Abstract
Moyamoya disease is a progressive steno-occlusive condition of the main intracranial arteries that results in the compensatory formation of fragile moyamoya vessels at the base of the brain. RNF213 is the most significant susceptibility gene and is often found with the p.Arg4810Lys founder variant in East Asian patients. We identified three putatively deleterious variants of this gene from three pediatric patients: two were novel, and one was a recurrent missense variant previously reported in other pediatric patients.
Similar content being viewed by others
Moyamoya disease (MMD) is a cerebrovascular disease presenting unique angiographic and clinical features. The nature of this disease is progressive cerebral ischemia due to steno-occlusive lesions around the circle of Willis, which is compensated for by the formation of collateral parenchymal vessels (called ‘moyamoya’) penetrating toward the base of the brain.1 Clinical manifestations can vary with the degree of ischemia, ranging from headache to more severe symptoms such as epilepsy, transient ischemic attack, and cerebral infarction. Additionally, a rupture of the fragile moyamoya vessels can also cause intracranial hemorrhage.2 Epidemiological and genetic studies have demonstrated that MMD is a multifactorial disease. The recurrence rates in relatives were markedly lower than that expected from simple Mendelian inheritance.3 Likewise, the Japanese p.Arg4810Lys founder variant (c.14429G>A, rs112735431) in RNF213 showed markedly reduced penetrance despite its extremely high effect size (217.0, additive model).4 We previously demonstrated allelic heterogeneity of RNF213 in the susceptibility to MMD, and the other susceptibility variants found in pedigrees with MMD also showed reduced penetrance. It is thought that the susceptibility variants of RNF213 require additional environmental or genetic factors for the development of MMD.4,5
In the present study, we analyzed three Japanese pediatric patients showing typical radiological and clinical features of MMD, although they were confirmed to be negative for the p.Arg4810Lys founder variant of RNF213.
The first patient was a 12-year-old boy. He presented with transient motor weakness of the right upper extremity while eating hot noodles at 11 years of age. Conventional angiography revealed bilateral steno-occlusive changes at the terminal portions of the internal carotid arteries (ICAs) with moyamoya vascular networks. Bilateral superficial temporal artery to middle cerebral artery anastomoses were performed and resulted in a good postoperative outcome (Figure 1a–c).
Radiological studies of the patients. Conventional angiography of the first patient shows bilateral steno-occlusive changes of the ICAs with moyamoya vascular networks (a), which are also indicated by low-signal-intensity flow voids in the bilateral basal ganglia in T1-weighted magnetic resonance imaging (MRI) (b). Post-operative magnetic resonance angiography (MRA) shows bilateral bypasses between the superficial temporal arteries and the middle cerebral arteries (c). T2-weighted MRI of the second patient shows bilateral cerebral infarctions (d). MRA shows bilateral stenosis of the ICA terminals, which is more severe on the right side (e). MRA 4 years after surgery shows stenoses in the right vertebral artery and the proximal portion of the left ICA, in addition to the disease progression on the right side (f). Three-dimensional-computed tomography angiography (3D-CTA) shows diffuse stenosis along the descending aorta involving the left renal artery (g). MRA shows a steno-occlusive change at the right ICA terminal with moyamoya vascular networks (h). T2*-weighted MRI of the third patient shows a low-signal-intensity hemorrhagic scar in the right frontal lobe (i).
The second patient was a 5-year-old boy. He suffered bilateral cerebral infarctions at the age of one (Figure 1d). Magnetic resonance angiography (MRA) demonstrated that bilateral steno-occlusive changes at the ICA terminals were more prominent on the right side with an early moyamoya vessel formation (Figure 1e). Four years after a right-side encephalo-duro-arterio-synangiosis (EDAS), the stage of the ipsilateral lesion progressed with distinct moyamoya vessel formation, although the contralateral lesion at the ICA terminal improved (Figure 1f). Multiple stenoses further developed not only in the proximal intracranial arteries (Figure 1f), but also in the descending aorta involving the left renal artery (Figure 1g). Such extracranial vascular involvement has often been reported in MMD patients.6,7
The third case was a monozygotic twin brother. He was asymptomatic; however, he was diagnosed with MMD after his twin brother died of intraventricular hemorrhage due to MMD at 11 years of age. MRA demonstrated that the right ICA terminal was steno-occlusive with a moyamoya vascular network (Figure 1h). A right side EDAS was performed to prevent stroke. However, he developed an intracerebral hemorrhage at the right frontal lobe years after the surgery (Figure 1i).
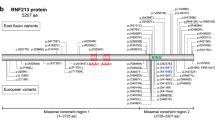
These three patients had no familial history of MMD, except for the deceased monozygotic twin brother of the third patient, and none had an underlying disease reported to complicate MMD. Radiological findings met the diagnostic criteria established by the Research Committee on Moyamoya Disease of the Ministry of Health, Labour, and Welfare, Japan,8 which were revised to include patients with both bilateral and unilateral lesions in 2015.2 They were not included in our previous genetic study of MMD in which resequencing analysis of RNF213 was performed.4 The genetic analysis of MMD patients was approved by the ethics committees of Tokyo Women’s Medical University and Tokyo Medical and Dental University. After obtaining written informed consent, genomic DNA was extracted from their peripheral blood samples. Polymerase chain reaction-based direct Sanger sequencing was performed on all coding exons of RNF213 (NM_001256071.1), as previously described,4 and the p.Arg4810Lys variant was confirmed to be negative in the three patients. From this analysis, one of three rare heterozygous, nonsynonymous variants of RNF213 was detected in each of the three patients: p.(Glu996Lys) (c.2986G>A), p.(His4058Pro) (c.12173A>C) and p.(Arg4062Gln) (c.12185G>A) (Figure 2a, b). They were not listed in the Exome Aggregation Consortium (http://exac.broadinstitute.org/) version 0.3.1 data set,9 the NCBI dbSNP147 (http://www.ncbi.nlm.nih.gov/snp) or the Human Genetic Variation Database (http://www.hgvd.genome.med.kyoto-u.ac.jp/).10 Their HumDiv and HumVar statuses of PolyPhen-2 11 were consistently deleterious according to wANNOVAR (http://wannovar.wglab.org/),12 reflecting the substitutions at the evolutionarily conserved amino-acid residues (Figure 2c).13 Two of the three variants, p.(His4058Pro) and p.(Arg4062Gln), were regarded as highly deleterious because their C-scores (23.2 and 19.05, respectively) from Combined Annotation Dependent Depletion (http://cadd.gs.washington.edu/)14 version 1.0 were more than 14.67, which was the robust cutoff for MMD susceptibility variants other than p.R4810K, which was defined in our previous study.4 The p.(Glu996Lys) variant in the first patient had a higher C-score (8.1) than the p.Arg4810Lys founder variant (6.746). Furthermore, this variant was consistently judged as deleterious even in other commonly used functional predictions such as SIFT15 and LRT,16 both of which categorized p.(His4058Pro) as tolerated. To the best of our knowledge, the p.(Glu996Lys) and p.(His4058Pro) variants have not been reported earlier in literature. The p.(Arg4062Gln) variant has been reported in another Japanese MMD pedigree with two childhood-onset patients sharing this same variant.4 This p.(Arg4062Gln) variant has also been reported in a European MMD patient.13
(a) Protein structure of RNF213, based on Q63HN8 (RN213_HUMAN) in the InterPro (http://www.ebi.ac.uk/interpro/) database. (b) DNA sequence chromatograms of the three heterozygous missense variants detected in the present study. (c) Multiple alignment of the amino-acid sequences of RNF213 from different species, using ClustalW version 2.1 (ftp://ftp.ebi.ac.uk/pub/software/clustalw2/2.1/). RefSeq (https://www.ncbi.nlm.nih.gov/refseq/) or Ensembl (http://www.ensembl.org/index.html) protein identification was shown for each RNF213 homolog. AAA, ATPases associated with diverse cellular activities; RING, really interesting new gene; FYVE, the acronym for the four cysteine-rich proteins: Fab1, YOTB, Vac1, and EEA1; PHD, plant homeodomain.
RNF213 encodes a 591-kDa protein that acts both as an AAA-type ATPase and an E3 ubiquitin ligase via two AAA+ modules and a RING finger domain, respectively.17 Several studies demonstrated that RNF213 was also associated with systolic blood pressure,18 intracranial aneurysms19 and coronary artery disease,20 suggesting that this molecule may play an important role in vascular construction. In fact, recent studies revealed its cooperative functions in angiogenic signaling pathways, such as Akt and Wnt signaling pathways, in vascular endothelial cells.17,21,22 However, the detailed biochemical functions of this molecule still remain largely unknown. The two highly deleterious variants of the three we studied, the p.(His4058Pro) and p.(Arg4062Gln) variants, were located within the RING/FYVE/PHD-type zinc finger domain according to the InterPro (http://www.ebi.ac.uk/interpro/) database (Figure 2a). Therefore, these two variants may affect the nucleotide-binding, protein-binding or E3 ubiquitin ligase activities of this domain, and are high-priority candidates for future functional analysis to elucidate the pathological role of the susceptibility variants of RNF213. Furthermore, it is important to accumulate improved knowledge about the variously reported RNF213 variants in MMD patients because there is little information on genotype–phenotype correlation.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
References
Tarasów E, Kułakowska A, Lukasiewicz A, Kapica-Topczewska K, Korneluk-Sadzyńska A, Brzozowska J et al. Moyamoya disease: diagnostic imaging. Pol J Radiol 2011; 76: 73–79.
Kim JS . Moyamoya disease: epidemiology, clinical features, and diagnosis. J Stroke 2016; 18: 2–11.
Fukuyama Y, Kanai N, Osawa M . Clinical genetic analysis on the moyamoya disease. In: Yonekawa Y (ed). The Research Committee on Spontaneous Occlusion of the Circle of Willis (Moyamoya Disease) of the Ministry of Health and Welfare Japan: Annual Report 1992. Ministry of Health and Welfare Japan: Tokyo, Japan, 1992, pp 141–146.
Moteki Y, Onda H, Kasuya H, Yoneyama T, Okada Y, Hirota K et al. Systematic validation of RNF213 coding variants in Japanese patients with moyamoya disease. J Am Heart Assoc 2015; 4: e001862.
Mukawa M, Nariai T, Onda H, Yoneyama T, Aihara Y, Hirota K et al. Exome sequencing identified CCER2 as a novel candidate gene for moyamoya disease. J Stroke Cerebrovasc Dis 2017; 26: 150–161.
Yamada I, Himeno Y, Matsushima Y, Shibuya H . Renal artery lesions in patients with moyamoya disease: angiographic findings. Stroke 2000; 31: 733–737.
Cecchi AC, Guo D, Ren Z, Flynn K, Santos-Cortez RL, Leal SM et al. RNF213 rare variants in an ethnically diverse population with moyamoya disease. Stroke 2014; 45: 3200–3207.
Research Committee on the Pathology and Treatment of Spontaneous Occlusion of the Circle of Willis Health Labour Sciences Research Grant for Research on Measures for Infractable Diseases. Guidelines for diagnosis and treatment of moyamoya disease (spontaneous occlusion of the circle of Willis). Neurol Med Chir 2012; 52: 245–266.
Lek M, Karczewski KJ, Minikel EV, Samocha KE, Banks E, Fennell T et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 2016; 536: 285–291.
Higasa K, Miyake N, Yoshimura J, Okamura K, Niihori T, Saitsu H et al. Human genetic variation database, a reference database of genetic variations in the Japanese population. J Hum Genet 2016; 61: 547–553.
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P et al. A method and server for predicting damaging missense mutations. Nat Methods 2010; 7: 248–249.
Yang H, Wang K . Genomic variant annotation and prioritization with ANNOVAR and wANNOVAR. Nat Protoc 2015; 10: 1556–1566.
Liu W, Morito D, Takashima S, Mineharu Y, Kobayashi H, Hitomi T et al. Identification of RNF213 as a susceptibility gene for moyamoya disease and its possible role in vascular development. PLoS ONE 2011; 6: e22542.
Kircher M, Witten DM, Jain P, O'Roak BJ, Cooper GM, Shendure J . A general framework for estimating the relative pathogenicity of human genetic variants. Nat Genet 2014; 46: 310–315.
Kumar P, Henikoff S, Ng PC . Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 2009; 4: 1073–1081.
Chun S, Fay JC . Identification of deleterious mutations within three human genomes. Genome Res 2009; 19: 1553–1561.
Koizumi A, Kobayashi H, Hitomi T, Harada KH, Habu T, Youssefian S . A new horizon of moyamoya disease and associated health risks explored through RNF213. Environ Health Prev Med 2016; 21: 55–70.
Koizumi A, Kobayashi H, Liu W, Fujii Y, Senevirathna ST, Nanayakkara S et al. P.R4810K, a polymorphism of RNF213, the susceptibility gene for moyamoya disease, is associated with blood pressure. Environ Health Prev Med 2013; 18: 121–219.
Zhou S, Ambalavanan A, Rochefort D, Xie P, Bourassa CV, Hince P et al. RNF213 is associated with intracranial aneurysms in the French-Canadian population. Am J Hum Genet 2016; 99: 1072–1085.
Morimoto T, Mineharu Y, Ono K, Nakatochi M, Ichihara S, Kabata R et al. Significant association of RNF213 p.R4810K, a moyamoya susceptibility variant, with coronary artery disease. PLoS ONE 2017; 12: e0175649.
Ohkubo K, Sakai Y, Inoue H, Akamine S, Ishizaki Y, Matsushita Y et al. Moyamoya disease susceptibility gene RNF213 links inflammatory and angiogenic signals in endothelial cells. Sci Rep 2015; 5: 13191.
Scholz B, Korn C, Wojtarowicz J, Mogler C, Augustin I, Boutros M et al. Endothelial RSPO3 controls vascular stability and pruning through non-canonical WNT/Ca(2+)/NFAT signaling. Dev Cell 2016; 36: 79–93.
Data Citations
Akagawa, Hiroyuki HGV Database http://dx.doi.org/10.6084/m9.figshare.hgv.1747 (2017)
Akagawa, Hiroyuki HGV Database http://dx.doi.org/10.6084/m9.figshare.hgv.1750 (2017)
Akagawa, Hiroyuki HGV Database http://dx.doi.org/10.6084/m9.figshare.hgv.1753 (2017)
Acknowledgements
We thank the patients and their families for making this study possible. We also thank the Exome Aggregation Consortium and the groups that provided exome variant data for comparison. A full list of contributing groups can be found at http://exac.broadinstitute.org/about. This work was supported by JSPS KAKENHI Grant Numbers 16K10740 (HA), 17K16629 (MM) and the Japan Intractable Diseases Research Foundation (MM).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Akagawa, H., Mukawa, M., Nariai, T. et al. Novel and recurrent RNF213 variants in Japanese pediatric patients with moyamoya disease. Hum Genome Var 5, 17060 (2018). https://doi.org/10.1038/hgv.2017.60
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/hgv.2017.60
This article is cited by
-
Difference in Clinical Phenotype, Mutation Position, and Structural Change of RNF213 Rare Variants Between Pediatric and Adult Japanese Patients with Moyamoya Disease
Translational Stroke Research (2023)
-
Comprehensive investigation of RNF213 nonsynonymous variants associated with intracranial artery stenosis
Scientific Reports (2020)
-
Novel missense variants in the RNF213 gene from a European family with Moyamoya disease
Human Genome Variation (2019)