Abstract
Purpose
Laboratory-generated genomic test reports are used to convey complex, and frequently multivariant or uncertain, information about disease risk to medical genetics professionals as well as to nonspecialist clinicians, patients, and family members. However, few guidelines exist to guide the content and format of genomic test reports, and little is known about variation in current reporting practices.
Methods
We conducted a structured content analysis of hereditary cancer gene panel test reports obtained from 16 United States–based CLIA-certified laboratories, including reports describing a variant of uncertain significance (VUS) only and reports with both a VUS and pathogenic or likely pathogenic (P/LP) test result.
Results
Report content and format varied widely across laboratories and between VUS and VUS + P/LP reports from the same laboratory, with regard to the inclusion and visual prominence of key content as well as in terms of overall length and readability.
Conclusion
Test report heterogeneity is likely to reflect both the lack of comprehensive reporting guidelines and disagreements between laboratories about the salience of specific types of information to test interpretation and use. Future research should explore the impact of reporting differences on clinician interpretation and shared decision making.
Similar content being viewed by others
Introduction
Genetic testing to identify heritable contributions to disease risk is increasingly common, with diverse test products including multigene panels and whole-exome or -genome sequencing now in routine use. The laboratory-generated test report is the primary vehicle for communicating test results and forms the basis for the initial return conversation with the medical geneticist or genetic counselor, then subsequently serves as a key resource for patients as they coordinate with other health-care providers and convey potential risks to family members. Complex, lengthy, or incautiously formatted genomic test reports are likely to pose challenges for all users, but most particularly generalist readers,1 interfering with the effective translation of genomic information for patient care and disease prevention.2,3,4
Surprisingly, few guidelines address the test-report practices of clinical genomic laboratories.1,3,5,6 Existing genomic testing standards, which focus largely on aspects of analytical validation and variant annotation, inform report content but do not address formatting or other report features relevant to patient-centered delivery.7,8 A lack of structured fields for capturing structured genomic information in current electronic health record (EHR) systems9,10 probably also contributes to inconsistent approaches to genomic test reporting. While some previous research has explored options for improved test reports targeted to patients3,11 and nongenetic health-care providers,12,13 to date no comprehensive analysis has attempted to describe current clinical genomic test-reporting practices or measure progress toward meeting existing recommendations.
Inconsistent or variably enacted reporting standards take on heightened significance in the face of the growing complexity of results generated by multigene panels and related genome-scale tests. Such tests hold the potential to identify many relevant genetic changes simultaneously and to identify pathogenic (P), likely pathogenic (LP), likely benign (LB), and/or benign (B) variants, as well as inconclusive variants of uncertain clinical significance (VUS) in the same report. VUS results, which are often difficult for generalist clinicians to interpret and patients to understand,14 are a common result from gene panel tests,15 yet previous research suggests wide variation in their reporting.16 However, no guidelines address best practices for VUS reporting, and, again, little is known about differences in content or format of test reports that include VUS results.
The goal of this study was therefore to describe variation in the content and format of laboratory reports generated from one type of gene panel test, hereditary cancer gene panel tests, with a focus on reports that include a VUS result, with the aim of characterizing the current range of reporting practices and identifying priorities for standardization as the field progresses.
Materials and methods
We conducted a content analysis to evaluate and compare the content and format of hereditary cancer panel genes test reports. We employed an approach to content analysis that was partly deductive (where initial codes are derived from observations reported in previous research) and partly inductive (where codes arose from features of the reports we examined). Two types of test reports were considered: those that report only a VUS and those that report a VUS as well as at least one P or LP variant (VUS + P/LP).
Sample
Our analysis focused exclusively on reports of clinical hereditary cancer panel gene tests consisting of two or more genes performed in a Clinical Laboratory Improvement Amendments (CLIA)-certified laboratory. We did not consider reports describing results of targeted, single-site tests or gene panel tests conducted by non-CLIA-certified labs.
To obtain test reports for our analysis we first identified 29 CLIA-certified US laboratory groups (commercial and academically affiliated) via a search of the National Center for Biotechnology Information’s Genetic Test Registry (https://www.ncbi.nlm.nih.gov/gtr/) for laboratories providing hereditary cancer genetic testing. We then sent email invitations to each laboratory, asking for a copy of two de-identified hereditary cancer gene panel sample reports: one with a VUS and one with a VUS and at least one P/LP result. Sample reports were provided by 14 laboratories (48% response rate). Nonresponders were mostly small, academically affiliated laboratories. In addition, de-identified hereditary cancer gene panel laboratory reports from two of the nonresponding laboratories were obtained from a review of patient records conducted in the context of a different institutional review board–approved study (University of Washington STUDY00000175). We estimate that together these reports cover over 80% of the cancer genetic-testing market share. Reports included here were collected between October 2016 and March 2017.
In total, laboratory reports were obtained from 16 different (3 academic and 13 commercial) laboratories performing tests of between 2 and 79 genes. VUS and VUS + P/LP reports were obtained from 11 laboratories, VUS-only reports from 3 laboratories and VUS + P/LP–only reports from 2 laboratories. Ten of 14 VUS-only reports reported a single VUS, three reported two VUSs, and one reported three VUSs. Each VUS + P/LP report contained one VUS and one P/LP variant.
To ensure that sample reports (i.e., those provided directly by the laboratory) were representative of actual patient reports, we compared de-identified patient reports to their corresponding sample reports from six laboratories. In all but one case the sample report included the same content and formatting as the de-identified patient report from the same lab; the one exception was a laboratory for which VUS-related content was considerably shorter in the actual patient report than in the sample report. In the latter case, we used the sample report for analysis.
Codebook
We developed a codebook to capture the content and format of the sampled laboratory reports. The choice of codes was directed in part by a sample cardiomyopathy panel report provided as a supplement to the 2013 American College of Medical Genetics and Genomics (ACMG) clinical laboratory standards article7 and partly guided by literature in patient-centered reporting.1,16,17 We began by test-coding two laboratory reports and edited the codebook by consensus. The process of iterative refinement of the codebook continued as we coded more reports and until no new themes were identified. The final codebook was used by two independent coders—S.M. coded all the laboratory reports and a second coder coded a randomly chosen subset of five sample reports; intercoder agreement was 80%. All areas of coder disagreement were reconciled and the codebook finalized to reflect the reconciled understanding. The final codebook was divided into two main sections: (i) report content and (ii) format:
Report content
Content codes were based on the 2013 ACMG standards.7 We noted the presence or absence of specific variant features, including gene name, zygosity, complementary DNA nomenclature, protein nomenclature, exon number, clinical relevance, and transcript variant. The number of variants reported and their classification (VUS or P/LP) were noted. We also coded for the presence or absence of distinct report sections as presented in the ACMG sample report:7 “Test Performed,” “Test Indication,” “DNA Variants,” “Interpretive Summary,” “Recommendations,” “Individual Variant Annotations,” “Test Background,” “Test Method,” “Limitations,” and “References.” VUS-related recommendations were captured verbatim and organized into thematic groups.
Report format
Three main features of report format were captured by the codebook: style, length, and readability.
Style
Style codes captured the visual prominence of text, result location, and text formatting. The visual prominence of section-specific content was coded on a scale of 1 to 3, with 3 being most prominent (i.e., content contained a heading, heading was bolded/colored/larger font size/capitalized or was preceded by blank space) and 1 being least prominent (content was embedded in other text or included under a different section heading). Location of the result (VUS or VUS + P/LP) on laboratory reports was noted by measuring the vertical distance of associated text from the top of the first page in centimeters. Text formatting of the main result section was also noted, for example, bold, color, and font size.
Length
Length of the total report, as well as of defined subsections, was measured using character and page counts. Total character count excluded header, footers, and references; supplementary information on the laboratory reports was included in estimates of total character count and page count if the information provided was specific to the reported variant.
Readability
The Flesch-Kincaid reading grade level of report content (minus headers, footers, and references) was assessed using the tool at http://www.readability-score.com.
Data analysis
Data were analyzed in R (version 3.3.2) and summarized using descriptive statistics (N, mean, median, and range). The Mann–Whitney U-test was used to compare means of nonparametric data.
Results
Report content
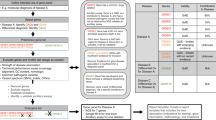
Figure 1 summarizes the presence and visual prominence of specific report sections corresponding to those suggested by the ACMG Laboratory Standards for Next Generation Sequencing example report7. Further information about the content observed within the 10 report sections is given below:
Section-specific content of hereditary cancer gene panel laboratory reports. Section headings as indicated in the American College of Medical Genetics and Genomics sample report.7
Test performed
Most reports (13 of 16) clearly indicated the name of the specific laboratory test performed. However, in many cases proprietary test names did not specify the number of genes included as part of the test panel. Instead, a list of individual genes included in the panel was provided elsewhere in the report.
Test indication
Indication was infrequently included in test reports (6 of 16 reports). When included, the indication for testing was typically described using a generic statement such as “personal or family history suggestive of hereditary predisposition for cancer.”
DNA variant
Sequence variant results were present in every report. However, the nomenclature used to describe variants varied considerably across laboratories. None of the laboratories included all seven of the variant features suggested by the ACMG. Seven laboratories each included six of the seven features, and five other laboratories each included only five features. Four laboratories each included only four types of variant features in their reports (Supplementary Table S1 online). In contrast to the ACMG sample, these pieces of information were not always easily discernible, as they were frequently embedded in other text. Gene name, protein change, DNA variant, zygosity, and interpretation were most commonly included in the main results section and therefore visually distinct.
Interpretive summary
All but one laboratory included high-level interpretive statements, as opposed to detailed variant annotations, to explain the clinical significance of the DNA variant(s) identified. A positive test result was usually closely followed by an explanation of the risk it confers for a specific type of cancer.
“A variant associated with increased risk for colon cancer was identified in APC.” (Laboratory 8)
“The … alteration in TP53 results in a stop codon and is therefore predicted to be deleterious. This result is consistent with a diagnosis of Li Fraumeni syndrome for this individual.” (Laboratory 10)
For VUS results, interpretative summary statements most often simply restated the finding, i.e., “A [hetero/homo]-zygous VUS was found in [name of gene].” In other cases, more meaningful explanations were provided:
“Inconclusive: based on currently available information, it is unclear whether variant is pathogenic or benign.” (Laboratory 7)
“A clear explanation of the patient’s condition was not found due to only variants of unknown significance.” (Laboratory 12)
Inconsistent terminology was used in summarizing interpretations across laboratories. For example, in VUS-only reports, interpretive summaries were often written as “Positive for a variant with unknown significance” or “Negative; additional findings: a variant of uncertain significance” or “Inconclusive: variant of unknown significance detected” or even “Complex (see result and interpretation).”
Result recommendation
Reports always provided some recommendation in response to a test result. Recommendations for pathogenic variants almost always included detailed management guidelines, implications for relatives, and option(s) for targeted testing of relatives.
Follow-up actions suggested in response to VUS results are summarized in Table 1. Reports often included a statement on genetic counseling (14/16) and mentioned targeted testing for relatives (10/16). Half of all laboratories invited patients to enroll in a variant reclassification program and/or other genetic research program. Only 4 of 16 reports explicitly noted that VUS results should not be used for medical management decisions and only 3 of 16 laboratories stated that predictive testing in relatives was contraindicated.
In VUS + P/LP reports, it was often difficult to tell if a given recommendation was related to the VUS or P/LP result. Some reports organized their recommendations under strongly worded headings such as “Clinical management,” whereas others used unremarkable headings such as “Notes,” “Suggestion,” or similar. In others, recommendations were included within other sections, making those recommendations difficult to identify on first inspection. Finally, a handful of reports did not include any additional recommendations beyond the suggestion that genetic counseling should be pursued.
Variant annotation
A section describing the scientific evidence used to classify each sequence variant was nearly always included in the laboratory reports (14/16). Generally, the section was written in highly technical language, using numerous acronyms. Sometimes this section was merged with the interpretive summary or recommendation.
Test background
A short background summary of the test and its application was included in only two laboratory reports (laboratories 4 and 5) although such a background section is suggested in the ACMG standards. One other laboratory had a section labeled “Test background,” but the information included in the section described the test indication rather than the test background. A website link to test background was included in only one report.
Test method, limitation, and reference
Content describing the “test method,” also labeled “assay method” or ”analysis description,” was almost always either present in the report itself (n = 14) or on the laboratory website linked to the report (n = 2). In one report, all three of these sections were only available online. “Limitation” was usually included as a separate section or included under methods, and a list of “references” was present in half (8/16) of the laboratory reports. In some cases full in-text citations were included instead of a separate list of references.
Report format
Style
Formatting of the main results section varied widely: 10 of 16 laboratories chose to draw a box around the results, 5 of 16 highlighted the area in color, 10 of 16 wrote part or all of the result in larger font size, 6 of 16 bolded the text, and 6 of 16 used capitalized fonts (data not shown). On occasion, color was used either to highlight text or as text color, but no two laboratories used exactly the same color—red or blue were used for a P/LP variant, green for a benign variant, and anything from yellow through gray to no color for a VUS.
Overall length
Reports varied widely in overall length, as measured using total character count. On average, reports were 1,648 characters (range: 280–3,867 characters) and 5 pages long (range: 2–13 pages). The longer reports contained detailed supplementary sections on risk management, information on screening guidelines, and frequently asked questions.
Readability
The average Flesch-Kincaid reading grade levels of VUS and VUS + P/LP reports were 10.3 and 9.98; average ranges: 7.4–12 and 7.1–12.2 respectively. Most laboratory reports were written at a tenth-grade reading level, which is considerably higher than the fourth- to sixth-grade reading levels recommended for patient health materials by the American Medical Association17 and the National Library of Medicine.18
VUS versus VUS + P/LP report format
We compared VUS and VUS + P/LP reports with regard to length as well as the location of information about either the VUS or VUS and pathogenic variant in the report.
Overall length
The median length of reports was 5 pages, with a range of 2–13 pages. On average, VUS-only reports tended to be shorter than VUS + P/LP reports (mean page length: 4.9 and 6 pages, respectively); however, the difference was not statistically significant. VUS + P/LP reports were longer than VUS-only reports in six laboratories and equal in length in four laboratory reports.
On average, VUS reports were 173 characters shorter (mean: 1,570, range: 280–3,358) than VUS + P/LP reports (mean = 1,743, range: 362–3,867). This difference was not statistically significant (Mann–Whitney W = 81, p = 0.31). The number of characters used to explain each type of variant—as calculated by tabulating the character length of results, variant interpretation, and recommendations sections specific to that variant—was also not significantly different between VUS and VUS + P/LP reports (data not shown).
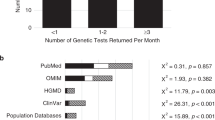
For laboratories that reported both VUS and P/LP results on a single report, 12 of 13 reports dedicated more characters to explaining P/LP variants than to explaining VUS results (Figure 2); laboratory 10 was the exception. P/LP variants were explained on average using 30.6% of total report length, whereas only 15.7% of report length was devoted to the explanation of VUS.
Location of VUS and P/LP variants
All laboratories reported variants within the first 15 cm of the top of the first page of the report. Most VUS + P/LP—containing reports (10 of 13) reported the VUS result after the P/LP result (Figure 3). The distribution of variant location in the two groups differed significantly (Mann–Whitney W = 42.5, p = 0.011). There were multiple-page-long gaps between the presentation of the P/LP and VUS results on some reports (up to 4.5 pages).
Location of text describing pathogenic/likely pathogenic (P/LP) variant and variant of uncertain significance (VUS) results, respectively, in VUS + P/LP hereditary cancer gene panel laboratory reports ( n = 13), measured as the distance from the top of page 1. Approximate page numbers (estimated using dimensions of US letter-sized paper, i.e., 8.5 × 11 in.) are shown on the vertical axis.
Discussion
This analysis demonstrates considerable variation in the ways in which United States–based CLIA-certified laboratories report the results of hereditary cancer gene panels, including wide differences in the treatment of uncertain (i.e., VUS) findings. Differences were observed in both the content and format of variant presentation, recommendations offered, and the relative emphasis placed on P/LP versus VUS. The observed heterogeneity is likely to reflect both the lack of comprehensive report guidelines and informal (and possibly unrecognized) disagreements among laboratories about the salience of specific types of information germane to test interpretation and use. Whatever the explanation, such heterogeneity is likely to challenge generalist clinicians and may discourage patients from engaging in meaningful conversations with their health-care providers and families about test findings. There is clearly an urgent need for greater standardization and changes to reporting that will incorporate key elements of patient-centered design.
In other areas of laboratory practice, attention to patient-centered communication has suggested key areas for improvement of laboratory test reports including content standardization, use of consistent terminology, and clarity of communication.19 In this study, there was wide variation in laboratory test report content, particularly with respect to “Test Indication” and “Test Background,” which were frequently absent. Furthermore, no laboratory report included all seven of the DNA variant elements suggested by the 2013 ACMG clinical laboratory standards.7 Similarly, terminology used to describe the DNA variant in the “Interpretive Summary” varied, with some laboratories providing substantive interpretations and other laboratories only reiterating the main finding in a sentence. Some of the observed variability may reflect lack of consensus on elements or be due to the fact that the rationale for certain components, such as “Background,” is not clear in the ACMG standards. Whatever the explanation, such variation is likely to impact nonspecialists and less literate patients, who may not be able to make full use of test findings for clinical management.
“Interpretative Summary” and “Recommendation” sections, which are most critical for patient and provider comprehension, ideally should be written at a reading level more consistent with universal readership. Of course, it is always challenging to convey highly technical information in a manner accessible to patients with limited genetics background and low literacy. The disparity between language used in laboratory reports and the reading level of the user20,21 could, however, be somewhat alleviated by using more informative section headings (for example, “Recommendation” versus “Notes”). Relegating technical details to online delivery (with links accessible to interested readers) might also help aid readability, but care should be taken to ensure that essential information is conveyed for those patients22 or providers23 who may not otherwise access such information.24
In addition to general content and format variability, we also observed highly inconsistent reporting of uncertain test results. Not only were VUS findings variously described and explained, clinical guidance about the fact that medical management decisions should not be based on a VUS was not present in a majority of the laboratory reports. We also observed inconsistency in other types of VUS-related recommendations which could confuse or otherwise disadvantage inexpert readers. Variation in the treatment of uncertain test results may reflect ambivalence in the medical genetics community about whether and how such information should be conveyed to patients. While it could be appropriate to limit discussion and/or description of uncertain results for which no clinical action is recommended, abridged reports that don’t fully explain VUS results can also invite misinterpretation or misunderstanding on the part of both nonspecialist clinicians and their patients.14,16
Currently, the tangible impact of these differences and apparent deficiencies is unknown. Providers have expressed ambivalence about the presence of explicit clinical recommendations on genetic test reports—while some welcomed recommendations, others felt that clinical decision making should be left to individual providers.5,6 In many cases providers are able to choose a laboratory whose reporting practices they prefer15 and laboratories can, of course, tailor reports based on feedback from those providers, perpetuating reporting heterogeneity. However, with the increasing use of genetic test reports by multiple members of the health-care team, greater consistency and adherence to accepted standards may be needed. Reporting heterogeneity could also challenge efforts to reclassify uncertain variants if, for example, members of the same family received different information about a VUS unique to their family from different laboratories.
Much of the reporting heterogeneity we observed could be effectively addressed by the adoption of synoptic reporting,6,25 i.e., a report format in which the information elements are presented in a predefined, structured tabular form.26 Using standardized data fields to report results from gene test panels could benefit patients and providers by helping them quickly locate important information as well as facilitating integration of genetic information into the EHR10 as previously recommended for molecular pathology reports.25,27 Synoptic reporting would be particularly important for reporting variant data (e.g., complementary DNA or protein change, variant classification, and zygosity), facilitating subsequent information-seeking and sharing, irrespective of the classification status. While EHR systems are being developed to accommodate such reporting, paper-report formatting conventions can make information retrieval easier for readers immediately, via the use of visually distinct section headings (through text formatting or color), as suggested previously.1,28 Although color and formatting cannot be retained in the EHR as coded text, formatting is still useful for viewing scanned copies of laboratory reports that are uploaded into EHRs and for lower-socioeconomic-status patients who are less likely to use EHR-based patient portals to view reports.29,30
Limitations of the study include a focus on one or two laboratory reports only from each laboratory, as well as an incomplete sampling of United States–based CLIA-certified laboratories, which may mean the reports we analyzed were not representative of the full range of cancer gene panel reports currently available to patients. The different variants reported may have had variable amounts of supporting evidence, which would have affected report content. In addition, the cross-sectional study design allowed us to examine only a single snapshot of the ever-evolving nature of report formats. However, because our analysis was based, in part, on an assessment of features recommended in the 2013 ACMG standards,7 we believe that enough time has elapsed to allow us to assess the extent to which laboratories have (or have not) responded to those recommendations. Finally, because we only explored cancer gene panel reports, our observations may not reflect content or format variations in play in other types of gene panel tests.
Future research should explore the impact of reporting differences on test interpretation and use by various stakeholders, including clinicians from a variety of specialties, primary-care providers, patients, and their family members. It will also be important to examine other types of genomic laboratory reports, including gene panels focused on diseases other than cancer as well as reports of whole-exome or whole-genome tests, to determine whether the heterogeneity observed here is common to other classes of test report. A better understanding of the generalizable and unique features of laboratory-test reporting in this area will help direct the development and adoption of comprehensive reporting guidelines that will promote broad accessibility and uptake in diverse health-care settings.
The National Quality Forum advises laboratories to “design work so that it is easy to do it right and hard to do it wrong.”31 Standardization of report content and format will make reporting easier for laboratories to “do right,” reduce heterogeneity in gene panel test reporting, and promote better comprehension and use of gene test information by generalist health-care providers and their patients.
References
Haga SB, Mills R, Pollak KI et al. Developing patient-friendly genetic and genomic test reports: formats to promote patient engagement and understanding. Genome Med 2014;6:58.
Kelman A, Robinson CO, Cochin E et al. Communicating laboratory test results for rheumatoid factor: what do patients and physicians want? Patient Prefer Adherence 2016;10:2501–2517.
Stuckey H, Williams JL, Fan AL et al. Enhancing genomic laboratory reports from the patients’ view: A qualitative analysis. Am J Med Genet A 2015;167A:2238–2243.
Perrotta PL & Karcher DS. Validating laboratory results in electronic health records: a College of American Pathologists Q-Probes study. Arch Pathol Lab Med 2016;140:926–931.
Williams JL, Rahm AK, Stuckey H et al. Enhancing genomic laboratory reports: A qualitative analysis of provider review. Am J Med Genet A 2016;170A:1134–1141.
Lubin IM, McGovern MM, Gibson Z et al. Clinician perspectives about molecular genetic testing for heritable conditions and development of a clinician-friendly laboratory report. J Mol Diagn 2009;11:162–171.
Rehm HL, Bale SJ, Bayrak-Toydemir P et al. ACMG clinical laboratory standards for next-generation sequencing. Genet Med 2013;15:733–747.
Richards S, Aziz N, Bale S et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015;17:405–424.
Sitapati A, Kim H, Berkovich B et al. Integrated precision medicine: the role of electronic health records in delivering personalized treatment. Wiley Interdiscip Rev Syst Biol Med 2017;9:e1378.
Shirts BH, Salama JS, Aronson SJ et al. CSER and eMERGE: current and potential state of the display of genetic information in the electronic health record. J Am Med Inform Assoc 2015;22:1231–1242.
Kaphingst KA, McBride CM, Wade C et al. Patients’ understanding of and responses to multiplex genetic susceptibility test results. Genet Med 2012;14:681–687.
Dorschner MO, Amendola LM, Shirts BH et al. Refining the structure and content of clinical genomic reports. Am J Med Genet C Semin Med Genet 2014;166C:85–92.
Scheuner MT, Hilborne L, Brown J & Lubin IM. Board motRMGTRA. A report template for molecular genetic tests designed to improve communication between the clinician and laboratory. Genet Test Mol Biomarkers 2012;16:761–769.
Vos J, Otten W, van Asperen C, Jansen A, Menko F & Tibben A. The counsellees’ view of an unclassified variant in BRCA1/2: recall, interpretation, and impact on life. Psychooncology 2008;17:822–830.
Maxwell KN, Hart SN, Vijai J et al. Evaluation of ACMG-guideline-based variant classification of cancer susceptibility and non-cancer-associated genes in families affected by breast cancer. Am J Hum Genet 2016;98:801–817.
Eccles DM, Mitchell G, Monteiro AN et al. BRCA1 and BRCA2 genetic testing-pitfalls and recommendations for managing variants of uncertain clinical significance. Ann Oncol 2015;26:2057–2065.
Weiss BD. Health Literacy: A Manual for Clinicians. American Medical Association, American Medical Foundation: Chicago, IL, 2003.
MedlinePlus. How to write easy-to-read health materials. https://medlineplus.gov/etr.html. Accessed 27 December, 2017.
Mossanen M, True LD, Wright JL, Vakar-Lopez F, Lavallee D & Gore JL. Surgical pathology and the patient: a systematic review evaluating the primary audience of pathology reports. Hum Pathol 2014;45:2192–2201.
Helitzer D, Hollis C, Cotner J & Oestreicher N. Health literacy demands of written health information materials: an assessment of cervical cancer prevention materials. Cancer Control 2009;16:70–78.
Hoffmann T & McKenna K. Analysis of stroke patients’ and carers’ reading ability and the content and design of written materials: recommendations for improving written stroke information. Patient Educ Couns 2006;60:286–293.
Norman CD & Skinner HA. eHealth literacy: essential skills for consumer health in a networked world. J Med Internet Res 2006;8:e9.
Shirts BH, Larsen N & Jackson BR. Utilization and utility of clinical laboratory reports with graphical elements. J Pathol Inform 2012;3:26.
Shirts BH, Gundlapalli AV & Jackson B. Pilot study of linking Web-based supplemental interpretive information to laboratory test reports. Am J Clin Pathol 2009;132:818–823.
Tack V, Dufraing K, Deans ZC, van Krieken HJ & Dequeker EM. The ins and outs of molecular pathology reporting. Virchows Arch;e-pub ahead of print 26 March 2017.
Leslie KO & Rosai J. Standardization of the surgical pathology report: formats, templates, and synoptic reports. Semin Diagn Pathol 1994;11:253–257.
Hammond EH & Flinner RL. Clinically relevant breast cancer reporting: using process measures to improve anatomic pathology reporting. Arch Pathol Lab Med 1997;121:1171–1175.
Valenstein PN. Formatting pathology reports: applying four design principles to improve communication and patient safety. Arch Pathol Lab Med 2008;132:84–94.
Kruse RL, Koopman RJ, Wakefield BJ et al. Internet use by primary care patients: where is the digital divide? Fam Med 2012;44:342–347.
Swartz SH, Cowan TM & Batista IA. Using claims data to examine patients using practice-based Internet communication: is there a clinical digital divide? J Med Internet Res 2004;6:e1.
Kizer KW. To err is human: Reducing medical errors. National Quality Forumhttp://www.ehcca.com/presentations/Kizer.pdf. Accessed 23 June 2017.
Acknowledgments
This research was supported by grants from the National Institutes of Health National Human Genome Research Institute (R21 HG007578 and R21HG008513).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
B.H.S. works at the Department of Laboratory Medicine, University of Washington, which is a laboratory that contributed reports for this study. The other authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Makhnoon, S., Shirts, B.H., Bowen, D.J. et al. Hereditary cancer gene panel test reports: wide heterogeneity suggests need for standardization. Genet Med 20, 1438–1445 (2018). https://doi.org/10.1038/gim.2018.23
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2018.23
Keywords
This article is cited by
-
Helping Patients Understand and Cope with BRCA Mutations
Current Oncology Reports (2022)
-
Variants of uncertain significance in the era of high-throughput genome sequencing: a lesson from breast and ovary cancers
Journal of Experimental & Clinical Cancer Research (2020)
-
Variant of Uncertain Significance-Related Uncertainty in Breast Cancer Genomics
Current Breast Cancer Reports (2020)