth and grazing rates of the herbivorous dinoflagellate Gymnodinium sp from the open subarctic Pacific Ocean. Mar Ecol Prog Ser 78: 103–113.
Suttle CA, Chan AM . (1993). Marine cyanophages infecting oceanic and coastal strains of Synechococcus—abundance, morphology, cross-infectivity and growth characteristics. Mar Ecol Prog Ser 92: 99–109.
Tettelin H, Masignani V, Cieslewicz MJ, Donati C, Medini D, Ward NL et al. (2005). Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial ‘pan-genome’. Proc Natl Acad Sci USA 102: 13950–13955.
Toledo G, Palenik B . (1997). Synechococcus diversity in the California current as seen by RNA polymerase (rpoC1) gene sequences of isolated strains. Appl Environ Microbiol 63: 4298–4303.
Toledo G, Palenik B . (2003). A Synechococcus serotype is found preferentially in surface marine waters. Limnol Oceanogr 48: 1744–1755.
Toledo G, Palenik B, Brahamsha B . (1999). Swimming marine Synechococcus strains with widely different photosynthetic pigment ratios form a monophyletic group. Appl Environ Microbiol 65: 5247–5251.
Waterbury JB, Valois FW . (1993). Resistance to co-occurring phages enables marine Synechococcus communities to coexist with cyanophages abundant in seawater. Appl Environ Microbiol 59: 3393–3399.
Waterbury JB, Watson SW, Guillard RRL, Brand LE . (1979). Widespread occurrence of a unicellular, marine, planktonic, cyanobacterium. Nature 277: 293–294.
Waterbury JB, Watson SW, Valois FW, Franks DG . (1986). Biological and ecological characterization of the marine unicellular cyanobacterium Synechococcus. Can Bull Fish Aquat Sci 214: 71–120.
Wilson WH, Joint IR, Carr NG, Mann NH . (1993). Isolation and molecular characterization of five marine cyanophages propagated on Synechococcus sp. strain WH7803. Appl Environ Microbiol 59: 3736–3743.
Winant CD, Bratkovich AW . (1981). Temperature and currents on the southern California shelf—a description of the variability. J Phys Oceanogr 11: 71–86.
Worden AZ, Nolan JK, Palenik B . (2004). Assessing the dynamics and ecology of marine picophytoplankton: the importance of the eukaryotic component. Limnol Oceanogr 49: 168–179.
Zeidner G, Preston CM, Delong EF, Massana R, Post AF, Scanlan DJ et al. (2003). Molecular diversity among marine picophytoplankton as revealed by psbA analyses. Environ Microbiol 5: 212–216.
Zubkov MV, Fuchs BM, Tarran GA, Burkill PH, Amann R . (2003). High rate of uptake of organic nitrogen compounds by Prochlorococcus cyanobacteria as a key to their dominance in oligotrophic oceanic waters. Appl Environ Microbiol 69: 1299–1304.
Zwirglmaier K, Heywood JL, Chamberlain K, Woodward EMS, Zubkov MV, Scanlan DJ . (2007). Basin-scale distribution patterns lineages in the Atlantic Ocean. Environ Microbiol 9: 1278–1290.
Zwirglmaier K, Jardillier L, Ostrowski M, Mazard S, Garczarek L, Vaulot D et al. (2008). Global phylogeography of marine Synechococcus and Prochlorococcus reveals a distinct partitioning of lineages among oceanic biomes. Environ Microbiol 10: 147–161.
Acknowledgements
Funding for this project was provided through a National Science Foundation grant (NSF OCE05-26979) to BP and Ron Burton (SIO). We thank Ron Burton for comments on the experimental protocols and data analysis. We also thank J McGowan, M Carter and M Hilbern for collecting and providing access to the Scripps Pier Chlorophyll Program dataset, funded by NOAA and COTS through the Southern California Coastal Ocean Observing System (SCCOOS).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Rights and permissions
About this article
Cite this article
Tai, V., Palenik, B. Temporal variation of Synechococcus clades at a coastal Pacific Ocean monitoring site. ISME J 3, 903–915 (2009). https://doi.org/10.1038/ismej.2009.35
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2009.35
Keywords
This article is cited by
-
Spatio-temporal patterns of Synechococcus oligotypes in Moroccan lagoonal environments
Scientific Reports (2023)
-
Seasonal dynamics in picocyanobacterial abundance and clade composition at coastal and offshore stations in the Baltic Sea
Scientific Reports (2022)
-
Spatial Distribution Patterns of Bacterioplankton in the Oxygen Minimum Zone of the Tropical Mexican Pacific
Microbial Ecology (2020)
-
Parallel phylogeography of Prochlorococcus and Synechococcus
The ISME Journal (2019)
-
Latitudinal and Vertical Variation of Synechococcus Assemblage Composition Along 170° W Transect From the South Pacific to the Arctic Ocean
Microbial Ecology (2019)