Abstract
The exploration of the microbial world has been an exciting series of unanticipated discoveries despite being largely uninformed by rational estimates of the magnitude of task confronting us. However, in the long term, more structured surveys can be achieved by estimating the diversity of microbial communities and the effort required to describe them. The rates of recovery of new microbial taxa in very large samples suggest that many more taxa remain to be discovered in soils and the oceans. We apply a robust statistical method to large gene sequence libraries from these environments to estimate both diversity and the sequencing effort required to obtain a given fraction of that diversity. In the upper ocean, we predict some 1400 phylotypes, and a mere fivefold increase in shotgun reads could yield 90% of the metagenome, that is, all genes from all taxa. However, at deep ocean, hydrothermal vents and diversities in soils can be up to two orders of magnitude larger, and hundreds of times the current number of samples will be required just to obtain 90% of the taxonomic diversity based on 3% difference in 16S rDNA. Obtaining 90% of the metagenome will require tens of thousands of times the current sequencing effort. Although the definitive sequencing of hyperdiverse environments is not yet possible, we can, using taxa-abundance distributions, begin to plan and develop the required methods and strategies. This would initiate a new phase in the exploration of the microbial world.
Similar content being viewed by others
Introduction
The microbial world is vast, with 1030 organisms present on the Earth (Whitman et al., 1998), diverse (Curtis et al., 2002) and can only be observed through relatively tiny samples at discrete points in space and time (Sloan et al., 2007). Thus, it remains largely unexplored and is likely to remain so without quantitative statistical tools to estimate the magnitude of the task. Although very exciting and impressively large surveys are now being undertaken in marine (Huber et al., 2007; Rusch et al., 2007) and terrestrial environments (Roesch et al., 2007), none of these studies provides an exhaustive census of the microbial taxa in the samples. Sampling is still dictated by budgets and technologies and not an assessment of what is required to gain an authoritative picture of the diversity or to detect an organism of a known abundance. Without such an assessment, it is impossible to devise a rational strategy for the exploration of the microbial world, and until such a strategy is evinced, the systematic documentation of the microbial world will be impossible to plan and metagenomics studies will be conducted ‘blind’ (Curtis and Sloan, 2005). Taxa-abundance distributions (TADs) are central to this task (Curtis et al., 2002; Curtis and Sloan, 2005; Sloan et al., 2007), as estimates of richness alone do not give a realistic impression of the sequencing effort required to reveal the ‘unseen’ taxa (Curtis et al., 2006). However, lack of data has confounded the best efforts to home in on plausible distributions. The advent of high throughput sequencing technologies is changing this and it has now become possible to apply rigorous statistical methods to fit TADs to the data. This in turn permits us to determine the sequencing effort required to document a gene or all genes in a given environment.
Here, we apply a Bayesian approach, central to which is the definition of the likelihood, the probability of observing the data given the model parameters, in this case, the diversity and TAD (Chao and Bunge, 2002; Bunge et al., 2006; Hong et al., 2006). We looked at three data sets obtained through shotgun sequencing and 454 pyrosequencing that comprise an enormous number of sequence reads: the Global Ocean Survey (GOS) data from the upper oceans given by Rusch et al. (2007), the deep-sea vent data of Huber et al. (2007) and the soil data of Roesch et al. (2007). Our operational taxonomic units (OTUs) are based on differences in 16S rDNA sequences read either from a short variable region (Huber et al., 2007) or from longer shotgun sequences (Rusch et al., 2007). Taxa are defined as clusters of sequences that differ by at most 3% of sites, this approximates species (Konstantinidis and Tiedje, 2005; Hanage et al., 2006), and facilitates comparisons with other studies (Schloss and Handelsman, 2005; Huber et al., 2007). Our method yields the most informed estimates yet of diversity in these environments and applying the same method to the data sets facilitates comparison between them. In addition, we employ a novel technique to estimate not just the diversity in these systems but the degree of sampling required to catalogue that diversity.
Materials and methods
Data sets
Three data sets were used in this study, two sets of 16S rDNA tag sequences from pyrosequencing and a data set consisting of 16S cluster abundances from the GOS obtained through personal communication with Aaron Halpern (Rusch et al., 2007). The sequence data sets consisted of the deep-sea vent data of Huber et al. (2007), downloaded from the supplementary material, and the soil data of Roesch et al. (2007). See both papers for details of sample preparation, DNA amplification and pyrosequencing. The deep-sea vent data consist of tag sequences from two locations denoted by FS312 and FS396 separated into bacteria (FS312b and FS396b) and archaea (FS312a and FS396a). Because TADs can differ between communities and between bacteria and archaea, we treated each of these four samples separately.
The soil sequence data of Roesch et al. (2007) were obtained through personal communication with Eric W Triplett. These data derived from four different locations (Brazil, Florida, Illinois and Canada), and we treated each location as a separate sample. We only used bacterial sequences and did not consider the archaeal sequences separately because their sample sizes were relatively small (Illinois had the largest number of archaea samples, at 4530). To aid comparison with the deep-sea vent data and to reduce noise, we trimmed these sequences as described by Huber et al. (2007). Therefore, in total we analysed nine samples, four from each of the sequence data sets and the GOS data.
Generation of sample-abundance distributions
To reduce the size of the pyrosequencing samples, the unique tag sequences were identified and their frequencies were determined. The reduced samples were then aligned (Edgar, 2004), a distance matrix was calculated and the program dotur was used to cluster sequences (Sogin et al., 2006). OTUs were defined as the clusters at 97% sequence identity. This level of identity in 16S rRNA genes has been shown to be approximately equivalent to a 70% DNA–DNA reassociation value and produces OTUs that are reasonable proxies to species (Stackebrandt and Goebel, 1994; Konstantinidis and Tiedje, 2005; Hanage et al., 2006). This definition also allows comparison with other studies (Huber et al., 2007). The abundance of each taxa was then calculated as the sum of the constituent tag frequencies.
The GOS shotgun sequence data was already clustered when it was provided to us (Rusch et al., 2007). Assemblies were searched for conserved regions associated with 16S rDNA genes. These gene fragments were then aligned, distances generated in a similar fashion to above and then were clustered at 97% sequence identity. The contribution of each assembly to the cluster abundance was weighted by the proportion of the 16S gene present, and the average copy number of the assembly. This gave continuous weights for each cluster. To approximate number of 16S reads per cluster and thereby generate discrete taxa abundances, we divided these weights by half and rounded up to the nearest integer, a half being approximately the length of a typical read, 900 base pairs, divided by the length of the 16S gene, 1600 base pairs. For all nine samples, the taxa abundances were converted into sample-abundance distributions, that is, the number of taxa observed with a given abundance. Table 1 contains summary information on the nine microbial samples.
Definition of the likelihood
The first step in deriving the likelihood is to convert the continuous distribution of taxa abundances in the community into a discrete distribution of probabilities for the number of times an arbitrary taxon appears in the sample. We begin by denoting the normalized TAD by T(λ∣θ), where θ is a vector of parameters. We assume that each time an individual is sampled (with replacement), the probability that it is from a given taxon is equal to that taxon abundance, λ, divided by the total population number N. The number of times a taxon appears in the sample will then be approximately Poisson-distributed with mean λL/N, where L is the sample size. The unknown taxa abundances in the community will comprise a realization of independent samples from the TAD. We can integrate over these realizations to give a probability Pn that we will observe a taxon n times in the sample:

where μ=L/N is the sampling frequency (Pielou, 1969; Chao and Bunge, 2002; Etienne and Olff, 2005; Hong et al., 2006; Green and Plotkin, 2007). In general, μ will be hard to specify because of the difficulty in defining the number of individuals, N, that constitute a community. Fortunately, most abundance distributions are invariant under rescaling from community abundance λ to sample abundance λ → x=λ μ (Pielou, 1969). Therefore, we can write

where θ′ are rescaled parameters that will be a function of μ, hence we can fit for the parameters θ′ without knowing the value of μ. The discrete distribution Pn(θ ′) is a continuous mixture of Poisson distributions (Johnson et al., 2005).
If we let the richness, the total number of taxa in the community, be S, then each of these taxa has a probability Pn of appearing n times in the sample, including not appearing at all with probability P0. These probabilities are the same for all taxa, but this does not imply that each taxon has the same abundance in the community, rather that a priori all taxa are equivalent in the sense that their abundances are drawn from the same distribution. Therefore, the likelihood of observing the sample abundances has a multinomial distribution (Chao and Bunge, 2002; Hong et al., 2006). Let fi be the number of taxa observed with a given abundance i, and denote all these frequencies by the vector f=(f1,f2,…fL). The largest possible observed abundance is equal to the number of individuals in the sample L. The number of observed taxa D will be equal to ∑i=1Lfi so that there are f0=S−D unobserved taxa, then,

is the likelihood.
Calculating the discrete probability distribution in the sample
The expression for the probability that an arbitrary taxon will be represented by n individuals in the sample (Equation (2)) will depend on the TAD fitted, and consequently so will the diversity estimates and sampling efforts. To explore this, we used four different distributions: two two-parameter distributions, the log-normal and inverse Gaussian, and two three-parameter distributions, the log-Student's t and Sichel distributions. The log-normal has been frequently used to fit both microbial and macrobial abundance distributions (Pielou, 1969; Curtis et al., 2002). The log-Student's t distribution (or log-t distribution) is a generalization of this that has heavier tails but approaches log-normality as the ‘degrees of freedom’ parameter becomes infinite (Lange et al., 1989). The inverse Gaussian is a highly skewed distribution that has been applied previously to microbial abundances (Hong et al., 2006). The Sichel distribution is its three-parameter generalization (Sichel, 1974). Other distributions, for example, the exponential, gamma and a mixture of exponentials, were also tried, but they were a poor fit to the TADs. In the case of the inverse Gaussian and the Sichel distributions, the integral in Equation (2) can be performed analytically giving the probabilities Pn in terms of modified Bessel functions (Johnson et al., 2005). For the log-normal and log-Student's t distributions, no such closed expression is possible and instead numerical integration was used to calculate the Pn values.
Bayesian fitting to the sample-abundance distributions
To fit the sample abundances, we used a simple Bayesian approach. In Bayesian statistics, the ‘posterior distribution’ is the probability of the parameters given the data; it is proportional to the likelihood of the data multiplied by the prior probabilities of the parameters (Gelman et al., 2004). The likelihood of the sample abundances given the parameters of the underlying TAD is derived above. We used non-informative improper prior distributions. To sample from the posterior distribution, a Metropolis algorithm was used to perform Markov chain Monte-Carlo (Gilks et al., 1996). For each fit, three Markov chain Monte-Carlo runs from overdispersed starting parameters were performed and checked for convergence (Gelman, 1996). We found that a run length of 250 000 steps and a burn-in period of 100 000 was sufficient to ensure convergence. All results quoted in the paper are collated over the last 150 000 steps of all three runs. Table 2 gives the diversity estimates obtained from fitting the four TADs to the nine samples. These are calculated as the medians of the sampled S values together with 95% confidence intervals. This method of diversity estimation, which can be viewed as parametric, as it assumes a form for the TAD (Hong et al., 2006), infers the true diversity in the community together with confidence intervals, in contrast to non-parametric estimators such as those of Chao, which generate a lower bound (Chao, 1987; Hong et al., 2006; Sloan et al., 2008).
Model comparison
We used the deviance information criterion (DIC) to compare fits between models (Spiegelhalter et al., 2002). The DIC is defined as the sum of the deviance (−2 times the negative log-likelihood) averaged over the posterior distribution, d̄, and the effective number of parameters, pD. The deviance is related to the more familiar Chi-squared statistic of non-linear regression; a smaller deviance indicates a better fit. The pD term penalizes more complex models. For our non-hierarchical models, the DIC is simply d̄+3 for the two parameter abundance distributions, and d̄+4 for their three-parameter generalizations (the extra parameter is the diversity S). Models with smaller DIC values are preferred. When quoting fitted diversities in Table 2, we give the model ranking in terms of DIC, and we also highlight the best-fitting model and all those models that had DIC values within six of the best fit, except that we did not highlight any three parameter model that failed to decrease the DIC value by at least one over its nested two-parameter model. All highlighted models should be considered plausible candidates for fitting the data.
Calculating the 90% sampling effort
At a new sampling frequency, μ′=L′/N, the observed number of taxa will be drawn from a binomial distribution with mean (Chao and Bunge, 2002):

As stated above, we can use the transformation λ → x=λμ′ and express P0 as a function of the rescaled parameters, θ″. Remarkably, we can calculate these as a function of our original fitted parameters without knowledge of the community size. The exact procedure depends on the abundance distribution used. We will illustrate it for the log-normal distribution, which has two parameters M and V corresponding to the mean and variance of the log-transformed abundance, respectively. The variable V is unchanged under the transformation λ → x, only the mean changes M′=log(μ)+M, similarly at the new sampling frequency M″=log(μ′)+M. Therefore, simply combining and rearranging gives M″ as a function of the sample sizes and the fitted parameter, M″=log(L′/L)+M′. We applied similar procedures to all the fitted abundance distributions to determine the 90% sampling effort: the sample size with a mean observed diversity of 90% of the taxa present in the community. This method is conceptually similar to that of Schloss and Handelsman (2006), who generated artificial communities with abundance distributions similar to observed communities, and sampled from them. However, by avoiding the labour of generating and resampling large populations, we can generate sampling efforts from across the entire posterior distribution of fitted parameter values. In addition, we avoid the difficult problem of specifying the community size, N. For each microbial sample, and for each distribution, we calculated the sampling level expected to obtain 90% of the diversity (90% sampling effort) for 4500 sets of parameter values taken from the posterior distribution of fitted values. In Table 3, we list the medians of these sampling efforts together with 90% confidence intervals.
The proportion of the metagenome sequenced
In metagenomics, the aim is not to characterize the species present, but to obtain the aggregate genome or metagenome of the community through random reads distributed throughout the microbial genomes. Given a fitted abundance distribution, we can calculate the expected proportion of the metagenome obtained for a given number of randomly distributed reads. If there were only a single taxon present, then the expected proportion of the genome not sequenced (assuming reads of fixed length R base pairs randomly distributed throughout the genome) is an exponentially decreasing function of the coverage c=RL/G, where L is the read number and G is the genome size. In a community, assuming all taxa have the same genome length, then the number of reads from any given taxon will be weighted by its relative abundance, therefore the coverage becomes RLλ/GN≡Rμλ/G. Averaging over all taxa, this gives

for the expected fraction of the metagenome sequenced, with γ=G/R being the number of reads required to span the genome. Comparing this with Equation (1), we see that M=1−P0(μ/γ,θ). Thus, having calculated the expected sample size to capture 90% of the species diversity, we can multiply this by γ to give the expected sample size to accrue 90% of the genetic diversity.
Results and discussion
Table 1 contains the sample size in number of reads, the number of taxa observed, an estimate of the lower bound on richness obtained using Chao's estimator and estimates of diversity and 90% sampling effort using the best-fitting TAD for the GOS data (Rusch et al., 2007), for the four different soils samples (Roesch et al., 2007), and for the two deep-sea vent sites separated into bacteria and archaea (Huber et al., 2007). Figure 1a gives our estimates of the diversity of OTUs at these sites from the four different TADs, and Figure 1b gives our estimates of the sampling in reads required to observe 90% of the OTU diversity. The numerical values for these diversities and sampling efforts (in multiples of the current sample size) are given in Tables 2 and 3 together with 95% confidence intervals. Reassuringly, the lower bounds on our estimates for richness are higher than those given by Chao's estimator at all the sites and the median values of richness are significantly higher, irrespective of the distribution used. On the basis of the DIC values, it was easy to distinguish the best-fitting TADs for all data sets including the GOS where data were collated from multiple samples at different locations. Significantly, these distributions are a very good fit to the whole range of abundances (Figure 2); we did not need to right-truncate the sample to rare species as Hong et al. (2006) did when fitting to much smaller samples.
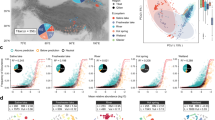
(a) Bayesian parametric diversity estimates from fits of abundance distributions to the samples summarized in Table 1. Estimates are given as medians with a 95% confidence interval (log-normal, red; inverse Gaussian, green; log-Student's t, blue; Sichel, yellow). These figures are given in Table 2. For each sample, the estimates are ordered according to the Bayesian DIC measure of fit (Spiegelhalter et al., 2002). The distributions that were significantly better than all others are highlighted in black, where two distributions fitted equally well both are highlighted. The Chao estimates (solid lines) and number of observed taxa (filled circles) are also shown. (b) The sampling effort (as 16S reads) necessary to sample 90% of taxa present (see main text). This is also given as medians and confidence intervals over the posterior distribution of fitted parameters. These figures are given in Table 3. Distributions are colour-coded as in panel (a), actual sample size (number of reads) is shown as a solid line.
Bayesian fits of abundance distributions to the nine samples summarized in Table 1. Diversity estimates from these fits are given in Table 1, and Table 2 with confidence intervals. Data points were aggregated to reduce noise such that the aggregate counts were at least 20. For each sample, only the best-fitting distribution is shown—identified on individual panels. Fits are the posterior average of predicted frequencies (SPn). Both axes have been scaled logarithmically.
Marine surface plankton
The planktonic bacterial communities in the upper ocean sampled in the GOS appear to be the least diverse. The best-fitting, Sichel distribution, estimates 1420 OTUs in the marine plankton biota, which is approximately 400 more taxa than estimated previously (Table 1). Thus, with 811 taxa defined so far, they are a little over half way through a complete census. However, doubling the sample size will be insufficient to identify the remaining taxa because they are rare and harder to find than the first 811. Again on the basis of the Sichel distribution, we estimate that to obtain 90% of the diversity in the 16S rRNA gene, approximately 20 000 partial or full genes would be required, which one might expect from 35 000 reads of that gene. Given that untargeted shotgun sequencing is being applied to the entire metagenome in the GOS, approximately 30 000 000 reads would be required to achieve this and simultaneously reveal 90% of the total genetic diversity. This corresponds to approximately 5 times the current sequencing effort (Table 1), which is eminently achievable. However, if the aim is to assemble complete genomes, which requires larger levels of coverage to obtain overlapping sequences, then many more reads than this will be necessary.
Soil
Until recently, soil environments were regarded as being the most diverse (Torsvik et al., 2002). Some controversial (Bunge et al., 2006) estimates of diversity exceeded 106 different prokaryotes per gram of soil (Gans et al., 2005). Soils are a prime example of an environment where estimates of diversity have been compromised by undersampling. Therefore, very large samples of Roesch et al. (2007) are particularly welcome. They estimated total diversity using a portion of the 16S rRNA gene and the sampling effort required to recover 90% of that diversity. They asserted that soil samples can be easily characterized using pyrosequencing; for a soil sample from Brazil, they estimate that the maximum diversity is 5021 OTUs and that to capture 90% of that diversity it would require a modest 226 388 reads. However, their non-parametric estimates of diversity are known to be conservative and their alternative method of extrapolating a Michaelis-Menten (M-M) curve, fitted by non-linear regression to the mean diversity estimates obtained from sub-sampling is even more conservative (Gotelli and Colwell, 2001). In Figure 3, we plot the observed rarefaction curve for the Brazilian soil data along with that of the best-fitting M-M curve and those for our fitted TADs. The M-M fit is clearly unsatisfactory; it does not fit the expected sub-sampled diversities as well as the abundance distributions, and it extrapolates to an asymptotic level of diversity (3375), which is smaller than the lower bound given by the Chao estimator. Given the statistically robust nature of the Chao estimator's lower bounds, this strongly suggests that even the more sophisticated two-part M-M curves used by Roesch et al. (2007) are inferior to the non-parametric estimators. In contrast, our parametric estimates of soil diversity are significantly higher (Table 1). It is clear from Figure 1b and Table 3 that substantial extra sampling will be necessary to characterize these communities. In particular, the diversity in the Canadian soil sample is large; our predictions range from 20 000 to 140 000 taxa (Table 2). The best-fitting Sichel distribution predicts that a median of 551 sample sizes or just over 29 million reads will be required to obtain 90% of these taxa, which contrasts with the equivalent figure of 713 000 reads obtained by Roesch et al. (2007) from extrapolation of the rarefaction curve. Our figure would require at least 70 runs of a Roche FLX genome sequencer, a considerable effort in terms of time and money. In addition, the upper limit of our prediction is ten times this value. It is unlikely that all soil communities can be easily characterized by current pyrosequencing technologies (Roesch et al., 2007). However, we know now the performance that would be required to attain this goal. We anticipate that the systematic and authoritative sequencing of the soil will enable us to see patterns obscured previously by inadequate sampling. It has, for example, not escaped our attention that the diversity in this soil data set apparently increases from south to north.
Estimated rarefaction curves for the Brazil soil data set of Roesch et al. (2007). The main figure shows the mean observed diversity as a function of read number obtained by applying Equation (3) to the log-normal (dashed line) and inverse Gaussian (dot-dash) fits. Results are averaged over the posterior distributions of the fitted parameters. A least-squares Michaelis-Menten (M-M) fit (dotted) to the diversities obtained by sub-sampling is also shown (Gotelli and Colwell, 2001). The M-M fit has an asymptotic diversity of 3775.4 and half that maximum diversity will be observed at 9305.9 reads. The inset shows the same curves together with the mean expected diversity from sub-sampling (solid). The diversities from sub-sampling and fitting the abundance distributions coincide almost exactly.
Deep-sea vents
In 2006, Sogin et al. showed that in samples from deep-sea diffuse hydrothermal vents, there was a surprisingly large phylogenetic diversity in the rare bacterial taxa. Huber et al. (2007) returned to the same vents in an attempt to further define the extent of microbial diversity and to fully resolve the archaeal and bacterial communities. The number of 16S RNA tag sequences that they obtained is an order of magnitude higher than for any other environment. However, on the basis of their non-parametric diversity estimates and rarefaction curves, they conclude that yet further sampling is required. It can be seen from Figure 1 and Tables 2 and 3 that our estimates of diversity and the sampling effort to accrue 90% of the taxa in these deep-sea vents depends strongly on the TAD assumed, especially at the FS396 site where the sample size is lower than the FS312 site. This reinforces the conclusion of Huber et al. that the environments are still undersampled and that to be absolutely certain of the underlying TAD will require an increase in sampling frequency. However, our best-fitting distributions, the log-normal and its generalization, the log-t distribution, are significantly better models of the data than the inverse-Gaussian or Sichel distributions, which gives us more confidence in predictions made on their basis. Thus, our best estimates are that there are approximately 50 000 bacterial taxa at FS312 and 300 000 bacterial taxa at FS396. This would mean that even at the better-sampled FS312 site sample sizes would need to increase 280 times, requiring 120 million reads, just to obtain 90% of the diversity in the 16S rRNA tag sequences. Even for the relatively taxa-poor archaea, sampling levels at this site would need to increase 70-fold to obtain 90% of the tag sequence diversity. Suppose that a metagenomics data set was to be compiled from 1000 base pair random insert clone libraries for bacteria at the FS312 deep-sea vent. To obtain 90% of the bacterial genetic diversity, 1011 reads would be required, assuming a conservative marine bacterial genome length of just 1 000 000 base pairs (Giovannoni et al., 2005). For assembly, this figure is a lower limit that assumes that assembly is possible with infinitesimal overlap, but even for this to be practical, new, even more powerful, sequencing technologies would be required.
Further applications
In this study, we have focused on bacteria and archaea; however, our approach has wider applicability. High-throughput sequencing technologies could, and no doubt soon will, be used to investigate the diversity of eukaryotic microbes such as fungi, protists and microalgae in environmental samples through amplification and sequencing of hypervariable regions of ribosomal genes (Montero et al., 2008). Virus genomes lack conserved regions; consequently, metagenomics is necessary to identify the types present (Angly et al., 2006; Fierer et al., 2007). In either case, the potential exists for large data sets to be collected, and our method for predicting true diversity and sampling efforts could be used to inform that collection. Indeed, our statistical methods can be applied to any community for which a sample-abundance distribution is available, regardless of the origin of that data, for instance, clone libraries could be used, although their typically small sizes will result in uncertain estimates. In fact, we used the Barro Colorado Island tropical tree data set to validate our methods (Hubbell et al., 1999). This provided a completely characterized community of over 200 000 individuals, from which we randomly sub-sampled much smaller numbers (1000) to mimic low sampling frequencies. From these sub-samples, we estimated the total community diversity and 90% sampling effort, and the true values of both were within the 95% confidence intervals of our predictions. This illustrates the robustness and generality of our method.
Concluding remarks
Access to the results of high-throughput sequencing data has allowed us to make the best informed estimates of diversity to date for very diverse environments such as soils and deep-sea vents and the sequencing efforts required to uncover them. These results are striking but ultimately less significant than the methods, mathematical and molecular from which they were derived. For the methods evinced in this article, transform the intriguing observations made using high-throughput sequencing into testable hypotheses about the distribution and extent of the microbial diversity in these environments.
It is imperative that we now determine whether we can or cannot predict the diversity and TAD of an environment using a conjunction of mathematics and the new generation of sequencing technology. If we can, then studies of microbial diversity can move into a new phase in which the estimation and description of microbial diversity becomes a rational and planned activity. The systematic mapping of the extent of the diversity of the microbial world can become a reality and its systematic exploration can become plausible. The clearer picture this new approach will offer us will be a foundation for a more sophisticated and predictive understanding of real microbial communities.
We have called previously for a microbial survey analogous to a geological survey (Curtis, 2006), reasoning that microbes will have at least as much, if not more, impact on our environment and society in the next century as geology has had in the past 200 years. The advent of novel sequencing technologies and adequate mathematical tools make this proposal a tangible and fundable reality.
References
Angly FE, Felts B, Breitbart M, Salamon P, Edwards RA, Carlson C et al. (2006). The marine viromes of four oceanic regions. PLoS Biol 4: 2121–2131.
Bunge J, Epstein SS, Peterson DG . (2006). Comment on ‘Computational improvements reveal great bacterial diversity and high metal toxicity in soil’. Science 313: 918.
Chao A . (1987). Estimating the population-size for capture recapture data with unequal catchability. Biometrics 43: 783–791.
Chao A, Bunge J . (2002). Estimating the number of species in a Stochastic abundance model. Biometrics 58: 531–539.
Curtis TP . (2006). Microbial ecologists: it's time to ‘go large’. Nat Rev Microb 4: 488.
Curtis TP, Head IM, Lunn M, Woodcock S, Schloss PD, Sloan WT . (2006). What is the extent of prokaryotic diversity? Philos T Roy Soc B 361: 2023–2037.
Curtis TP, Sloan WT . (2005). Exploring microbial diversity—a vast below. Science 309: 1331–1333.
Curtis TP, Sloan WT, Scannell JW . (2002). Estimating prokaryotic diversity and its limits. Proc Natl Acad Sci USA 99: 10494–10499.
Edgar RC . (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797.
Etienne RS, Olff H . (2005). Confronting different models of community structure to species-abundance data: a Bayesian model comparison. Ecol Lett 8: 493–504.
Fierer N, Breitbart M, Nulton J, Salamon P, Lozupone C, Jones R et al. (2007). Metagenomic and small-subunit rRNA analyses reveal the genetic diversity of bacteria, archaea, fungi, and viruses in soil. Appl Environ Microbiol 73: 7059–7066.
Gans J, Wolinsky M, Dunbar J . (2005). Computational improvements reveal great bacterial diversity and high metal toxicity in soil. Science 309: 1387–1390.
Gelman A . (1996). Inference and monitoring convergence. In: Gilks WR, Richardson S, Spiegelhalter DJ (eds). Markov chain Monte Carlo in practice. Chapman & Hall: London, UK, pp 131–143.
Gelman A, Carlin JB, Stern HS, Rubin DB . (2004). Bayesian Data Analysis. Chapman & Hall: London, UK.
Gilks WR, Richardson S, Spiegelhalter DJ . (1996). Introducing Markov chain Monte Carlo. In: Gilks WR, Richardson S, Spiegelhalter DJ (eds). Markov Chain Monte Carlo in Practice. Chapman & Hall: London, UK, pp 1–20.
Giovannoni SJ, Tripp HJ, Givan S, Podar M, Vergin KL, Baptista D et al. (2005). Genome streamlining in a cosmopolitan oceanic bacterium. Science 309: 1242–1245.
Gotelli NJ, Colwell RK . (2001). Quantifying biodiversity: procedures and pitfalls in the measurement and comparison of species richness. Ecol Lett 4: 379–391.
Green JL, Plotkin JB . (2007). A statistical theory for sampling species abundances. Ecol Lett 10: 1037–1045.
Hanage WP, Fraser C, Spratt BG . (2006). Sequences, sequence clusters and bacterial species. Philos T Roy Soc B 361: 1917–1927.
Hong SH, Bunge J, Jeon SO, Epstein SS . (2006). Predicting microbial species richness. Proc Natl Acad Sci USA 103: 117–122.
Huber JA, Mark Welch D, Morrison HG, Huse SM, Neal PR, Butterfield DA et al. (2007). Microbial population structures in the deep marine biosphere. Science 318: 97–100.
Hubbell SP, Foster RB, O'Brien ST, Harms KE, Condit R, Wechsler B et al. (1999). Light-gap disturbances, recruitment limitation, and tree diversity in a neotropical forest. Science 283: 554–557.
Huse SM, Huber JA, Morrison HG, Sogin ML, Mark Welch D . (2007). Accuracy and quality of massively parallel DNA pyrosequencing. Genome Biol 8: R143.
Johnson NL, Kemp AW, Kotz S . (2005). Univariate Discrete Distributions. John Wiley & Sons: Hoboken, New Jersey.
Konstantinidis KT, Tiedje KM . (2005). Genomic insights that advance the species definition for prokaryotes. Proc Natl Acad Sci USA 102: 2567–2572.
Lange KL, Little RJA, Taylor JMG . (1989). Robust statistical modeling using the t-distribution. J Am Stat Assoc 84: 881–896.
Montero CI, Shea YR, Jones PA, Harrington SM, Tooke NE, Witebsky FG et al. (2008). Evaluation of Pyrosequencing® technology for the identification of clinically relevant non-dematiaceous yeasts and related species. Eur J Clin Microbiol Infect Dis; Online First DOI 10.1007/s10096 008–01510-x.
Pielou EC . (1969). An Introduction to Mathematical Ecology. Wiley: New York, NY.
Roesch LF, Fulthorpe RR, Riva A, Casella G, Hadwin AKM, Kent AD et al. (2007). Pyrosequencing enumerates and contrasts soil microbial diversity. ISME J 1: 283–290.
Rusch DB, Halpern AL, Sutton G, Heidelberg KB, Williamson S, Yooseph S et al. (2007). The sorcerer II global ocean sampling expedition: northwest Atlantic through eastern tropical Pacific. PLoS Biol 5: 398–431.
Schloss PD, Handelsman J . (2005). Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microb 71: 1501–1506.
Schloss PD, Handelsman J . (2006). Toward a census of bacteria in soil. PLoS Comp Biol 2: 786–793.
Sichel HS . (1974). Distribution representing sentence-length in written prose. J Roy Stat Soc A 137: 25–34.
Sloan WT, Quince C, Curtis TP . (2008). The Uncountables. In: Zengler K (ed). Accessing Uncultivated Microorganisms: from the Environment to Organisms and Genomes and Back. ASM Press: Washington, DC, pp 35–54.
Sloan WT, Woodcock S, Lunn M, Head IM, Curtis TP . (2007). Modeling taxa-abundance distributions in microbial communities using environmental sequence data. Microbial Ecol 53: 443–455.
Sogin ML, Morrison HG, Huber JA, Mark Welch D, Huse SM, Neal PR et al. (2006). Microbial diversity in the deep sea and the underexplored ‘rare biosphere’. Proc Natl Acad Sci USA 103: 12115–12120.
Spiegelhalter DJ, Best NG, Carlin BR, van der Linde A . (2002). Bayesian measures of model complexity and fit. J Roy Stat Soc B 64: 583–616.
Stackebrandt E, Goebel BM . (1994). A place for DNA-DNA reassociation and 16S ribosomal-RNA sequence analysis in the present species definition in bacteriology. Int J Syst Bacteriol 44: 846–849.
Torsvik V, Ovreas L, Thingstad TF . (2002). Prokaryotic diversity—magnitude, dynamics, and controlling factors. Science 296: 1064–1066.
Whitman WB, Coleman DC, Wiebe WJ . (1998). Prokaryotes: the unseen majority. Proc Natl Acad Sci USA 95: 6578–6583.
Acknowledgements
We thank Aaron Halpern, Alberto Riva, Doug Rusch and Eric W Triplett for providing the soil sequence and GOS cluster data sets. We also thank Sue Huse and Mitchell Sogin for providing a copy of the quickdist program, and Rampal S Etienne for an algorithm used in fitting the log-normal distribution. We thank Mark Bailey, Stephen Giovannoni, and two anonymous reviewers for helpful comments on an earlier version of this article. Chris Quince is supported by a Lord Kelvin Adam Smith Research Fellowship from the University of Glasgow, and Bill Sloan is supported by an Engineering and Physical Sciences Advanced Research Fellowship.
The Bayesian diversity estimation software used in this study is available from the corresponding author upon request or can be downloaded from the website: http://people.civil.gla.ac.uk/~quince/Software/BDES.html.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Quince, C., Curtis, T. & Sloan, W. The rational exploration of microbial diversity. ISME J 2, 997–1006 (2008). https://doi.org/10.1038/ismej.2008.69
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2008.69
Keywords
This article is cited by
-
Direct quantification of ecological drift at the population level in synthetic bacterial communities
The ISME Journal (2021)
-
Clade-specific diversification dynamics of marine diatoms since the Jurassic
Nature Ecology & Evolution (2018)
-
Abundance determines the functional role of bacterial phylotypes in complex communities
Nature Microbiology (2018)
-
Hill number as a bacterial diversity measure framework with high-throughput sequence data
Scientific Reports (2016)
-
The activated sludge ecosystem contains a core community of abundant organisms
The ISME Journal (2016)