Abstract
Plant breeding, one of the oldest agricultural activities, parallels human civilization. Many crops have been domesticated to satisfy human's food and aesthetical needs, including numerous specialty horticultural crops such as fruits, vegetables, ornamental flowers, shrubs, and trees. Crop varieties originated through selection during early human civilization. Other technologies, such as various forms of hybridization, mutation, and transgenics, have also been invented and applied to crop breeding over the past centuries. The progress made in these breeding technologies, especially the modern biotechnology-based breeding technologies, has had a great impact on crop breeding as well as on our lives. Here, we first review the developmental process and applications of these technologies in horticultural crop breeding. Then, we mainly describe the principles of the latest genome-editing technologies and discuss their potential applications in the genetic improvement of horticultural crops. The advantages and challenges of genome-editing technologies in horticultural crop breeding are also discussed.
Similar content being viewed by others
Introduction
Plant breeding is one of the oldest agricultural activities, and it parallels human civilization. Human society started to progress from hunting and gathering to agriculture approximately 11 000 years ago,1 which represents a shift from mobile collection to settled production. Since then, efforts to produce more food for survival have never stopped. By the end of the 18th century, more than 1000 species of plants had been domesticated around the world, of which approximately 100–200 now constitute the major components of the human diet, such as rice, wheat, maize, potato, yam, coconut, banana, etc.1 Interestingly, the majority of these domesticated crops by numbers are horticultural crops, which have been domesticated for satisfying human's special diet, medical needs, and aesthetic purposes. To obtain higher yield and better quality varieties, many approaches have been used in crop breeding, such as various forms of hybridization breeding, mutation breeding, and transgenic breeding. Through the application of these newer technologies, many new crop varieties with novel traits have been generated. Here, we briefly summarize the development of breeding technologies in horticultural crops, we then introduce the latest genome-editing technologies, and, finally, we discuss their potential applications and challenges in breeding horticultural crops.
Principles and applications of traditional breeding technologies in horticultural crop breeding
Hybridization breeding technology
Hybridization breeding has long been used by humankind. In early days, naturally hybridized individuals with desirable traits such as larger fruit/nut, better taste, or higher yield were intentionally selected and preserved. Later, people observed the differences between the male and female reproductive organs of plants and learned that new offspring with superior traits could be generated by artificial mating, or crosspollination. This marked the emergence of plant hybridization breeding, a hallmark of modern agriculture/horticulture. Through intentional hybridization, breeders could combine useful traits from two or more sources in one individual plant in one or more generations. One of the most successful applications of hybridization breeding is the utilization of heterosis, a phenomenon in which a hybrid (F1) progeny is typically superior with respect to size, growth characteristics, and yield compared with either parents.2 Many fruit and vegetable crops are generated by hybridization and selection, such as garden strawberry (Fragaria × ananassa), apple (Malus × domestica), sweet orange (Citrus sinensis), tomato (Solanum lycopersicum), and squash (Cucurbita maxima). Another application of hybridization breeding is the generation of seedless horticultural crops, such as watermelon, by employing diploid and tetraploid parents. However, crop hybridization breeding has limitations that are difficult to overcome. First, hybridization can only be successfully conducted between two compatible plants in the same or closely related species/genus. Second, when plants are hybridized, many favorable traits of interest are transferred along with undesirable traits such as poor quality or low yield potential.3 Due to the tight linkage of these genes, it may take several generations to segregate the undesired genes by back-crossing. Third, but not least, the breeding of many woody horticultural crops such as apple and walnut can take as many as 20–30 years to assort several favorable traits together in a single individual. This requires enormous amounts of labor and land resources, although fast track breeding techniques4 and molecular marker-assisted selection5 may accelerate breeding and selection processes.
Mutation breeding technology
During the process of crop evolution, spontaneous variations with new characteristics sometimes occurred and these have been preserved. Utilization of these variations in crop breeding, such as the semi-dwarf variation of cereal crops, has greatly improved grain yield; this is known as the “green revolution.”6 New cultivars selected from spontaneous mutations are particularly productive in perennial horticultural crops, such as the new red-skinned Fuji apple,7 large-berry tetraploid Kyoho grape,8 and many unique-looking ornamentals. This method of taking advantage of natural gifts is still widely utilized today. However, this method has at least three shortcomings. First, many of these types of mutations are not inheritable and may frequently reverse back to the original phenotypes. Second, many of these mutations give rise to chimeras and require “purification,” frequently through the use of successive grafting. Third, there is a low frequency of these chance mutations in nature.9
The low frequency of natural mutation can be overcome by artificially exposing various plant materials, such as seeds, cuttings, pollen, or tissue cultured calli, to either physical or chemical mutagens. This method was first discovered in the early 20th century when plant biologists found that exposing seeds to certain chemical compounds or radioactive rays could increase the frequency of genetic variations. This discovery later brought about plant mutation breeding. Though little was known about the molecular or genetic basis at that time, mutagens were rapidly applied to plant breeding, and a wide variety of genetic variations have been induced in most of the economically important crop species, including agricultural crops, fruit crops, and ornamental flowers, etc.10 However, despite this considerable increase in the frequency of mutations, mutation is a random, non-specific process, and the majority of mutations are deleterious and chimeric. Thus, obtaining varieties with a desirable trait requires a large population, efficient screening, and frequent follow-up purification of chimeras.
Transgenic technology in horticultural crop breeding
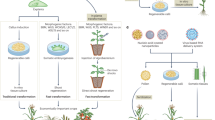
The gene is the basis of plant breeding. During early stages, breeders selected new phenotypes with meritorious traits without knowing the genotype. The advent of molecular genetics has paved a wide road for modern biotechnological breeding. By knowing the details how desirable/undesirable traits are inherited and genetically controlled, molecular biologists can precisely manipulate the gene encoding a trait to create novel phenotypes through DNA recombinant technologies. Known as transgenic technology, the beauty of this breeding technology is that it can transfer the cloned gene regardless of the source or recipient of the genes. The key step in transgenic technology is the integration of desired foreign genes into the host plant genome. At present, there are three main plant transformation methods, i.e., the Agrobacterium-mediated method, particle bombardment method, and protoplast transformation method. The Agrobacterium-mediated method is the easiest and most convenient method. However, many horticulture crops are not susceptible to Agrobacterium and therefore cannot be transformed using this method. The bombardment and protoplast methods can overcome the host-dependent shortage of the Agrobacterium-mediated method. However, these methods also have shortcomings, e.g., the bombardment method requires specific facilities and the protoplast method is dependent on handling skills.11 Nevertheless, protoplast transformation and the following regeneration of a whole plant from a single cell is an alternative strategy for many horticulture crops. Through transgenic approaches, breeders can introduce genes encoding for new traits into plants, such as resistance to certain pests or diseases, even with the genes from viral, bacterial, or distant plant species. The primary advantages are its precision and the improvement of a trait without altering the genetic constitution of an elite genotype. This is particularly useful for many perennial horticultural crops because they are highly heterozygous.
The first field trials of genetically engineered plants were performed in France and the USA in 1986.12 FlavrSavr tomato was the first transgenic food that was approved for sale in the USA in 1994.13 A glycoside hydrolase gene that encodes polygalacturonase, which can dissolve cell-wall pectin in the plant cell wall, is silenced in the FlavrSavr tomato, resulting in slower softening and decay. This characteristic permits the transgenic tomato to be picked at a later stage of maturity with a greater development of flavor compounds and therefore better taste.14 Different from FlavrSavr tomato, another successful transgenic story in horticultural crops is the viral disease resistant papaya.15 In addition to tomato and papaya, many horticultural crop varieties have been generated using transgenic technology and have been released (Table 1). Although transgenic technology has achieved great success in supplementing crop breeding and has considerable commercial value, this technology faces some technical challenges. For instance, there are many economically important plant species, or elite varieties of particular species, that remain highly recalcitrant to genetic transformation and regeneration.3 In addition to the technical obstacles, transgenic technology has faced increasing opposition in recent years because of the likely unpredictable risks to the environment and food safety, even though many of these claims are baseless. However, more useful technologies have been developed to address these concerns.16,17
Genome-editing technologies
In the past decade, new technologies commonly referred to as genome-editing technologies have emerged. These technologies rely on engineered endonucleases (EENs) that cleave DNA in a sequence-specific manner because of the presence of a sequence-specific DNA-binding domain or RNA sequence.18,19 Through recognition of the specific DNA sequence, these nucleases can efficiently and precisely cleave the targeted genes. The double-strand breaks (DSBs) of DNA consequently result in cellular DNA repair mechanisms, including homology-directed repair (HDR) and error-prone non-homologous end joining breaks (NHEJ),20 leading to gene modification at the target sites.
Zinc finger nucleases (ZFNs) in gene modification
ZFNs are the first generation EENs that were developed following the discovery of the functional principles of the Cys2-His2 zinc finger (ZF) domains.21 Each Cys2-His2 ZF domain consists of 30 amino-acid residues that fold into a βββ configuration.21,22 Crystal structure analysis indicates that Cys2-His2 ZF proteins bind to DNA by inserting the α-helix into the major groove of the double helix.23 Each ZF protein is able to recognize 3 contiguous nucleotide bases within the DNA substrate. As shown in Figure 1, a generic ZFN monomer is fused by two functional distinct domains: an artificially prepared Cys2-His2 ZF domain at the N-terminal and a nonspecific DNA cleavage domain of the Fok I DNA restriction enzyme at the C-terminal. The dimerization of the Fok I domain is crucial for its enzymatic activity.24 Therefore, a ZFN dimer composed of two 3- or 4-ZF domains will recognize an 18- or 24-base target sequence that, statistically, forms a unique site in the genomes of most organisms. Since the first report in 1996, ZFNs have been successfully applied to gene modification mainly in animals such as human cells,25 zebrafish,26,27 and plants such as Arabidopsis,28 tobacco,29 and maize.30 However, obtaining functional ZFNs requires an extensive and time-consuming screening process.31 Further, ZFNs have other limitations, such as off-target effect22 or even toxic to the host cells. These shortcomings limit the application of ZFNs in plant genome editing. Until now, there have been no reports on ZFN applications in horticultural crops.
Schematic illustration of the ZFN structure and the principle of ZFN-mediated genomic modifications. The target site of the ZFN is recognized by the “left” and “right” half monomer that each consist of a tandem array of engineered ZFPs, and each engineered ZFP can recognize a nucleotide triplet (shown in different colors). The ZFN monomer is comprised of an N-terminal domain containing a NLS (red), a recognition domain that usually comprises tandem ZFPs (in different colors) and a C-terminal function domain that comprises the Fok I endonuclease. Recognition of the target sequence by the left and right ZFPs results in dimerization of the Fok I endonuclease, which is critical for the activity of the ZFNs. DNA cleavage takes place between the two ZFP recognition sites that contain a spacer sequence that is usually 6 bp long. Induced DSB of the target DNA are repaired either by NHEJ or HDR, resulting in gene mutation around the cleavage sites. NLS, nuclear localization signal; ZFP, zinc finger proteins; DSB, double-strand breaks; NHEJ, non-homologous end joining; HDR, homology-directed repair. Mutation#x002A;, the red color box region contains nucleotide deletion, insertion or substitution. Figure modified from Gaj et al. (2013), Figure 119 and Moore et al. (2012), Figure 1.100
TALENs in gene modification
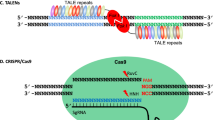
Recently, a new EEN, i.e., transcription activator-like effector nucleases (TALENs), has rapidly emerged as an alternative to ZFNs for genome editing.32 The broad applications of TALENs were based on the recognition of the functional principles of the type III transcription activator-like (TAL) effectors that are secreted by the plant pathogenic bacteria Xanthomonas.33 After been pumped into host cells, the TAL effectors enter the nucleus and bind to effector-specific sequences in the host gene promoters and activate transcription.34 AvrBs3 was the first investigated TAL effector protein found in 1989.35 However, the recognition mechanism of the TAL effectors was not deciphered until 2009 by two independent research groups.36,37 The DNA recognition property of the TAL effectors is mediated by tandem amino acid repeats (34 residues in length). Two hypervariable amino acids known as repeat-variable di-residues (RVDs) located at the 12th and 13th position in each repeat determine the binding specificity of the TAL effectors.38,39 HD, NG, NI, and NN are the four most common RVDs, accounting for each of the four nucleotides C, T, A, G, respectively. Similar to ZFN, the TALEN monomer is also fused by two independent domains: a customizable DNA-binding domain at the N-terminal and a nonspecific Fok I nuclease domain at the C-terminal (Figure 2). Due to easier manipulation, the genes modified by TALENs have been successfully used in both animal and plant species within three years of deciphering their function. These species include zebrafish,40,41 rat,42 human cells,43,44 rice,45 wheat,46 Arabidopsis,47, 48, 49 and horticultural crops such as potato50 and tomato.51
The structure of TALEN and the principle of TALEN-mediated genomic modifications. The target site of TALEN is recognized by the “left” and “right” half monomer that each consist of a tandem repeat of TALE repeats. Each TALE repeat comprises a 34 amino acid (aa) unit that differs at two hypervariable aa located at the 12th and 13th position, known as RVD, which determine the recognition specificity of each repeat. The TALEN monomer consists of an N-terminal domain containing a nuclear localization signal (NLS, red), a recognition domain typically composed of tandem TALE repeats (in different colors), and a C-terminal function domain that comprises the Fok I endonuclease. Simultaneous bindings of the left and right TALE enable dimerization of the Fok I cleavage domain, resulting in DSBs of the target DNA. Induced DSBs of the target DNA are repaired either by NHEJ or HDR resulting in gene mutations that include nucleotide insertion, deletion, or substitution around the cleavage site. TALE, transcription activator-like effector; NLS, nuclear localization signal; RVD, repeat-variable di-residues; DSB, double-strand breaks; NHEJ,non-homologous end joining; HDR, homology-directed repair. Mutation*, red color box regions contain nucleotide deletion, insertion or substitution. Figure modified from Gaj et al. (2013), Figure 119 and Moore et al. (2012), Figure 1.100
CRISPR/Cas in gene modification
More recently, a new class of genome-editing technology, i.e., the CRISPR (clustered regularly interspaced short palindromic repeats)/Cas (CRISPR-associated) system, has been developed. The principle of the CRISPR/Cas system was derived from a type II prokaryotic organism adaptive immune system.52 CRISPRs were firstly identified in the Escherichia coli genome in 1987 as an unusual sequence element consisting of a series of 29-nucleotide repeats separated by unique 32-nucleotide “spacer” sequences.53,54 Later, repetitive sequences with a similar repeat-spacer-repeat pattern were identified in other bacterial and archaeal genomes, but the functions of these repeats remained obscure until 2005 when three independent research groups found the spacer sequence was identical to some part of the viral and plasmid sequence.55, 56, 57 Further investigations indicated that CRISPRs function through an RNA interference-like mechanism to recognize and cleave foreign DNA.58 As shown in Figure 3, the type II CRISPR/Cas from Streptococcus pyogenes, a short CRISPR RNA (crRNA), is able to recognize a complementary stretch of nucleotides in alien DNA and determines the sequence specificity. In addition, a transactivating crRNA (tracrRNA) is required to form a ribonucleoprotein complex with Cas9 nuclease to generate site-specific DSBs.52,59 Later, investigators found that the components of crRNA and tracrRNA could be combined into a single RNA chimera, which was termed as guide RNA (gRNA).60 Efficient cleavage also requires the presence of the protospacer adjacent motif (PAM) in the complementary strand following the recognition sequence.52
Schematic illustration of the CRISPR/Cas9 system structure and principle of CRISPR/Cas9-mediated genomic modifications. The synthetic guide RNA (sgRNA) contains a region (usually 20 bp in length) complementary to the target site on the genomic loci and stem loops that mediate the binding of the Cas9 protein. The protospacer adjacent motif (PAM, NGG) required for cleavage is indicated in red, the Cas9 protein is shown by the brown circle, and the cleavage sites located 3 bp from the PAM motif are indicated by scissors. Induced DSBs of the target DNA are repaired either by NHEJ or HDR resulting in gene mutations that include nucleotide insertion, deletion or substitution around the cleavage sites. sgRNA, synthetic guide RNA; DSB, double-strand breaks; NHEJ, non-homologous end joining; HDR, homology-directed repair. Mutation*, red color box region contains nucleotide deletion, insertion or substitution. Figure modified from Xie and Yang (2013), Figure 1.76
In some cases, the CRISPR/Cas method may introduce unwanted off-target mutations.61, 62, 63 To eliminate the potential off-target effect of the CRISPR/Cas system, different efforts have been attempted. Hou et al. reported that a Cas9 protein from Neisseria meningitides could recognize a longer target sequence than the one from S. pyogenes, which consequently improved the target specificity of the CRISPR/Cas system.64 In addition to changing the Cas9 protein from different bacteria, other strategies have also been applied to reduce the off-target effects, such as utilization of Cas9 nickase65 and manipulation of the length of the recognition sequence in gRNA.66 These strategies dramatically reduced the off-target effects of the CRISPR/Cas system and will greatly improve the specificity of this system.
Since the first report in early 2013, this technology has been widely applied in gene modification in both animals and plants, such as zebrafish,67,68 mice,69 human cells,60,70,71 Arabidopsis,72,73 tobacco,72 rice,74, 75, 76 wheat,74 and sweet orange.77 In plants, the modified traits include changing plant architecture, e.g., Miao et al. changed the tiller angle by modifying the LAZY1 gene in rice,75 and Shan et al. changed the plant color to white by modifying the OsSPD gene in rice.74
Comparison of zfns, talens, and crispr/cas
All three genome-editing technologies—ZFNs, TALENs, and CRISPR/Cas—are able to induce DSBs at specific sites in the genome, which might be repaired by NHEJ or HDR that results in gene mutations at the target site. Compared with ZFNs and TALENs, CRISPR/Cas offers a few advantages as shown in Table 2. First and foremost, the CRISPR/Cas system is based on simple RNA/DNA hybrids that confer sequence specificity.59 Investigators can easily target a different gene by replacing a 20-bp complementary nucleotide sequence that will modify the new target gene. In contrast, ZFNs and TALENs are based on the protein-guided recognition mechanism, in which the targeting of a specific DNA sequence requires the modular assembly of pairs of recognition proteins units while the vector system is being constructed, which is time-consuming and tedious work. Therefore, ZFNs and TALENs are much more expensive than the CRISPR/Cas method and have not been widely adopted by the plant research community until now. Second, CRISPR/Cas can simultaneously introduce multiple gene disruptions,78 which allows researchers to edit multiple genes in one plant line through a single transformation without the time-consuming post-transgenic hybridization and screening processes. In summary, the CRISPR/Cas method is considered to be the most efficient, least expensive, and most user-friendly among the three genome-editing technologies.79
Potential applications and challenges of genome-editing technologies in horticultural crop breeding
Horticultural crops are plants with unique characteristics that have been domesticated and further bred to satisfy human's special dietary, medicinal, or esthetic needs.80 Therefore, the goals of horticultural crop breeding have always focused on the improvement of the unique qualities of these plants. Despite the enormous diversity of horticultural crops, many breeding goals are similar, for example, increasing the concentrations of some unique secondary metabolites; extending the shelf life of fruits, vegetables, and cut flowers; altering the plant architecture of fruit trees, ornamental flowers, and trees; improving yield potential; and enhancing plant pest and disease resistance, similar to other agronomic crops.
Some fruits and vegetables contain high contents of plant secondary metabolites that are beneficial to human health, such as anthocyanins that are reported to inhibit certain cancers, age-related degenerative diseases and cardiovascular diseases.81, 82, 83, 84 In addition, in combination with carotenoids and chlorophylls, anthocyanins are responsible for the coloration of fruits and vegetables. The accumulation of anthocyanin pigments in fruits, flowers, and vegetables is an important indicator of their ripeness and quality.85 Studies in model plants as well as in horticultural crops have revealed the conserved anthocyanin biosynthetic and regulatory pathway. Central to this process is the activity of MYB-bHLH-WD repeat (MBW) complexes that regulate the transcription of anthocyanin genes.86 Further studies identified that the MYB factors in the complex are the key regulators that determine the patterning and spatial localization of anthocyanins. In petunia, R2R3-MYB and MYB27 function as anthocyanin synthesis repressors, and MYB27 RNAi lines exhibit an obvious pigmented phenotype compared with the wild type and MYB27 overexpression lines.86 Therefore, new crop varieties with high concentrations of anthocyanin could be generated by modifying some of these genes through genome-editing technologies.
As mentioned above, the “green revolution” of agricultural crops has greatly increased grain yield, which demonstrates the importance of plant architecture. Breeding semi-dwarf statured fruit/nut trees could also revolutionize horticultural crop industries because these trees have the potential to increase productivity via higher density plantings and reduced labor cost, resulting in reduced land, water, pesticide, and fertilizer use.87 Studies have revealed the close relationship between plant architecture and phytohormones. For example, gibberellin (GA) affects plant height, while strigolactone (SL) influences shoot branching. Disruption of the function of the genes in the GA or SL biosynthetic or signaling transduction pathway could result in various phenotypes such as dwarf6 and excessive outgrowth of axillary buds,88,89 respectively. Thus, new horticultural crops with semi-dwarf phenotypes or more branches could be generated by the disruption of the functions of these gene homologs using genome-editing technologies.
Shelf life is one of the key traits that determines the quality of fruit, flowers and vegetables. Research indicates that the plant hormone ethylene plays a very important role in the process of flower wilting and fruit ripening.90 Inhibition of ethylene biosynthesis and blocking ethylene signal transduction can delay flower senescence in carnation91 and petunia,92 respectively. Thus, new horticultural crops with a longer shelf life could be generated by disrupting the key genes involved in the ethylene biosynthetic or signaling transduction pathway through genome-editing technologies.
Plant disease caused by microorganisms is another major factor that reduces shelf life and the quality and yield of horticultural crops. Powdery mildew, which is one of the most common plant diseases, is caused by different Erysiphales fungal species.93 Studies in barley showed that the gene MILDEW-RESISTANCE LOCUS (MLO) encodes a protein that represses defenses against powdery mildew disease.94 Phylogenic studies indicated that the MLO gene family is conserved in the plant kingdom.95 Loss-of-function mlo alleles in barley, Arabidopsis, tomato, and pea lead to broad-spectrum and durable resistance to the powdery mildew pathogens in these species.46,96 In 2013, Jiwan et al. reported that antisense expression of the peach MLO gene in strawberry (Fragaria × ananassa) conferred cross-species resistance to Fragaria-specific powdery mildew.97 These studies indicate the conserved biological function of this gene family. Very recently, Wang et al. used TALEN technology to modify the three homoeo-alleles of MLO in hexaploid bread wheat.46 Gain-of-function mutants exhibit heritable resistance to powdery mildew, and this highlights the potential application of genome-editing technologies in the modification of the MLO alleles in horticultural crops to generate disease resistant varieties.
The essential prerequisite of genome editing is the availability of precise genomic information and gene functions. Most of the traits mentioned above are based on genetic functional studies in model plants. Therefore, information from model plants can only serve as a reference. The lack of molecular information on horticultural crops has greatly restricted breeding efficiency, but this is drastically improving. Many horticultural crops have been whole-genome sequenced,98 including grapevine, papaya, strawberry, sweet orange, etc. In addition, a considerable higher number of various types of transcriptomes of horticultural crops are now available.99 These vast genomic data will surely facilitate elucidating the molecular control of important traits in horticultural crops and as a result, identify the precise target gene sequences for genome editing. The reference genomes, along with transcriptomes and resequencing data in many horticultural crops, may also offer unlimited targets for genome editing for characterizing the potential functions of these genes, which in turn, can help to design better gene/genome-editing strategies, especially by employing CRISPR/Cas technology. Currently, in addition to the lack of well-characterized target gene information, another major limiting factor that restricts the broad application of gene/genome-editing technologies to horticultural crops is the same challenge faced in transgenic breeding technology: Many horticultural crops remain highly recalcitrant to transformation and regeneration process, and this has also restricted molecular characterization of horticultural traits.
Conclusions and perspectives
Compared with traditional breeding methods, genome-editing technologies provide obvious advantages, as shown in Table 3. Traditional breeding technologies have allowed breeding and the selection of hundreds and thousands of unique horticultural crops with improved traits, from better qualities and extended shelf life, to novel color- and shaped fruits, vegetables, and ornamental flowers and trees. However, the long breeding cycles, high heterozygosities, lack of various degrees of preciseness in hybridization, and low frequencies of desirable mutations have made new varietal development highly resource-demanding. Transgenic technology, to some degree, is a versatile technology with unlimited application potential. This type of technology can overcome the incompatibility barriers between species by integrating foreign genes into target plant genomes, or even introducing synthetic artificial genes to generate new varieties with desired traits. However, in recent years, breeding new varieties, especially the main dietary crops, using transgenic technology has faced increasing opposition from the public. In addition, before release or commercialization, a transgenic crop variety requires years of risk assessments that result in considerable increases in time and cost.
The latest genome-editing technologies, particularly CRISPR/Cas, promise to be more efficient and precise to edit genes when the genome sequences for target genes are known. These technologies could generate new varieties through mutation breeding. However, these technologies could be as direct and efficient as transgenic methods and could be used to generate new varieties without introducing foreign genes into the plant genome in many cases. Therefore, new crop varieties generated using these methods could be considered as non-transgenic crops that might be more acceptable in countries where transgenic plants are rejected by the public.
Although there are many challenges that need to be resolved, we are optimistic that these hurdles will be removed and the site-specific, time-saving, and high-efficient genome-editing technologies, especially the CRISPR/Cas technology, will undoubtedly be incorporated into horticultural plant breeding, The last, but most important point that should be taken into consideration is the understanding and acceptance aspects of the public to new horticultural crop varieties generated using genome-editing technology. It is necessary to establish a policy for this new biotechnology and to distinct the boundaries between traditional genetically modified organisms and genome-edited organisms. Ultimately, genome-editing technology, in combination with other breeding technologies, will result in more nutritious, colorful, tasteful, and esthetic fruits, vegetables, and ornamental flowers and trees and make our lives more healthy, beautiful and enjoyable.
References
Xu Y . Molecular plant breeding. CABI, 2010.
Dirks R, Van Dun K, De Snoo CB, Van Den Berg M, Lelivelt CL, Voermans W et al. Reverse breeding: a novel breeding approach based on engineered meiosis. Plant Biotechnol J 2009; 7: 837–845.
Liu Y, Yang H, Sakanishi A . Ultrasound: mechanical gene transfer into plant cells by sonoporation. Biotechnol Adv 2006; 24: 1–16.
van Nocker S, Gardiner SE . Breeding better cultivars, faster: applications of new technologies for the rapid deployment of superior horticultural tree crops. Hor Res 2014; 1 14022.
Longhi S, Giongo L, Buti M, Surbanovski N, Viola R, Velasco R et al. Molecular genetics and genomics of the Rosoideae: state of the art and future perspectives. Hor Res 2014; 1.
Peng J, Richards DE, Hartley NM, Murphy GP, Devos KM, Flintham JE, et al. ‘Green revolution’ genes encode mutant gibberellin response modulators. Nature 1999; 400: 256–2825.
Ban Y, Honda C, Hatsuyama Y, Igarashi M, Bessho H, Moriguchi T . Isolation and functional analysis of a MYB transcription factor gene that is a key regulator for the development of red coloration in apple skin. Plant Cell Physiol 2007; 48: 958–970.
Alleweldt G, Possingham JV . Progress in grapevine breeding. Theor Appl Genet 1988; 75: 669–673.
Parry MA, Madgwick PJ, Bayon C, Tearall K, Hernandez-Lopez A, Baudo M et al. Mutation discovery for crop improvement. J Exp Bot 2009; 60: 2817–2825.
Shu QY . Turning plant mutation breeding into a new era: molecular mutation breeding. In Induced plant mutations in the genomics era. Rome: FAO, 2009: 425–427.
Hansen G, Wright MS . Recent advances in the transformation of plants. Trends Plant Sci 1999; 4: 226–231.
James C, Krattiger AF . Global review of the field testing and commercialization of transgenic plants: 1986 to 1995. ISAAA Briefs 1996; (1).
Bruening G, Lyons J . The case of the FLAVR SAVR tomato. Calif Agr 2000; 54: 6–7.
Clark D, Klee H, Dandekar A . Despite benefits, commercialization of transgenic horticultural crops lags. Calif Agr 2004; 58:89–98.
Azad MA, Amin L, Sidik NM . Gene technology for papaya ringspot virus disease management. The Scientific World J 2014; 2014: 768038.
Luo KM, Duan H, Zhao DG, Zheng XL, Deng W, Chen YQ et al. ‘GM-gene-deletor’: fused loxP-FRT recognition sequences dramatically improve the efficiency of FLP or CRE recombinase on transgene excision from pollen and seed of tobacco plants. Plant Biotechnol J 2007; 5: 263–274.
Ding J, Duan H, Deng ZN, Zhao DG, Yi GJ, Li Y . Molecular strategies for addressing gene flow problems and their potential applications in abiotic stress tolerant transgenic plants. Crit Rev Plant Sci 2014; 33: 190–204.
Carroll D . Genome engineering with targetable nucleases. Annu Rev Biochem 2014; 83: 409–439.
Gaj T, Gersbach CA, Barbas CF, 3rd. ZFN, TALEN, and CRISPR/Cas-based methods for genome engineering. Trends Biotechnol 2013; 31: 397–405.
Wyman C, Kanaar R . DNA double-strand break repair: all's well that ends well. Annu Rev Genet 2006; 40: 363–383.
Pabo CO, Peisach E, Grant RA . Design and selection of novel Cys2His2 zinc finger proteins. Annu Rev Biochem 2001; 70: 313–340.
Cathomen T, Joung JK . Zinc-finger nucleases: the next generation emerges. Mol Ther 2008; 16: 1200–1207.
Pavletich NP, Pabo CO . Zinc finger-DNA recognition: crystal structure of a Zif268-DNA complex at 2.1 A. Science 1991; 252: 809–817.
Kim YG, Cha J, Chandrasegaran S . Hybrid restriction enzymes: zinc finger fusions to Fok I cleavage domain. Proc Natl Acad Sci USA 1996; 93: 1156–1160.
Perez EE, Wang J, Miller JC, Jouvenot Y, Kim KA, Liu O et al. Establishment of HIV-1 resistance in CD4+ T cells by genome editing using zinc-finger nucleases. Nat Biotechnol 2008; 26: 808–816.
Doyon Y, McCammon JM, Miller JC, Faraji F, Ngo C, Katibah GE et al. Heritable targeted gene disruption in zebrafish using designed zinc-finger nucleases. Nat Biotechnol 2008; 26: 702–708.
Meng X, Noyes MB, Zhu LJ, Lawson ND, Wolfe SA . Targeted gene inactivation in zebrafish using engineered zinc-finger nucleases. Nat Biotechnol 2008; 26: 695–701.
Osakabe K, Osakabe Y, Toki S . Site-directed mutagenesis in Arabidopsis using custom-designed zinc finger nucleases. Proc Natl Acad Sci USA 2010; 107: 12034–12039.
Townsend JA, Wright DA, Winfrey RJ, Fu F, Maeder ML, Joung JK et al. High-frequency modification of plant genes using engineered zinc-finger nucleases. Nature 2009; 459: 442–445.
Shukla VK, Doyon Y, Miller JC, DeKelver RC, Moehle EA, Worden SE, et al. Precise genome modification in the crop species Zea mays using zinc-finger nucleases. Nature 2009; 459: 437–441.
Hsu PD, Zhang F . Dissecting neural function using targeted genome engineering technologies. ACS Chem Neurosci 2012; 3: 603–610.
Joung JK, Sander JD . TALENs: a widely applicable technology for targeted genome editing. Nature reviews Mol Cell Biol 2013; 14: 49–55.
Boch J, Bonas U . Xanthomonas AvrBs3 family-type III effectors: discovery and function. Annu Rev Phytopathol 2010; 48: 419–436.
Bogdanove AJ, Schornack S, Lahaye T . TAL effectors: finding plant genes for disease and defense. Curr Opin Plant Biol 2010; 13: 394–401.
Bonas U, Stall RE, Staskawicz B . Genetic and structural characterization of the avirulence gene avrBs3 from Xanthomonas campestris pv. vesicatoria. Mol Gen Genet 1989; 218: 127–136.
Moscou MJ, Bogdanove AJ . A simple cipher governs DNA recognition by TAL effectors. Science 2009; 326: 1501.
Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S et al. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 2009; 326: 1509–1512.
Mak AN, Bradley P, Cernadas RA, Bogdanove AJ, Stoddard BL . The crystal structure of TAL effector PthXo1 bound to its DNA target. Science 2012; 335: 716–719.
Deng D, Yan C, Pan X, Mahfouz M, Wang J, Zhu JK et al. Structural basis for sequence-specific recognition of DNA by TAL effectors. Science 2012; 335: 720–723.
Sander JD, Cade L, Khayter C, Reyon D, Peterson RT, Joung JK et al. Targeted gene disruption in somatic zebrafish cells using engineered TALENs. Nat Biotechnol 2011; 29: 697–698.
Huang P, Xiao A, Zhou M, Zhu Z, Lin S, Zhang B . Heritable gene targeting in zebrafish using customized TALENs. Nat Biotechnol 2011; 29: 699–700.
Tesson L, Usal C, Menoret S, Leung E, Niles BJ, Remy S et al. Knockout rats generated by embryo microinjection of TALENs. Nat Biotechnol 2011; 29: 695–696.
Miller JC, Tan S, Qiao G, Barlow KA, Wang J, Xia DF et al. A TALE nuclease architecture for efficient genome editing. Nat Biotechnol 2011; 29: 143–148.
Hockemeyer D, Wang H, Kiani S, Lai CS, Gao Q, Cassady JP et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nat Biotechnol 2011; 29: 731–734.
Li T, Liu B, Spalding MH, Weeks DP, Yang B . High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat Biotechnol 2012; 30: 390–392.
Wang Y, Cheng X, Shan Q, Zhang Y, Liu J, Gao C et al. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 2014; 32: 947–951.
Christian M, Cermak T, Doyle EL, Schmidt C, Zhang F, Hummel A et al. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 2010; 186: 757–761.
Cermak T, Doyle EL, Christian M, Wang L, Zhang Y, Schmidt C et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res 2011: gkr218.
Li L, Piatek MJ, Atef A, Piatek A, Wibowo A, Fang X et al. Rapid and highly efficient construction of TALE-based transcriptional regulators and nucleases for genome modification. Plant Mol Biol 2012; 78: 407–416.
Sawai S, Ohyama K, Yasumoto S, Seki H, Sakuma T, Yamamoto T et al. Sterol side chain reductase 2 is a key enzyme in the biosynthesis of cholesterol, the common precursor of toxic steroidal glycoalkaloids in potato. Plant cell 2014; 26: 3763–3774.
Lor VS, Starker CG, Voytas DF, Weiss D, Olszewski NE . Targeted mutagenesis of the tomato PROCERA gene using transcription activator-like effector nucleases. Plant Physiol 2014; 166: 1288–1291.
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E . A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012; 337: 816–821.
Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A . Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol 1987; 169: 5429–5433.
Wiedenheft B, Sternberg SH, Doudna JA . RNA-guided genetic silencing systems in bacteria and archaea. Nature 2012; 482: 331–338.
Pourcel C, Salvignol G, Vergnaud G . CRISPR elements in Yersinia pestis acquire new repeats by preferential uptake of bacteriophage DNA, and provide additional tools for evolutionary studies. Microbiology 2005; 151: 653–663.
Bolotin A, Quinquis B, Sorokin A, Ehrlich SD . Clustered regularly interspaced short palindrome repeats (CRISPRs) have spacers of extrachromosomal origin. Microbiology 2005; 151: 2551–2561.
Mojica FJ, Diez-Villasenor C, Garcia-Martinez J, Soria E . Intervening sequences of regularly spaced prokaryotic repeats derive from foreign genetic elements. J Mol Evol 2005; 60: 174–182.
Deveau H, Garneau JE, Moineau S . CRISPR/Cas system and its role in phage-bacteria interactions. Annu Rev Microbiol 2010; 64: 475–493.
Walsh RM, Hochedlinger K . A variant CRISPR-Cas9 system adds versatility to genome engineering. Proc Natl Acad Sci USA 2013; 110: 15514–15515.
Mali P, Yang L, Esvelt KM, Aach J, Guell M, DiCarlo JE et al. RNA-guided human genome engineering via Cas9. Science 2013; 339: 823–826.
Fu Y, Foden JA, Khayter C, Maeder ML, Reyon D, Joung JK et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat Biotechnol 2013; 31: 822–826.
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 2013; 31: 827–832.
Pattanayak V, Lin S, Guilinger JP, Ma E, Doudna JA, Liu DR . High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat Biotechnol 2013; 31: 839–843.
Hou Z, Zhang Y, Propson NE, Howden SE, Chu L-F, Sontheimer EJ et al. Efficient genome engineering in human pluripotent stem cells using Cas9 from Neisseria meningitidis. Proc Natl Acad Sci USA 2013; 110: 15644–15649.
Cho SW, Kim S, Kim Y, Kweon J, Kim HS, Bae S et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res 2014; 24: 132–141.
Fu YF, Sander JD, Reyon D, Cascio VM, Joung JK . Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat Biotechnol 2014; 32: 279–284.
Chang N, Sun C, Gao L, Zhu D, Xu X, Zhu X et al. Genome editing with RNA-guided Cas9 nuclease in zebrafish embryos. Cell Res 2013; 23: 465–472.
Hwang WY, Fu Y, Reyon D, Maeder ML, Tsai SQ, Sander JD et al. Efficient genome editing in zebrafish using a CRISPR-Cas system. Nat Biotechnol 2013; 31: 227–229.
Shen B, Zhang J, Wu H, Wang J, Ma K, Li Z et al. Generation of gene-modified mice via Cas9/RNA-mediated gene targeting. Cell Res 2013; 23: 720–723.
Tsai SQ, Wyvekens N, Khayter C, Foden JA, Thapar V, Reyon D et al. Dimeric CRISPR RNA-guided Fok I nucleases for highly specific genome editing. Nat Biotechnol 2014; 32: 569–576.
Guilinger JP, Thompson DB, Liu DR . Fusion of catalytically inactive Cas9 to Fok I nuclease improves the specificity of genome modification. Nat Biotechnol 2014; 32: 577–582.
Li JF, Norville JE, Aach J, McCormack M, Zhang D, Bush J et al. Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 2013; 31: 688–691.
Feng Z, Mao Y, Xu N, Zhang B, Wei P, Yang DL et al. Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proc Natl Acad Sci USA 2014; 111: 4632–4637.
Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol 2013; 31: 686–688.
Miao J, Guo D, Zhang J, Huang Q, Qin G, Zhang X et al. Targeted mutagenesis in rice using CRISPR-Cas system. Cell Res 2013; 23: 1233–1236.
Xie K, Yang Y . RNA-guided genome editing in plants using a CRISPR-Cas system. Mol Plant 2013; 6: 1975–1983.
Jia HG, Wang N . Targeted genome editing of sweet orange using Cas9/sgRNA. PloS one 2014; 9: e93806.
Wang H, Yang H, Shivalila CS, Dawlaty MM, Cheng AW, Zhang F et al. One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 2013; 153: 910–918.
Nagamangala Kanchiswamy C, Sargent DJ, Velasco R, Maffei ME, Malnoy M . Looking forward to genetically edited fruit crops. Trends Biotechnol 2015; 33: 62–64.
Janick J . Horticultural plant breeding: past accomplishments, future directions. Proc. IS on Hort. in Asian-Pacific Region 2004; 694: 61–65.
Seeram NP, Adams LS, Hardy ML, Heber D . Total cranberry extract versus its phytochemical constituents: antiproliferative and synergistic effects against human tumor cell lines. J AgrFood Chem 2004; 52: 2512–2517.
Joseph JA, Shukitt-Hale B, Denisova NA, Bielinski D, Martin A, McEwen JJ et al. Reversals of age-related declines in neuronal signal transduction, cognitive, and motor behavioral deficits with blueberry, spinach, or strawberry dietary supplementation. J Neurosci 1999; 19: 8114–8121.
Renaud S, de Lorgeril M . Wine, alcohol, platelets, and the French paradox for coronary heart disease. Lancet 1992; 339: 1523–1526.
Hou DX, Fujii M, Terahara N, Yoshimoto M . Molecular mechanisms behind the chemopreventive effects of anthocyanidins. Journal Biomed Biotechnol 2004; 5: 321–325.
Allan AC, Hellens RP, Laing WA . MYB transcription factors that colour our fruit. Trends Plant Sci 2008; 13: 99–102.
Albert NW, Davies KM, Lewis DH, Zhang H, Montefiori M, Brendolise C et al. A conserved network of transcriptional activators and repressors regulates anthocyanin pigmentation in eudicots. Plant cell 2014; 26: 962–980.
Hollender CA, Dardick C . Molecular basis of angiosperm tree architecture. New Phytol 2015; 206: 541–556.
Gomez-Roldan V, Fermas S, Brewer PB, Puech-Pagès V, Dun EA, Pillot J-P et al. Strigolactone inhibition of shoot branching. Nature 2008; 455: 189–194.
Umehara M, Hanada A, Yoshida S, Akiyama K, Arite T, Takeda-Kamiya N et al. Inhibition of shoot branching by new terpenoid plant hormones. Nature 2008; 455: 195–200.
Reid MS, Wu M-J . Ethylene and flower senescence. Plant Growth Regul 1992; 11: 37–43.
Savin KW, Baudinette SC, Graham MW, Michael MZ, Nugent GD, Lu C-Y et al. Antisense ACC oxidase RNA delays carnation petal senescence. HortScience 1995; 30: 970–972.
Shibuya K, Barry KG, Ciardi JA, Loucas HM, Underwood BA, Nourizadeh S et al. The central role of PhEIN2 in ethylene responses throughout plant development in petunia. Plant Physiol 2004; 136: 2900–2912.
Goldberg NP . Powdery mildew. Compendium of pepper diseases APS Press: St Paul 2003: 19–20.
Buschges R, Hollricher K, Panstruga R, Simons G, Wolter M, Frijters A et al. The barley Mlo gene: a novel control element of plant pathogen resistance. Cell 1997; 88: 695–705.
Devoto A, Hartmann HA, Piffanelli P, Elliott C, Simmons C, Taramino G et al. Molecular phylogeny and evolution of the plant-specific seven-transmembrane MLO family. J Mol Evol 2003; 56: 77–88.
Pavan S, Schiavulli A, Appiano M, Marcotrigiano AR, Cillo F, Visser RGF et al. Pea powdery mildew er1 resistance is associated to loss-of-function mutations at a MLO homologous locus. Theor Appl Genet 2011; 123: 1425–1431.
Jiwan D, Roalson EH, Main D, Dhingra A . Antisense expression of peach mildew resistance locus O (PpMlo1) gene confers cross-species resistance to powdery mildew in Fragaria × ananassa. Transgenic Res 2013; 22: 1119–1131.
Bolger ME, Weisshaar B, Scholz U, Stein N, Usadel B, Mayer KFX . Plant genome sequencing – applications for crop improvement. Curr Opin Biotech 2014; 26: 31–37.
Zhang S, Chen W, Xin L, Gao Z, Hou Y, Yu X et al. Genomic variants of genes associated with three horticultural traits2 in apple revealed by genome re-sequencing. Horticulture Res 2014; 1: 14045.
Moore FE, Reyon D, Sander JD, Martinez SA, Blackburn JS, Khayter C et al. Improved somatic mutagenesis in zebrafish using transcription activator-like effector nucleases (TALENs). PLoS One 2012; 7: e37877.
Acknowledgements
We thank Dr Zong-Ming Cheng (University of Tennessee) and the reviewers for their critical review and instructive comments on our manuscript. This work was supported in part by the National Natural Science Foundation of China (grant number 31401849 to J-S X).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Xiong, JS., Ding, J. & Li, Y. Genome-editing technologies and their potential application in horticultural crop breeding. Hortic Res 2, 15019 (2015). https://doi.org/10.1038/hortres.2015.19
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/hortres.2015.19
This article is cited by
-
An Insight into Modern Targeted Genome-Editing Technologies with a Special Focus on CRISPR/Cas9 and its Applications
Molecular Biotechnology (2023)
-
Mapping of resistance to corn borers in a MAGIC population of maize
BMC Plant Biology (2019)
-
CRISPR/Cas9-mediated multiple gene editing in Brassica oleracea var. capitata using the endogenous tRNA-processing system
Horticulture Research (2019)