Abstract
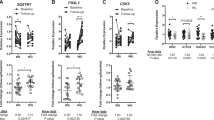
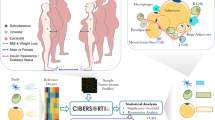
Obese subjects with a similar body mass index (BMI) exhibit substantial heterogeneity in gluco- and cardiometabolic heath phenotypes. However, defining genes that underlie the heterogeneity of metabolic features among obese individuals and determining metabolically healthy and unhealthy phenotypes remain challenging. We conducted unsupervised hierarchical clustering analysis of subcutaneous adipose tissue transcripts from 30 obese men and women ⩾40 years old. Despite similar BMIs in all subjects, we found two distinct subgroups, one metabolically healthy (group 1) and one metabolically unhealthy (group 2). Subjects in group 2 showed significantly higher total cholesterol (P=0.005), low-density lipoprotein cholesterol (P=0.006), 2-h insulin during oral glucose tolerance test (P=0.015) and lower insulin sensitivity (SI, P=0.029) compared with group 1. We identified significant upregulation of 141 genes (for example, MMP9 and SPP1) and downregulation of 17 genes (for example, NDRG4 and GINS3) in group 2 subjects. Intriguingly, these differentially expressed transcripts were enriched for genes involved in cardiovascular disease-related processes (P=2.81 × 10−11–3.74 × 10−02) and pathways involved in immune and inflammatory response (P=8.32 × 10−5–0.04). Two downregulated genes, NDRG4 and GINS3, have been located in a genomic interval associated with cardiac repolarization in published GWASs and zebra fish knockout models. Our study provides evidence that perturbations in the adipose tissue gene expression network are important in defining metabolic health in obese subjects.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Ahima RS, Lazar MA . Physiology. The health risk of obesity—better metrics imperative. Science 2013; 341: 856–858.
Bluher M . The distinction of metabolically 'healthy' from 'unhealthy' obese individuals. Curr Opin Lipidol 2010; 21: 38–43.
Phillips CM . Metabolically healthy obesity: definitions, determinants and clinical implications. Rev Endocr Metab Disord 2013; 14: 219–227.
Tchkonia T, Thomou T, Zhu Y, Karagiannides I, Pothoulakis C, Jensen MD et al. Mechanisms and metabolic implications of regional differences among fat depots. Cell Metab 2013; 17: 644–656.
Jo J, Gavrilova O, Pack S, Jou W, Mullen S, Sumner AE et al. Hypertrophy and/or hyperplasia: dynamics of adipose tissue growth. PLoS Comput Biol 2009; 5: e1000324.
Despres JP, Lemieux I . Abdominal obesity and metabolic syndrome. Nature 2006; 444: 881–887.
Van Gaal LF, Mertens IL, De Block CE . Mechanisms linking obesity with cardiovascular disease. Nature 2006; 444: 875–880.
Das SK, Sharma NK, Hasstedt SJ, Mondal AK, Ma L, Langberg KA et al. An integrative genomics approach identifies activation of thioredoxin/thioredoxin reductase-1-mediated oxidative stress defense pathway and inhibition of angiogenesis in obese nondiabetic human subjects. J Clin Endocrinol Metab 2011; 96: E1308–E1313.
Elbein SC, Kern PA, Rasouli N, Yao-Borengasser A, Sharma NK, Das SK . Global gene expression profiles of subcutaneous adipose and muscle from glucose-tolerant, insulin-sensitive, and insulin-resistant individuals matched for BMI. Diabetes 2011; 60: 1019–1029.
Keller MP, Attie AD . Physiological insights gained from gene expression analysis in obesity and diabetes. Annu Rev Nutr 2010; 30: 341–364.
Qatanani M, Tan Y, Dobrin R, Greenawalt DM, Hu G, Zhao W et al. Inverse regulation of inflammation and mitochondrial function in adipose tissue defines extreme insulin sensitivity in morbidly obese patients. Diabetes 2013; 62: 855–863.
Sharma NK, Langberg KA, Mondal AK, Das SK . Phospholipid biosynthesis genes and susceptibility to obesity: analysis of expression and polymorphisms. PLoS One 2013; 8: e65303.
Caraux G, Pinloche S . PermutMatrix: a graphical environment to arrange gene expression profiles in optimal linear order. Bioinformatics 2005; 21: 1280–1281.
Tusher VG, Tibshirani R, Chu G . Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA 2001; 98: 5116–5121.
Huang dW, Sherman BT, Lempicki RA . Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009; 4: 44–57.
Guilherme A, Virbasius JV, Puri V, Czech MP . Adipocyte dysfunctions linking obesity to insulin resistance and type 2 diabetes. Nat Rev Mol Cell Biol 2008; 9: 367–377.
Newton-Cheh C, Eijgelsheim M, Rice KM, de Bakker PI, Yin X, Estrada K et al. Common variants at ten loci influence QT interval duration in the QTGEN Study. Nat Genet 2009; 41: 399–406.
Pfeufer A, Sanna S, Arking DE, Muller M, Gateva V, Fuchsberger C et al. Common variants at ten loci modulate the QT interval duration in the QTSCD Study. Nat Genet 2009; 41: 407–414.
Qu X, Jia H, Garrity DM, Tompkins K, Batts L, Appel B et al. Ndrg4 is required for normal myocyte proliferation during early cardiac development in zebrafish. Dev Biol 2008; 317: 486–496.
Milan DJ, Kim AM, Winterfield JR, Jones IL, Pfeufer A, Sanna S et al. Drug-sensitized zebrafish screen identifies multiple genes, including GINS3, as regulators of myocardial repolarization. Circulation 2009; 120: 553–559.
Acknowledgements
This work was supported by grant R01 DK039311 and R01 DK090111 from the National Institutes of Health/NIDDK. We thank the Clinical Research Center staff of University of Arkansas for Medical Sciences for their outstanding support in the physiologic studies and assistance with data management. We thank Prof Siqun Zheng, Director, and the technical staff of the Center for Human Genomics, Wake Forest School of Medicine, especially Ms Shelly Smith and Dr Ge Li for their extensive support in gene expression analysis. We also thank Ethel Kouba (Internal Medicine, Endocrinology) and Karen Klein (Biomedical Research Services and Administration) for critical reading and editing of our manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on International Journal of Obesity website
Supplementary information
Rights and permissions
About this article
Cite this article
Das, S., Ma, L. & Sharma, N. Adipose tissue gene expression and metabolic health of obese adults. Int J Obes 39, 869–873 (2015). https://doi.org/10.1038/ijo.2014.210
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ijo.2014.210
This article is cited by
-
Common and ethnic-specific derangements in skeletal muscle transcriptome associated with obesity
International Journal of Obesity (2024)
-
Exploratory integrated analysis of circulating exosomal miRNA and tissue mRNA related to long-term physical activity for more than 25 years: a bioinformatics study
European Journal of Applied Physiology (2023)
-
Transcriptional, epigenetic and metabolic signatures in cardiometabolic syndrome defined by extreme phenotypes
Clinical Epigenetics (2022)
-
Genetic background of obesity
Pediatric Research (2021)
-
Distinct whole-blood transcriptome profile of children with metabolic healthy overweight/obesity compared to metabolic unhealthy overweight/obesity
Pediatric Research (2021)