Abstract
Similar to virtually all components of natural environments, microbial systems are inherently complex and dynamic. Advances in cultivation-independent molecular methods have provided a route to study microbial consortia in their natural surroundings and to begin resolving the community structure, dominant metabolic processes and inter-organism interactions. However, the utility of these methods generally scales inversely with community complexity. By applying genomics-enabled methods to the study of natural microbial communities with reduced levels of species richness, a relatively comprehensive understanding of the metabolic networks and evolutionary processes within these communities can be attained. In such well-defined model systems, it is also possible to link emergent ecological patterns to their molecular and evolutionary underpinnings, facilitating construction of predictive ecosystem models. In this study, we review over a decade of research on one such system—acid mine drainage biofilm communities. We discuss the value and limitations of tractable model microbial communities in developing molecular methods for microbial ecology and in uncovering principles that may explain behavior in more complex systems.
Similar content being viewed by others
Editor's note
Professor Jillian (Jill) Banfield was invited to contribute a Winogradsky review based on her recognized contributions to the field of microbial ecology using the acid mine drainage (AMD) biofilm as a model microbial community. During her studies of this defined community she and her co-workers established the new field of community proteogenomics, developed binning techniques for microbial community genome assemblies and recently determined the importance of CRISPR elements in shaping the community. Insight gained from the AMD biofilm community has led to increased understanding of how microbial populations evolve within communities and have established a paradigm for exploration of more complex microbial ecosystems.
Introduction
Model systems and the real world
Questions relating to microbial evolution and ecology are often approached using model organisms (e.g. for studying the relationship between sequence divergence and sexual isolation (Roberts and Cohan, 1993) and model consortia (e.g., to study the balance between bottom-up and top-down controls on community structure (Bohannan and Lenski, 2000)). While these studies have been invaluable in furthering our understanding of the natural world, they capture neither the role of the uncultivated majority within communities nor the fact that (micro)organisms generally occur in communities made up of natural populations with inherent microdiversity (that is, not clonal populations; reviewed in Wilmes et al. (2009b)).
Studies of natural microbial consortia
Analysis of whole ecosystems with all of their attendant diversity is a hard problem. There has been progress toward addressing ecological and evolutionary questions in situ using marker gene-based culture-independent methods (Horner-Devine et al., 2004). Despite some success, genomic heterogeneity between strains with nearly identical (Konstantinidis et al., 2006) and identical (Simmons et al., 2008) 16S rRNA genes complicates interpretation of marker-based patterns. Evidence that fine-scale heterogeneity is ecologically relevant in natural populations (for example, Hunt et al., 2008) underlines the need to extend analyses toward the whole genome level. However, the complexity and poorly defined structural and functional boundaries of most natural microbial communities have precluded most ecological and evolutionary studies from achieving the level of detail possible for laboratory isolate-based model systems.
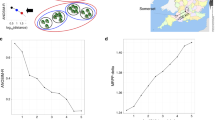
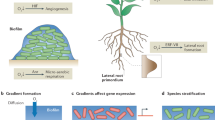
Clearly, systems with reduced complexity, either due to low species richness or dominance of one or a few populations, offer a special opportunity for analyses of community functioning. For over a decade, we have used molecular methods to study relatively low diversity biofilms that grow in the subsurface, where microbial activity stimulates pyrite (FeS2) dissolution to form acidic, metal-rich solutions (Figure 1a). Such solutions form naturally when sulfides are exposed to air, water and microbial communities, and are often an undesirable effluent from mining sites (acid mine drainage, AMD).
(a) Schematic illustrating important features that make the AMD system a good model for studying microbial communities (for example, relatively low species complexity, defined ecological succession patterns and trophic levels, tight biological–geochemical coupling, high biological productivity). (b) Picture of in situ biofilm developmental stages. (c) Rank–relative abundance curve for the AMD system communities. Sequencing reads from three Sanger sequencing libraries (∼100 Mb each) that overlapped with a 16S rRNA gene were identified based on the best match in a database specific for the AMD system using BLAST. Proteomics data were analyzed by assigning each identified protein to the corresponding organism and calculating its relative representation as a fraction of all proteins identified. Error bars indicate the s.d. for each rank across samples. Although diversity is lower in the AMD system, the relationship is similar to various more complex systems (see for example Fuhrman et al., 2008).
AMD biofilms: a tractable model microbial ecosystem
Background
At our field site within the Richmond Mine at Iron Mountain in northern California, chemoautotrophic biofilms establish and mature at the air–solution interface of streams and pools that overlie sediment composed of quartz and sulfide (primarily pyrite) minerals (Figure 1b). Carbon fixation in AMD biofilms is driven by oxidation of iron released from the dissolving pyrite surface, which is coupled to proton gradient-driven ATP formation. The by-product, ferric iron, reacts with pyrite surface sulfide/sulfur groups, stimulating further mineral dissolution (Figure 1a). In this environment, air, water and minerals sustain life, without measurable inputs from sunlight. Thus, the biofilms can be viewed as a relatively self-contained ecosystem in which there is tight and clearly defined coupling between inorganic and biological processes (Baker and Banfield, 2003; Druschel et al., 2004) (Figures 1 and 2). Because microbial activity drives mineral dissolution, similar consortia are harnessed for bioleaching-based recovery of metals from sulfide phases (Bosecker, 1997).
Ecosystem models beyond the ‘black box’. Combining insights from cultivation-dependent and -independent studies, metabolic roles can be assigned to the different members of the community (different symbols as per legend, symbol size indicates population size). Insights from population dynamics and proteomics data allow the determination of the relative roles of different populations (difference in thickness of the arrows, full lines=input; dotted lines=output). There is a division of metabolic roles as a function of biotic and abiotic environmental conditions. As an example, we here show shifts across different biofilm development stages. Parts of the nitrogen and sulfur cycles, the recycling of carbon after biofilm collapse and geochemical processes associated with the planktonic and sediment compartments still need to be linked to the specific populations involved. Quantifying fluxes of carbon and nitrogen through the system will be a focus of future research.
It has been about 15 years since our group and collaborators first performed molecular studies on AMD communities that grow underground in the Richmond Mine. Initial 16S rRNA gene clone library and fluorescent in situ hybridization-based analyses revealed lower complexity than most natural systems (Figure 1c). We attribute the relatively low diversity, now evident at all taxonomic and trophic levels (Baker et al., 2009), to the low solution pH (typically 0.3–1.2), high metal concentrations, and limited resource diversity. Low diversity and ability to resample defined structures and reproducible stages of biofilm succession over time (Wilmes et al., 2009a) make AMD communities amenable targets for detailed cultivation-independent studies (Figure 1).
Many of the features contributing to the reduced complexity of AMD systems separate them from most other environments. This raises the question whether processes and interactions within this unusual growth environment are representative of those that occur in other environments. For example, as discussed further below, reproducible patterns of community assembly are observed. This may distinguish the AMD system from others, since qits lower source diversity might drive these stable and predictable patterns (Curtis and Sloan, 2004). Nonetheless, evidence suggests community assembly in more diverse systems to be a non-random process, which can be predicted based on environmental parameters (Fuhrman et al., 2006; Horner-Devine et al., 2007). Moreover, despite conditions perceived by us as ‘extreme’, primary production rates in the AMD system are high, comparable on a g C m−2 y−1 basis to rates for many terrestrial and ocean ecosystems (Belnap et al., 2010). Rates are also similar on a per cell basis to those achieved by microbial primary production in the ocean (Belnap et al., 2010). Interestingly, this implies that carbon fixation rates in ∼100 μm thick biofilms are broadly equivalent to those achieved across the ocean photic zone. Comparable levels of ecosystem function across a very large range of species richness and physical scale increases our confidence that ecological principles inferred in the AMD system may predict patterns in more complex systems.
No model system can be expected to capture all features of other systems. Therefore, an important question is whether basic biological processes differ fundamentally across ecosystem types. To answer this, one may take advantage of model system tractability to generate simple and clearly defined hypotheses to be tested across a range of environments. By understanding unique features of the systems where each hypothesis is tested, one can gain insight into conserved features across systems and into contingencies specific to sets of systems. This philosophy echoes abovementioned approaches used in model organism-based research. Moreover, new methods are almost certainly best devised and tested in less complex systems, thereby providing templates for approaches to interrogate more complex and less well-defined environments.
Early research: addressing biological questions while developing novel approaches
The first molecular studies in the Richmond Mine AMD system changed our thinking of which lineages were directly responsible for mineral dissolution at low pH (<1.5), switching focus from the relatively culturable Acidithiobacillus ferrooxidans (γ-Proteobacteria class) to various less easily cultivated Leptospirillum species (Nitrospira class) (Schrenk et al., 1998). They also revealed the importance of archaeal mixotrophs belonging to the Thermoplasmata class (Edwards et al., 2000). Using 16S rRNA gene clone libraries and fluorescent in situ hybridization, these initial studies were also able to link aspects of community dynamics to environmental parameters. For example, seasonal drops in pH and increases in conductivity, resulting from the winter rain-driven displacement of highly concentrated AMD deep in the underground system, correlate to increases in the relative abundance of archaea (Edwards et al., 1999; Bond et al., 2000).
The ability to understand the physiological and ecological basis for spatiotemporal patterns in community structure is limited because many AMD community members are difficult to isolate. Arduous work to culture Ferroplasma and Leptospirillum species has classified these as mixotrophs and obligate chemolithoautotrophs, respectively (Johnson, 1998; Edwards et al., 2000; Dopson et al., 2004; Tyson et al., 2005). The need for understanding the metabolic roles of all community members motivated the development of approaches to recover genomic information directly from the environment (Stein et al., 1996). The comparatively low species richness of AMD biofilms made them ideal candidates for an effort to recover near-complete gene complements for all abundant organisms in a microbial community.
Community metagenomics
Using Sanger sequencing of a small-insert library (sequencing of both ends of ∼3 kb inserts) 76 Mbp of data were acquired from a biofilm community from the Richmond Mine. These data were assembled to generate near-complete composite genomes for Leptospirillum Group II and a Ferroplasma type II population, as well as partial genomic data sets for three other populations (Tyson et al., 2004). The Leptospirillum sequences provided the first genomic insights for a member of the Nitrospira phylum. The genomic inference that the Leptospirillum Group III population has the sole ability to fix N2 in the community was used to isolate this organism, now described as Leptospirillum ferrodiazotrophum (Tyson et al., 2005).
It is important to note that relatively deep sequence coverage (∼6–25 times) enabled reconstruction of near-complete genomic representations of the dominant organisms in several AMD communities without reference to isolate genomes. In fact, using ∼400 Mbp of Sanger sequencing data from multiple libraries (cumulatively, less than the equivalent of one plate of 454 titanium sequencing), it has been possible to recover ∼12 near-complete bacterial and archaeal population genomic data sets (Tyson et al., 2004; Lo et al., 2007; Dick et al., 2009). Several of these population genomic datasets have been analyzed to infer metabolic niches (Baker et al., submitted; Goltsman et al., 2009; Tyson et al., 2004) (Figure 2). The most detailed annotation has been performed for Leptospirillum Groups II and III, which allowed us to describe how these organisms fix carbon and generate energy through novel iron oxidation pathways (Goltsman et al., 2009).
Insights into the biology of lineages that are unrepresented in current isolate-based genomic databases are the major advantage of community genomics methods. At first, it was unclear how broadly applicable these techniques would be for the reconstruction of population genomic data sets from natural communities in other environments. However, as evidenced by multiple studies since, the approach has been successful across a range of levels of system complexity, either when applied directly (Legault et al., 2006; Martin et al., 2006; Woyke et al., 2006; Chivian et al., 2008) or to molecular enrichments that focus on a segment of the whole community (Baker et al., submitted; Hallam et al., 2006; Pernthaler et al., 2008).
Bacterial, archaeal and viral population metagenomics
Some of the most interesting findings from deep genomic sampling of AMD communities relate to evolutionary processes inferred from within-population genetic diversity. The dominant Leptospirillum Group II population displayed low polymorphism frequencies (0.08%), whereas a polymorphism rate of 2.2% was documented for the archaeal Ferroplasma Type II population (Tyson et al., 2004). Closer inspection of the sequencing reads revealed that the archaeal population consisted of multiple strains with mosaic genomes resulting from extensive homologous recombination involving approximately three closely related but distinct sequence types (Tyson et al., 2004). A detailed analysis of the Ferroplasma type I population revealed similar patterns. It was proposed that this process maintains diversity in the face of selection events, and thus confers increased population-level resilience to environmental perturbations (Allen et al., 2007). Despite some skepticism about the initial results (Delong, 2004), similar findings, mostly based on multilocus sequence typing, have emphasized the importance of recombination as an evolutionary process in other natural populations. These include findings for Sulfolobus icelandicus from hot springs (Whitaker et al., 2005), Halorubrum in hypersaline environments (Papke et al., 2004), and medically relevant organisms such as Pneumococcus (Hanage et al., 2005).
Although deeply sampled community genomic data sets were clearly treasure troves for molecular studies, using them to answer evolutionary questions required new methods to visualize and analyze population genetic data. Eppley et al. (2007b) developed the program ‘Strainer’ for this purpose. Strainer was used to comprehensively analyze sequence variation and recombination patterns in two Ferroplasma populations. These authors showed that both intra- and interpopulation recombination rates followed a log-linear relationship with sequence divergence (Eppley et al., 2007a). Eppley et al. also documented recombination across the species boundary, possibly initiated within genomic regions that shared unusually high sequence identity. This is in line with a recent model involving temporal fragmentation of speciation (Retchless and Lawrence, 2007).
Allen et al. (2007) evaluated the form of population-level heterogeneity by comparison of environmental population sequencing reads with the genome of a Ferroplasma acidarmanus fer1 isolate originating from the same site some years earlier. The finding of high levels of within-population variation resulting from phage, plasmid and transposase insertion and deletion are substantiated by data from other systems (Coleman et al., 2006; Cuadros-Orellana et al., 2007; Rusch et al., 2007).
Genome-wide determination of the rates of synonymous vs nonsynonymous polymorphisms in the dominant and deeply sampled bacterial population was used to evaluate evidence for neutral vs selective processes (Simmons et al., 2008). On the basis of the absence of strong indications of positive selection for the maintenance of variation, the authors suggested a population natural history in which allopatric speciation was followed by migration into the same location and recombination (Figure 3). This evolutionary model is in line with inferences made previously for the archaeal populations in this system (Allen et al., 2007).
Insights into links between evolutionary and ecological processes at different levels of resolution. (a) Although insights were derived for many archaeal and bacterial populations, this figure just focuses on the dominant Leptospirillum Group II bacteria. Two populations that derived from a common ancestor, as well as variants arising from recombination between them, are found in the Richmond Mine. Rectangles group the population(s) for which more details are shown in b–e. (b) The ecological distribution of the two populations and their recombinants primarily correlates with biofilm developmental stage (DS) and temperature (T). (c) Proteins encoded in hypervariable regions were rarely identified by proteomics, indicating that many have little or no role in environmental adaptation. Differential expression of shared genes is implicated in ecological divergence. (d) Population genomic analysis of single-nucleotide polymorphisms (SNPs) indicated variation within one population is not maintained by positive selection, but rather reflects allopatric divergence, migration and recombination, with some loss of variants. (e) Variation approaching the individual level is apparent in the clustered regularly interspaced short palindromic repeat (CRISPR) loci, which provide immunity against predation by specific viral/phage variants. Patterns of variation indicate that viral predation can shape population and community structure. Similar ‘hot spots’ of variability are found elsewhere in the genome, for example, at loci encoding cytochromes involved in Fe oxidation, raising questions about their potential role in fine-scale environmental adaptation.
Extremely high population-level diversity, nearing cell individuality, at the clustered regularly interspaced short palindromic repeat (CRISPR) locus, was an intriguing finding from two population genomic data sets for different Leptospirillum Group II strains (Tyson and Banfield, 2008) (Figure 3). The locus and associated proteins are now known to be involved in bacterial and archaeal defense against viruses and phage (Makarova et al., 2006; Barrangou et al., 2007; Brouns et al., 2008). Comparison of the set of spacer loci (regions between the repeats transcribed as CRISPR RNAs) and the organization within and between the Leptospirillum populations indicated unidirectional locus expansion and provided evidence for lateral gene transfer of the locus. Notably, inferences about locus and virus–host interaction dynamics based on data from these natural populations are completely consistent with those from laboratory studies in which loci in Streptococcus thermophilis are activated by phage challenge (Barrangou et al., 2007; Horvath et al., 2008).
Given that spacers in the CRISPR locus derive directly from viral genomes, they could be used to identify and assemble viral genomes from the community genomic data sets (Andersson and Banfield, 2008). The deeply sampled virus population genomic data sets revealed extensive recombination among closely related virus types. Correspondence of the recombination block size to the CRISPR spacer length for one AMD viral population suggests that viral recombination can result in evasion of the CRISPR-based host defense system (Andersson and Banfield, 2008). More recently, the CRISPR locus carried by plasmids in the AMD system was shown to target the protein machinery of the CRISPR locus of other plasmids (Goltsman et al., 2009), indicating a broader role of the defense system (for example, in plasmid–plasmid competition). Such analyses have begun to bridge the gap between evolution and ecology by providing insight into the molecular dynamics resulting from predator/prey interactions, and by conclusively linking phage/viral populations with their hosts (Figure 3).
Metagenomic analyses of less abundant organisms in communities
In obtaining a system-level understanding of ecological function in the AMD model system, most attention has been focused on the abundant organisms (comprising more than ∼10% of the community), on the basis of the assumption that the majority of biological processes will be carried out by a few abundant populations (for example, paralleling the macroeconomic Pareto principle; Dejonghe et al., 2001). Nevertheless, it has been shown that a relatively low-abundance organism (∼0.3% of all cells) can carry out significant fractions of metabolic fluxes (Musat et al., 2008). We have made progress with analysis of less abundant AMD community members. For example, Baker et al. (2006) discovered an enigmatic group of ultra-small archaea, represented by only a few sequence fragments, some of which contained rRNA genes <85% identical to any known sequence. Subsequent three-dimensional cryogenic electron tomography revealed that these tiny cells contain an average of only 92 ribosomes and have large internal tubular organelles of unknown function (Comolli et al., 2009). Intriguingly, these community members seem to be highly targeted by viruses, and often by multiple morphotypes simultaneously. The combination of cryo-electron microscopy and targeted metagenomic analyses (Baker et al., submitted) holds promise to unveil unexpected details of biology of low-abundance organisms.
The ability to interpret metagenomic sequence information to achieve ecological and evolutionary insights depends highly on methods to assign sequence fragments to organisms. Thus, we used curated composite genomes for nine archaeal, three bacterial and many virus populations, in combination with thousands of unassigned fragments from strain variants and low-abundance organisms to evaluate sequence-binning techniques. Dick et al. (2009) reported that a particularly effective method is based on emergent self-organizing maps (ESOM) of the tetranucleotide composition of genomic fragments (Teeling et al., 2004; Abe et al., 2005). This study also provided insights into the sources of genome signatures that distinguish coexisting populations (Dick et al., 2009). ESOM-based clustering revealed several clusters of genome fragments that, based on marker genes, could be assigned to specific low-abundance (percent-level) bacterial and archaeal populations. Despite the inability to assemble these fragments into large genomic stretches, insights into the biology of the minor community members can be achieved based on the analysis of the binned gene inventories (Dick et al., in preparation) (Figure 2).
Surveys of biofilm community composition from across many Richmond Mine environments, optimized by new genomic binning approaches, have documented ∼20 predominant taxa, with each community usually dominated by only 4–6 taxa. We now know that AMD biofilm communities include representatives from at least two archaeal and eight bacterial divisions, as well as plasmids and viral (phage) populations (Andersson and Banfield, 2008). In addition, microscopic and 18S rRNA-based analyses have documented several fungal and protist species in mature biofilms (Baker and Banfield, 2003; Baker et al., 2009). These results highlight the existence of multiple trophic levels with top-down predation, emergent patterns of organization (Denef et al., 2010; Wilmes et al., 2009a), succession, metabolic state switches on maturation (Mueller et al., submitted) and ecosystem stability founded on diversity (Denef et al., 2010) (Figures 1 and 2). These are the hallmarks of a complex biological system. For this reason, we conclude that the AMD system is not ‘simple’ (Handelsman et al., 2007). The already apparent parallels with other environments (reviewed above) are strong indications that ecological and evolutionary insights from the AMD system will indeed be broadly applicable.
Functional analysis of microorganisms in communities through proteomics
Possession of a gene does not equate to gene function. Community proteomic analyses (also referred to as environmental or metaproteomic analyses) provide a route to link genetic potential with activity. Community proteomic analysis was achieved with unprecedented depth by using the extensive set of gene predictions from an AMD biofilm to identify proteins through mass spectrometry-based methods capable of high mass accuracy measurements. In the initial study, Ram et al. (2005) extracted proteins from a biofilm similar to one for which genomic data were available, digested the proteins into peptides, separated the peptides by two-dimensional liquid chromatography, measured mass to charge ratio (m/z) values for peptides and their fragmentation products (high-throughput ‘shotgun’ proteomics) and identified >2000 proteins. This identified protein set comprised ∼50% of the predicted proteins from the dominant organism, but only ∼5% of proteins from the least abundant of the five genomically characterized members. Deep coverage of the proteomes of low-abundance members is often elusive, even in more recent analyses. Therefore, methodological advances, particularly those involving better protein/peptide separation methods (VerBerkmoes et al., 2009a, 2009b), will be essential for obtaining comprehensive proteomes from more diverse samples.
Despite current technical limitations, important insights have been gleaned from community proteomic studies. The first such study on the AMD system identified hundreds of proteins of unknown function that were previously classified as predicted proteins (Ram et al., 2005). It is notable that several were located in genomic regions associated with mobile elements that, in general, do not have many proteins that are identified by mass spectrometry. However, some proteins of unknown function were very abundant, and were targeted for detailed analyses using cultivation-independent biochemical methods. This study led to the characterization of two cytochromes involved in a new iron oxidation pathway (Jeans et al., 2008; Singer et al., 2008). Visualization of newly identified cytochromes in intact biofilms revealed high concentrations of the protein only at the interface between the lower surface of the biofilm and the solution, indicating spatially heterogeneous activity levels at the tens of microns scale (Wilmes et al., 2009a). Comparatively high levels of sequence variation have been documented in these cytochromes relative to the rest of the genome. This finding suggests that cytochrome sequence variation has a role in adaptation to redox gradients within the biofilms (Singer et al., submitted; Jeans et al., 2008; Singer et al., 2008). This type of ecological differentiation based on cytochrome fine-tuning is supported by experimental studies of Springs et al. (2002), which detected large (∼160 mV) changes in redox potential of cytochromes that differ by only a few amino-acid substitutions.
Proteomic data have been invaluable aids to metabolic interpretations based on genome reconstruction (Goltsman et al., 2009). For example, high abundances of pyruvate ferredoxin oxidoreductase and other functionally annotated proteins were used to deduce core metabolic pathways, including the route for carbon fixation, in Leptospirillum Groups II and III. Microscopy of intact biofilms showed that Leptospirillum Group II initiates biofilm colonization and that Leptospirillum Group III generally occurs as microcolonies or single cells dispersed throughout the biofilm (Wilmes et al., 2009a). Observations such as the high numbers of sensory genes and abundant expression of these as well as motility proteins in this organism are consistent with these ecological patterns (Goltsman et al., 2009) (Figure 2).
Ecological insights can also be achieved using proteomic analyses of large numbers of biofilm communities. For example, we have quantitatively analyzed protein-abundance patterns in specific organisms across a range of environmental (inorganic and biological) conditions. Notably, protein-abundance patterns of the dominant organism were most highly and significantly correlated with community composition, which is linked to changes in organismal membership over the course of ecological succession (Mueller et al., submitted). In addition, abundances of proteins from less well represented populations correlated with inorganic parameters, suggesting distinct environmental niches for these members.
Strain-resolved proteomics
Shotgun community proteomics potentially can be used to distinguish expression patterns for closely related orthologous proteins to understand the importance of strain-level differentiation. A single amino-acid substitution in a peptide almost always changes the peptide mass (isoleucine/leucine being the exception), precluding peptide identification and reducing protein identification rates (especially for relatively low-abundance proteins). Thus, as the sequence of an organism in a sample diverges from the sequences in reference databases, protein identification is restricted (identification at <85% amino-acid identity is generally precluded) and abundance metrics become inaccurate (Denef et al., 2007). The high mass accuracy of current mass spectrometers allows for the discrimination of all peptides originating from closely related organisms (thus proteins), so long as sequences for alternative peptides are available. To take advantage of this capability, we have created databases of potential protein variants in the AMD system by teasing apart sequence variation that underlies the composite sequences in metagenomic data sets.
In several studies, we have distinguished single (Ram et al., 2005) and groups of closely related candidate proteins in natural community samples (Lo et al., 2007; Wilmes et al., 2008; Denef et al., 2009). By performing this analysis at the peptide, protein and genome level, it is possible to infer the genotype of a microbial population in a genomically uncharacterized sample. A first application of this method revealed that proteins encoded in some genomic regions matched one candidate genome type, whereas other regions matched a second, closely related genotype (∼95% amino-acid identity). The approach, referred to as proteomics-inferred genome typing (PIGT), provided evidence for recombination involving sequence blocks of tens to hundreds of kilobases from two closely related Leptospirillum Group II populations (Lo et al., 2007). PIGT on a more extensive sample set led to detection of multiple new genotypes comprised of different mixtures of the end member strain sequences (>1000 proteins evaluated) and provided support for the notion that homologous recombination is used as a strategy for fine-scale environmental adaptation (Denef et al., 2009) (Figure 3). Very recently, the PIGT approach was expanded to carry out proteomic analyses of another system (Geobacter-dominated communities) for which metagenomic sequences were not available (Wilkins et al., 2009).
The ability to detect recombination directly in the environment using a high-resolution, culture-independent method is closely linked to the ability to detect changes in protein-abundance levels, and thus activity levels, with strain-level resolution. Previous studies have focused on the role of gene content differences between related strains/ecotypes/species in ecological divergence (for example, Kettler et al., 2007). In the AMD system, strain-resolved proteomics was applied to evaluate the relative roles of lateral gene transfer and changes in gene regulation in ecological divergence of two closely related populations (differing by 0.3% at the 16S rRNA gene level) (Figure 3). Proteins shared between populations were commonly identified and showed divergent expression levels, even when these populations co-occurred (Denef et al., 2010). Such findings illustrate that very closely related organisms can have distinct ecological roles, and indicate a key process in lineage divergence is proteome optimization, presumably driven by changes in genome regulation. The importance of looking beyond traditionally defined phylogenetic groups to understand ecological and evolutionary dynamics is currently being addressed in a variety of other systems as well (Hunt et al., 2008; Wilmes et al., 2008; Konstantinidis et al., 2009).
Bringing complex biofilm consortia into the laboratory for hypothesis testing
Characterization of a series of environmental communities, as described above, is insightful by itself and is also crucial for the generation of hypotheses that can then be tested in controlled laboratory settings. The power to address biological questions using laboratory-grown communities derived from natural inocula is well established (for example, Fernandez et al., 1999). For the AMD system, we have developed methods to reproducibly grow multispecies biofilms in laboratory bioreactors. Proteomic methods have been key to optimizing growth conditions and verifying that communities are comparable at the functional and compositional levels with natural biofilms (Belnap et al., 2010). An important advance in this study was achieved by mixing 15N-labeled laboratory-grown biofilms with the natural biofilms samples with which they were being compared. This allowed for relative quantification of peptides from laboratory-grown and natural communities using spectral peak areas detected in the mass spectrometry experiment. The ability to manipulate these laboratory-grown biofilms offers important experimental opportunities that are being exploited in ongoing research.
Future studies in the AMD model system
The overall tractability of the AMD system for molecular studies makes it a logical target for transcriptomic and metabolomic analyses, as well as studies that incorporate the roles of eukaryotes. But when such data are on hand, will it be possible to integrate metrics of activity and interaction into predictive models? There is no question that there are remaining barriers that need to be overcome before this goal is achieved in the AMD (and probably all other) systems. Among the most pressing of these is the gap in knowledge associated with the many proteins of unknown function. We hope that heterologous expression and subsequent biochemical screening of a large fraction of these, which have been targeted based on integration of metagenomic and proteomic data, can help to reveal their functions (Yakunin et al., 2004; Kuznetsova et al., 2005). Another challenge will be to define ecosystem indicators (for example, biomarkers in the form of metabolite, protein or transcript pools) that signify ecosystem state or the onset of a transition. Such methods can be developed for model systems such as AMD biofilms, and could ultimately find application in very complex environments where comprehensive analysis is neither affordable nor practical.
Scaling to more complex systems
It took 25 years to make the leap from the comprehensive sequencing of the first DNA virus, bacteriophage phi X174 (Sanger et al., 1977) to the human genome (Lander et al., 2001; Venter et al., 2001), which is approximately one million times larger. Just 3 years after the first AMD metagenomic data set was published, the amount of sequencing attributed to a sample set had increased about 100-fold (Tyson et al., 2004; Rusch et al., 2007) and soon the factor will exceed 10 000 (for example, the Terragenome project, which aims at obtaining a comprehensive metagenomic data set from soil Vogel et al., 2009).
The current expansion of sequencing capabilities clearly places almost any system within the reach of metagenomics (Wilmes et al., 2009b). In addition, subcompartments of very complex systems can be analyzed using molecular enrichment or screening methods (Hallam et al., 2006; Pernthaler et al., 2008) and single cell genomics (Marcy et al., 2007). Although there are some important issues regarding the accuracy of next-generation sequencing technologies, which currently limit their utility for population genetic analysis (Harismendy et al., 2009), the combination of new technical developments and co-assembly of multiple data types should address this problem (Reinhardt et al., 2009). Beyond this, the challenges with working with such vast data sets and complications due to the presence of many closely related genotypes are not yet fully appreciated. Bioinformatic tools and approaches, such as those being developed and refined through application to the AMD system, provide a starting point. However, it remains to be seen whether clear patterns and principles can be extracted from terragenome-scale efforts applied to complex, heterogeneous, and temporally varying environments. A limitation of the AMD system is that it may not anticipate the level of difficulty that will be encountered in this transition. However, progress in systems of intermediate (and even moderately high) diversity levels provide some encouragement that scaling of methods across complexity levels will be possible (Gill et al., 2006).
At the functional level, continuing advances in proteomics (Sowell et al., 2008; Verberkmoes et al., 2009a, 2009b), transcriptomics (Frias-Lopez et al., 2008) and imaging techniques (Huang et al., 2007; Behrens et al., 2008) will provide deeper insights into ecological and evolutionary questions for systems that are currently only being characterized using phylogenetic markers (Cruz-Martinez et al., 2009) or gene surveys (Tringe et al., 2005). Integration of these methods with cultivation-dependent methods will further our understanding of microbial and global ecosystem functioning (Mou et al., 2008).
Conclusion
The many features of the AMD system that render it particularly tractable for molecular-level, cultivation-independent analyses allow for identification of evolutionary mechanisms underlying strong and robust ecological patterns, despite the inherently noisy natural environment (Figure 1). Already, parallel findings in more complex systems are confirming the broader relevance of insights we have gathered regarding: (i) the character and ecological and functional roles of different forms of genomic diversity within and between populations (Figure 3); (ii) the interaction between bacteria, archaea, their phages and viruses (Figures 2 and 3); and (iii) relative rates of mutation and recombination (Figure 3). Such insights significantly advance our understanding of microbial physiology in the environment and the driving factors of community assembly, while laying the groundwork for the development of predictive models of community composition and metabolism in natural settings (Figures 1 and 2).
References
Abe T, Sugawara H, Kinouchi M, Kanaya S, Ikemura T . (2005). Novel phylogenetic studies of genomic sequence fragments derived from uncultured microbe mixtures in environmental and clinical samples. DNA Res 12: 281–290.
Allen EE, Tyson GW, Whitaker RJ, Detter JC, Richardson PM, Banfield JF . (2007). Genome dynamics in a natural microbial strain population. Proc Natl Acad Sci USA 104: 1883–1888.
Andersson AF, Banfield JF . (2008). Virus population dynamics and acquired virus resistance in natural microbial communities. Science 320: 1047–1050.
Baker BJ, Banfield JF . (2003). Microbial communities in acid mine drainage. FEMS Microbiol Ecol 44: 139–152.
Baker BJ, Comolli LR, Dick GJ, Hauser LJ, Hyatt D, Dill B, Land ML, VerBerkmoes NC, Hettich RL and Banfield JF . Enigmatic, ultra-small uncultivated Archaea. (Submitted)
Baker BJ, Tyson GW, Goosherst L, Banfield JF . (2009). Insights into the diversity of eukaryotes in acid mine drainage biofilm communities. Appl Environ Microbiol 75: 2192–2199.
Baker BJ, Tyson GW, Webb RI, Flanagan J, Hugenholtz P, Allen EE et al. (2006). Lineages of acidophilic archaea revealed by community genomic analysis. Science 314: 1933–1935.
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S et al. (2007). CRISPR provides acquired resistance against viruses in prokaryotes. Science 315: 1709–1712.
Behrens S, Losekann T, Pett-Ridge J, Weber PK, Ng W-O, Stevenson BS et al. (2008). Linking microbial phylogeny to metabolic activity at the single-cell level by using enhanced element labeling-catalyzed reporter deposition fluorescence in situ hybridization (EL-FISH) and nanoSIMS. Appl Environ Microbiol 74: 3143–3150.
Belnap CP, Pan C, VerBerkmoes NC, Power ME, Samatova NF, Carver RL et al. (2010). Cultivation and quantitative proteomic analyses of acidophilic microbial communities. ISME J 4: 520–530.
Bohannan BJM, Lenski RE . (2000). The relative importance of competition and predation varies with productivity in a model community. Am Nat 156: 329–340.
Bond PL, Druschel GK, Banfield JF . (2000). Comparison of acid mine drainage microbial communities in physically and geochemically distinct ecosystems. Appl Environ Microbiol 66: 4962–4971.
Bosecker K . (1997). Bioleaching: metal solubilization by microorganisms. FEMS Microbiol Rev 20: 591–604.
Brouns SJJ, Jore MM, Lundgren M, Westra ER, Slijkhuis RJH, Snijders APL et al. (2008). Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321: 960–964.
Chivian D, Brodie EL, Alm EJ, Culley DE, Dehal PS, DeSantis TZ et al. (2008). Environmental genomics reveals a single-species ecosystem deep within Earth. Science 322: 275–278.
Coleman ML, Sullivan MB, Martiny AC, Steglich C, Barry K, Delong EF et al. (2006). Genomic islands and the ecology and evolution of Prochlorococcus. Science 311: 1768–1770.
Comolli LR, Baker BJ, Downing KH, Siegerist CE, Banfield JF . (2009). Three-dimensional analysis of the structure and ecology of a novel, ultra-small archaeon. ISME J 3: 159–167.
Cruz-Martinez K, Suttle KB, Brodie EL, Power ME, Andersen GL, Banfield JF . (2009). Despite strong seasonal responses, soil microbial consortia are more resilient to long-term changes in rainfall than overlying grassland. ISME J 3: 738–744.
Cuadros-Orellana S, Martin-Cuadrado A-B, Legault B, D’Auria G, Zhaxybayeva O, Papke RT et al. (2007). Genomic plasticity in prokaryotes: the case of the square haloarchaeon. ISME J 1: 235–245.
Curtis TP, Sloan WT. (2004). Prokaryotic diversity and its limits: microbial community structure in nature and implications for microbial ecology. Curr Opin Microbiol 7: 221–226.
Dejonghe W, Boon N, Seghers D, Top EM, Verstraete W. (2001). Bioaugmentation of soils by increasing microbial richness: missing links. Environ Microbiol 3: 649–657.
Delong EF . (2004). Microbiology: reconstructing the wild types. Nature 428: 25–26.
Denef VJ, Kalnejais LH, Mueller RS, Wilmes P, Baker BJ, Thomas BC et al. (2010). Proteogenomic basis for ecological divergence of closely related bacteria in natural acidophilic microbial communities. Proc Natl Acad Sci USA 107: 2383–2390.
Denef VJ, Shah MB, Verberkmoes NC, Hettich RL, Banfield JF . (2007). Implications of strain- and species-level sequence divergence for community and isolate shotgun proteomic analysis. J Proteome Res 6: 3152–3161.
Denef VJ, Verberkmoes NC, Shah MB, Abraham P, Lefsrud M, Hettich RL et al. (2009). Proteomics-inferred genome typing (PIGT) demonstrates inter-population recombination as a strategy for environmental adaptation. Environ Microbiol 11: 313–325.
Dick G, Andersson A, Baker B, Simmons S, Thomas B, Yelton AP et al. (2009). Community-wide analysis of microbial genome sequence signatures. Genome Biol 10: R85.
Dopson M, Baker-Austin C, Hind A, Bowman JP, Bond PL . (2004). Characterization of ferroplasma isolates and Ferroplasma acidarmanus sp. nov., extreme acidophiles from acid mine drainage and industrial bioleaching environments. Appl Environ Microbiol 70: 2079–2088.
Druschel GK, Baker BJ, Gihring TM, Banfield JF . (2004). Acid mine drainage biogeochemistry at Iron Mountain, California. Geochem Trans 5: 13–32.
Edwards KJ, Bond PL, Gihring TM, Banfield JF . (2000). An archaeal iron-oxidizing extreme acidophile important in acid mine drainage. Science 287: 1796–1799.
Edwards KJ, Gihring TM, Banfield JF . (1999). Seasonal variations in microbial populations and environmental conditions in an extreme acid mine drainage environment. Appl Environ Microbiol 65: 3627–3632.
Eppley JM, Tyson GW, Getz WM, Banfield JF. (2007a). Genetic exchange across a species boundary in the archaeal genus Ferroplasma. Genetics 177: 407–416.
Eppley JM, Tyson GW, Getz WM, Banfield JF . (2007b). Strainer: software for analysis of population variation in community genomic datasets. BMC Bioinformatics 8: 398.
Fernandez A, Huang S, Seston S, Xing J, Hickey R, Criddle C et al. (1999). How stable is stable? Function versus community composition. Appl Environ Microbiol 65: 3697–3704.
Frias-Lopez J, Shi Y, Tyson GW, Coleman ML, Schuster SC, Chisholm SW et al. (2008). Microbial community gene expression in ocean surface waters. Proc Natl Acad Sci USA 105: 3805–3810.
Fuhrman JA, Hewson I, Schwalbach MS, Steele JA, Brown MV, Naeem S. (2006). Annually reoccurring bacterial communities are predictable from ocean conditions. Proc Natl Acad Sci USA 103: 13104–13109.
Fuhrman JA, Steele JA, Hewson I, Schwalbach MS, Brown MV, Green JL et al. (2008). A latitudinal diversity gradient in planktonic marine bacteria. Proc Natl Acad Sci USA 105: 7774–7778.
Gill SR, Pop M, Deboy RT, Eckburg PB, Turnbaugh PJ, Samuel BS et al. (2006). Metagenomic analysis of the human distal gut microbiome. Science 312: 1355–1359.
Goltsman DSA, Denef VJ, Singer SW, VerBerkmoes NC, Lefsrud M, Mueller RS et al. (2009). Community genomic and proteomic analysis of chemoautotrophic, iron-oxidizing ‘Leptospirillum rubarum’ (Group II) and Leptospirillum ferrodiazotrophum (Group III) in acid mine drainage biofilms. Appl Environ Microbiol 75: 4599–4615.
Hallam SJ, Konstantinidis KT, Putnam N, Schleper C, Watanabe Y-i, Sugahara J et al. (2006). Genomic analysis of the uncultivated marine crenarchaeote Cenarchaeum symbiosum. Proc Natl Acad Sci USA 103: 18296–18301.
Hanage WP, Fraser C, Spratt BG. (2005). Fuzzy species among recombinogenic bacteria. BMC Biol 3: 6.
Handelsman J, Tiedje JM, Alvarez-Cohen L, Ashburner M, Cann IKO, DeLong EF et al (2007). The New Science of Metagenomics: Revealing the Secrets of Our Microbial Planet. Committee on Metagenomics: Challenges and Functional Applications. The National Academies Press: Washington, DC.
Harismendy O, Ng P, Strausberg R, Wang X, Stockwell T, Beeson K et al. (2009). Evaluation of next generation sequencing platforms for population targeted sequencing studies. Genome Biol 10: R32.
Horner-Devine MC, Carney KM, Bohannan BJM. (2004). An ecological perspective on bacterial biodiversity. Proc Biol Sci 271: 113–122.
Horner-Devine MC, Silver JM, Leibold MA, Bohannan BJ, Colwell RK, Fuhrman JA et al. (2007). A comparison of taxon co-occurrence patterns for macro- and microorganisms. Ecology 88: 1345–1353.
Horvath P, Romero DA, Coute-Monvoisin AC, Richards M, Deveau H, Moineau S et al. (2008). Diversity, activity, and evolution of CRISPR loci in Streptococcus thermophilus. J Bacteriol 190: 1401–1412.
Huang WE, Stoecker K, Griffiths R, Newbold L, Daims H, Whiteley AS et al. (2007). Raman-FISH: combining stable-isotope Raman spectroscopy and fluorescence in situ hybridization for the single cell analysis of identity and function. Environ Microbiol 9: 1878–1889.
Hunt DE, David LA, Gevers D, Preheim SP, Alm EJ, Polz MF. . (2008). Resource partitioning and sympatric differentiation among closely related bacterioplankton. Science 320: 1081–1085.
Jeans C, Singer SW, Chan CS, VerBerkmoes NC, Shah M, Hettich RL et al. (2008). Cytochrome 572 is a conspicuous membrane protein with iron oxidation activity purified directly from a natural acidophilic microbial community. ISME J 2: 542–550.
Johnson DB. (1998). Biodiversity and ecology of acidophilic microorganisms. FEMS Microbiol Ecol 27: 307–317.
Kettler GC, Martiny AC, Huang K, Zucker J, Coleman ML, Rodrigue S et al. (2007). Patterns and implications of gene gain and loss in the evolution of Prochlorococcus. PLoS Genet 3: e231.
Konstantinidis KT, Ramette A, Tiedje JM . (2006). The bacterial species definition in the genomic era. Philos Trans R Soc Lond B Biol Sci 361: 1929–1940.
Konstantinidis KT, Serres MH, Romine MF, Rodrigues JLM, Auchtung J, McCue L-A et al. (2009). Comparative systems biology across an evolutionary gradient within the Shewanella genus. Proc Natl Acad Sci USA 106: 15909–15914.
Kuznetsova E, Proudfoot M, Sanders SA, Reinking J, Savchenko A, Arrowsmith CH et al. (2005). Enzyme genomics: application of general enzymatic screens to discover new enzymes. FEMS Microbiol Rev 29: 263–279.
Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, Baldwin J et al. (2001). Initial sequencing and analysis of the human genome. Nature 409: 860–921.
Legault BA, Lopez-Lopez A, Alba-Casado JC, Doolittle WF, Bolhuis H, Rodriguez-Valera F et al. (2006). Environmental genomics of ‘Haloquadratum walsbyi’ in a saltern crystallizer indicates a large pool of accessory genes in an otherwise coherent species. BMC Genomics 7: 171.
Lo I, Denef VJ, Verberkmoes NC, Shah M, Goltsman D, DiBartolo G et al. (2007). Strain-resolved community proteomics reveals that recombination shapes the genomes of acidophilic bacteria. Nature 446: 537–541.
Makarova KS, Grishin NV, Shabalina SA, Wolf YI, Koonin EV . (2006). A putative RNA-interference-based immune system in prokaryotes: computational analysis of the predicted enzymatic machinery, functional analogies with eukaryotic RNAi, and hypothetical mechanisms of action. Biol Direct 1: 7.
Marcy Y, Ouverney C, Bik EM, Losekann T, Ivanova N, Martin HG et al. (2007). Dissecting biological ‘dark matter’ with single-cell genetic analysis of rare and uncultivated TM7 microbes from the human mouth. Proc Natl Acad Sci USA 104: 11889–11894.
Martin HG, Ivanova N, Kunin V, Warnecke F, Barry KW, McHardy AC et al. (2006). Metagenomic analysis of two enhanced biological phosphorus removal (EBPR) sludge communities. Nat Biotechnol 24: 1263–1269.
Mou X, Sun S, Edwards RA, Hodson RE, Moran MA. (2008). Bacterial carbon processing by generalist species in the coastal ocean. Nature 451: 708–711.
Mueller RS, Denef VJ, Kalnejais LH, Suttle KB, Thomas BC, Wilmes P, Smith RL, Nordstrom DK, McCleskey RB, Shah MB, VerBerkmoes NC, Hettich RL and Banfield JF . Ecological Distribution and Population Physiology Defined By Proteomics in a Natural Microbial Community. (Submitted)
Musat N, Halm H, Winterholler Br, Hoppe P, Peduzzi S, Hillion F et al. (2008). A single-cell view on the ecophysiology of anaerobic phototrophic bacteria. Proc Natl Acad Sci USA 105: 17861–17866.
Papke RT, Koenig JE, Rodriguez-Valera F, Doolittle WF . (2004). Frequent recombination in a saltern population of Halorubrum. Science 306: 1928–1929.
Pernthaler A, Dekas AE, Brown CT, Goffredi SK, Embaye T, Orphan VJ . (2008). Diverse syntrophic partnerships from deep-sea methane vents revealed by direct cell capture and metagenomics. Proc Natl Acad Sci 105: 7052–7057.
Ram RJ, VerBerkmoes NC, Thelen MP, Tyson GW, Baker BJ, Blake II RC et al. (2005). Community proteomics of a natural microbial biofilm. Science 308: 1915–1920.
Reinhardt JA, Baltrus DA, Nishimura MT, Jeck WR, Jones CD, Dangl JL . (2009). De novo assembly using low-coverage short read sequence data from the rice pathogen Pseudomonas syringae pv. oryzae. Genome Res 19: 294–305.
Retchless AC, Lawrence JG. (2007). Temporal fragmentation of speciation in bacteria. Science 317: 1093–1096.
Roberts MS, Cohan FM. (1993). The effect of DNA sequence divergence on sexual isolation in Bacillus. Genetics 134: 401–408.
Rusch DB, Halpern AL, Sutton G, Heidelberg KB, Williamson S, Yooseph S et al. (2007). The Sorcerer II Global Ocean Sampling expedition: northwest Atlantic through eastern tropical Pacific. PLoS Biol 5: e77.
Sanger F, Air GM, Barrell BG, Brown NL, Coulson AR, Fiddes CA et al. (1977). Nucleotide sequence of bacteriophage phi X174 DNA. Nature 265: 687–695.
Schrenk MO, Edwards KJ, Goodman RM, Hamers RJ, Banfield JF . (1998). Distribution of Thiobacillus ferrooxidans and Leptospirillum ferrooxidans: implications for generation of acid mine drainage. Science 279: 1519–1522.
Simmons SL, DiBartolo G, Denef VJ, Goltsman DSA, Thelen MP, Banfield JF . (2008). Population genomic analysis of strain variation in Leptospirillum group II bacteria involved in acid mine drainage formation. PLoS Biol 6: e177.
Singer SW, Chan CS, Zemla A, VerBerkmoes NC, Hwang M, Hettich RL et al. (2008). Characterization of Cytochrome 579, an unusual cytochrome isolated from an iron-oxidizing microbial community. Appl Environ Microbiol 74: 4454–4462.
Singer SW, Erickson BK, VerBerkmoes NC, Hwang M, Shah MB, Hettich RL, Banfield JF, and Thelen MP . Post-translational modification and sequence variation of redox-active proteins correlate with biofilm lifecycle in natural microbial communities (submitted).
Sowell SM, Wilhelm LJ, Norbeck AD, Lipton MS, Nicora CD, Barofsky DF et al. (2008). Transport functions dominate the SAR11 metaproteome at low-nutrient extremes in the Sargasso Sea. ISME J 3: 93–105.
Springs SL, Bass SE, Bowman G, Nodelman I, Schutt CE, McLendon GL . (2002). A multigeneration analysis of cytochrome b(562) redox variants: evolutionary strategies for modulating redox potential revealed using a library approach. Biochemistry 41: 4321–4328.
Stein JL, Marsh TL, Wu KY, Shizuya H, DeLong EF . (1996). Characterization of uncultivated prokaryotes: isolation and analysis of a 40-kilobase-pair genome fragment from a planktonic marine archaeon. J Bacteriol 178: 591–599.
Teeling H, Waldmann J, Lombardot T, Bauer M, Glockner FO . (2004). TETRA: a web-service and a stand-alone program for the analysis and comparison of tetranucleotide usage patterns in DNA sequences. BMC Bioinformatics 5: 163.
Tringe SG, von Mering C, Kobayashi A, Salamov AA, Chen K, Chang HW et al. (2005). Comparative metagenomics of microbial communities. Science 308: 554–557.
Tyson GW, Banfield JF. (2008). Rapidly evolving CRISPRs implicated in acquired resistance of microorganisms to viruses. Environ Microbiol 10: 200–207.
Tyson GW, Chapman J, P H, Allen EE, Ram RJ, Richardson PM et al. (2004). Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature 428: 37–43.
Tyson GW, Lo I, Baker BJ, Allen EE, Hugenholtz P, Banfield JF . (2005). Genome-directed isolation of the key nitrogen fixer Leptospirillum ferrodiazotrophum sp. nov. from an acidophilic microbial community. Appl Environ Microbiol 71: 6319–6324.
Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, Sutton GG et al. (2001). The sequence of the human genome. Science 291: 1304–1351.
VerBerkmoes NC, Denef VJ, Hettich RL, Banfield JF . (2009a). Systems biology: functional analysis of natural microbial consortia using community proteomics. Nat Rev Microbiol 7: 196–205.
Verberkmoes NC, Russell AL, Shah M, Godzik A, Rosenquist M, Halfvarson J et al. (2009b). Shotgun metaproteomics of the human distal gut microbiota. ISME J 3: 179–189.
Vogel TM, Simonet P, Jansson JK, Hirsch PR, Tiedje JM, van Elsas JD et al. (2009). TerraGenome: a consortium for the sequencing of a soil metagenome. Nat Rev Microbiol 7: 252.
Whitaker RJ, Grogan DW, Taylor JW . (2005). Recombination shapes the natural population structure of the hyperthermophilic archaeon Sulfolobus islandicus. Mol Biol Evol 22: 2354–2361.
Wilkins MJ, VerBerkmoes NC, Williams KH, Callister SJ, Mouser PJ, Elifantz H et al. (2009). Proteogenomic monitoring of Geobacter physiology during stimulated uranium bioremediation. Appl Environ Microbiol 75: 6591–6599.
Wilmes P, Andersson AF, Lefsrud MG, Wexler M, Shah M, Zhang B et al. (2008). Community proteogenomics highlights microbial strain-variant protein expression within activated sludge performing enhanced biological phosphorus removal. ISME J 2: 853–864.
Wilmes P, Remis JP, Hwang M, Auer M, Thelen MP, Banfield JF . (2009a). Natural acidophilic biofilm communities reflect distinct organismal and functional organization. ISME J 3: 266–270.
Wilmes P, Simmons SL, Denef VJ, Banfield JF. (2009b). The dynamic genetic repertoire of microbial communities. FEMS Microbiol Rev 33: 109–132.
Woyke T, Teeling H, Ivanova NN, Huntemann M, Richter M, Gloeckner FO et al. (2006). Symbiosis insights through metagenomic analysis of a microbial consortium. Nature 443: 950–955.
Yakunin AF, Yee AA, Savchenko A, Edwards AM, Arrowsmith CH . (2004). Structural proteomics: a tool for genome annotation. Curr Opin Chem Biol 8: 42–48.
Acknowledgements
We thank Mr TW Arman, President, Iron Mountain Mines Inc., and Mr R Sugarek (US Environmental Protection Agency) for site access, and Mr R Carver and Mr D Dodds for on-site assistance. The contributions of many students, postdoctoral scientists, staff members and collaborators are gratefully acknowledged. The research was made possible by grant support from the US Department of Energy Genomics:GTL Program, the National Science Foundation LExEn and Biocomplexity Programs and the NASA Astrobiology Institute.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Denef, V., Mueller, R. & Banfield, J. AMD biofilms: using model communities to study microbial evolution and ecological complexity in nature. ISME J 4, 599–610 (2010). https://doi.org/10.1038/ismej.2009.158
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2009.158
Keywords
This article is cited by
-
Global trends and future prospects of acid mine drainage research
Environmental Science and Pollution Research (2023)
-
Functional Guilds, Community Assembly, and Co-occurrence Patterns of Fungi in Metalliferous Mine Tailings Ponds in Mainland China
Microbial Ecology (2023)
-
Mining of novel secondary metabolite biosynthetic gene clusters from acid mine drainage
Scientific Data (2022)
-
Microbial diversity in extreme environments
Nature Reviews Microbiology (2022)
-
Biofilm as an adaptation strategy to extreme conditions
Rendiconti Lincei. Scienze Fisiche e Naturali (2022)