Abstract
Purpose:
Familial amyotrophic lateral sclerosis has been linked to mutations in 15 genes, and it is believed these genes account for less than 20–30% of Chinese patients with familial amyotrophic lateral sclerosis. Of the 163 different superoxide dismutase 1 gene mutations in amyotrophic lateral sclerosis 1, only 6.1% of them were from individuals of Chinese origin. Therefore, to quickly learn the causative gene for patients with familial amyotrophic lateral sclerosis in a Chinese pedigree, we opted to apply whole-exome sequencing as a diagnostic tool.
Methods:
To avoid time-consuming screening of known familial amyotrophic lateral sclerosis candidate genes by PCR-Sanger sequencing, we conducted whole-exome sequencing toward selected individuals of a four-generation familial amyotrophic lateral sclerosis family.
Results:
Patients in the family showed autosomal dominant features, as well as a mean onset age of 35.3 years, and a mean duration of 2.1 years. By deep sequencing analysis, we identified a novel p.Cys146X SOD1 mutation in all examined patients. Genotype–phenotype and SOD1 structural model analysis revealed the effects of the Cys57–Cys146 disulfide bond formation and the C-terminal dimer contact region on the disease phenotypes.
Conclusion:
The Cys146X mutation causes familial amyotrophic lateral sclerosis with severe phenotypes. Whole-exome sequencing becomes an attractive diagnostic tool for identifying causative genes, particularly for neurological disorders with unusual phenotypes, pleiotropic malformations, multiple known candidate genes, and complicated inheritance patterns.
Genet Med 2012:14(9):823–826
Similar content being viewed by others
Introduction
Amyotrophic lateral sclerosis (ALS) is a progressive neurodegenerative disease involving both upper motor neurons and lower motor neurons (OMIM no. 105400).1 The prevalence of ALS is about 4–8:100,000, but only 10% or less of all cases are familial (FALS). The mean age of onset is 46 in FALS with mean duration about 2–5 years.2 To date, FALS has been linked to mutations in 15 different genes and four additional chromosomal loci.3,4,5,6,7,8
Given that FALS constitutes a clinically and genetically heterogeneous entity, identification of causative genes for new patients with FALS, using either conventional PCR-Sanger sequencing to screen multiple known candidate genes or whole-exome sequencing to explore potential novel genes, poses a challenge at the clinic level. A clinical implementation of genomics has to be scaled to the new reality of whole-genome analysis.9 With the advent of next-generation sequencing technologies, the ability to quickly and relatively inexpensively learn the sequence of an individual’s entire genome will soon be available to medical practitioners, patients, and consumers.10,11 In this study of a large FALS family, we opted to apply whole-exome sequencing and, as a consequence, have successfully identified a novel mutation of the superoxide dismutase 1 (SOD1) gene. We also explored the genotype–phenotype features of eight truncated SOD1 proteins by three-dimensional structural model analysis.
Materials and Methods
This study was approved by the ethics committee of the Institute of Genomic Medicine, Wenzhou Medical College, and informed consent was obtained from all human subjects or from their parents or guardians.
The patients (described in Figure 1a ), who originated from the Northern China pedigree (Chinese Han), had a mean onset age of 35.3 years (range: 24–52 years), about 10.7 years earlier than the mean age of previously reported FALS ( Supplementary Table S1 online). The mean duration for this pedigree was only 2.1 years (ranging 0.67–8.2). The proband (III-11) began with fasciculation and weakness in the right lower extremity at age 38 and died from respiratory failure 8.2 years later. Neurological examinations showed reduced strength in all four extremities with amyotrophia. Electromyography revealed active denervation discharge in the muscles of all limbs. Patients III-9 and III-10 showed similar clinical manifestations, with the exception of symptom and duration onset at different ages.
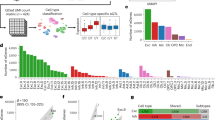
A novel Cys146X mutation in the SOD1 gene is identified in a Chinese FALS family. (a) Pedigree of the family with ALS. Males are represented by squares, females by circles, deceased by diagonals, and affected members by filled symbols. The proband (III-11) is indicated by an arrow. Asterisks (*) indicate that available DNAs were used for whole-exome sequencing. For simplicity, spouses and children of normal siblings in generations III and IV are not shown. Also, only the affected individuals and individuals visited in this study are numbered in the pedigree tree. Genotyping for IV-4, IV-5, IV-6, and IV-7 has not been performed. (b) Schematic representation of the filtering analysis of the exome data. The known SNP variants and InDels were excluded assuming the infrequency of the pathogenic variants. The heterozygous variants were screened for those affecting protein sequences. (c) A nonsense mutation Cys146X of SOD1 is confirmed by Sanger sequencing in the proband with heterozygous A/G at nucleotide 441 in exon 5. (d) Sanger sequence of the Cys146-containing region from the normal control (II-7). ALS, amyotrophic lateral sclerosis; FALS, familial amyotrophic lateral sclerosis; InDels, insertions or deletions; SNP, single-nucleotide polymorphism; SOD1, superoxide dismutase 1 gene.
Because of low-rate mutations of known ALS genes in Chinese FALS, unique phenotypes presented in these patients, and because relatively inexpensive in-house exome analysis was available, whole-exome sequencing was conducted on selected subjects ( Figure 1a ) using the Agilent SureSelect Exon Capture kit (48-Mb) (Agilent, Santa Clara, CA) and the Illumina HiSeq 2000 sequencer (Illumina, San Diego, CA). The reads were aligned for single-nucleotide polymorphism (SNP) calling and subsequent analysis ( Supplementary Table S2 online).12,13 Nonsynonymous/splice acceptor and donor site/insertions or deletions (NS/SS/InDel) variants in the dbSNP v130 (http://www.ncbi.nlm.nih.gov/projects/SNP/), HapMap samples, and 1000 Genome (http://www.ncbi.nlm.nih.gov/Ftp/) were removed. Synonymous changes were filtered using SIFT software (http://sift.jcvi.org). Sanger sequencing with specific primers was conducted to confirm the mutation of the patients.14
Results
Given that autosomal dominant FALS diseases are caused by heterozygous mutations, we assumed that the pathogenic cause for the affected individuals of this family is a single heterozygous mutation of the same gene. A total of 9,050 potential pathogenic mutations, e.g., SNP variants (missense, nonsense, and splice-site mutations) and 554 InDels (short coding insertions or deletions) were identified from three patients with shared heterozygous SNPs (870) and InDels (131) ( Supplementary Table S3 online, Figure 1b ). After removing the described variants that were also present in the controls, the shared SNPs and InDels were significantly reduced to 870 and 131, respectively ( Figure 1b ). Filtering the remaining variants against multiple databases, including the dbSNP (v130), HapMap, 1000 Genome, and 5,038 in-house exome data (from the Beijing Genomics Institute, Shenzhen, China), the candidate variants were reduced to 12 SNPs and three InDels. After the tolerated variants were removed using the SIFT program,15 three SNPs were retained, all mapped to the SOD1 gene-containing region within ~13 Mb on Chr. 21q22. A novel nonsense mutation (c.441T>A) of the SOD1 gene (GenBank acc. no., NM_000454.4) was further confirmed by Sanger sequencing ( Figure 1c,d ). This mutation changed the codon 146 TGT for cysteine into a stop codon TGA, resulting in a premature termination of the SOD1 polypeptide at the C-terminus (i.e., loss of the last eight residues, CGVIGIAQ). To determine whether the other two rare missense mutations (i.e., the c.6908T>C (p.Asn2303Ser) in the BRWD1 gene and the c.583G>A (p.Val195Met) in the KRTAP10-12 gene) are cosegregated with the ALS phenotype, and whether the pathogenesis of this disease can be attributed to these mutations, further examinations must be undertaken with DNA samples from the fourth generation of the family, as well as other unrelated affected individuals when available. However, bioinformatic analysis showed no evidence of these two variants to be the cause of ALS.
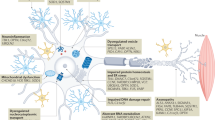
Each SOD1 monomer binds to one copper and one zinc ion; the disulfide bond (Cys57–Cys146) is required to anchor the loop of residues 49–56 ( Figure 2a ). Cys146 is located in the β8 strand of the C-terminal dimer contact region. Both the C-terminal region and the 49–56 loop contribute to the formation of the dimer interface. Cys146X is the only nonsense mutation found thus far involving the cysteine residues that forms a disulfide bridge ( Figure 2a,b ). As described earlier, the mean onset age for patients with Cys146X is 35.3, and the mean duration is 2.2 years. However, the majority of other mutations with truncated proteins were found in the active site loop between E121 and A145 ( Figure 2b , Supplementary Table S4 online),14 resulting in the truncation of 22–35 residues. These patients showed a mean age of 45.7 years at the onset, ~10.4 years later than our case. They also showed a mean duration of about 2.9 years, which is 0.7 years longer than our case but similar to those with the FALS missense mutations. An important observation to note includes a rare case where a Val29insA mutation in the β3 strand was found to contain a 33-residue polypeptide. Patients with this mutation showed an earlier mean onset age of 30.3 with incomplete penetrance. However, their mean duration of 12 years is much longer than those of other FALS patients with truncated mutations. These observations suggest that longer polypeptides in truncated SOD1 mutations are correlated with shorter durations of patients with FALS ( Supplementary Table S4 online).
Structure of SOD1 dimers and FALS-associated premature-stop mutations. (a) The monomer subunit A is colored deep salmon and the subunit B is green. The locations of Cys57 and Cys146, which form the intramolecular disulfide bond in each monomer, are shown in yellow. The C-terminal polypeptide 146–153, which is truncated in the Cys146X mutation, is shown in red. The loop between the residues 49 and 56 is shown in pink. The metal atoms in the active site are shown in spheres with copper in blue and zinc in gray. SOD1-related residues were positioned in the three-dimensional structural model using the PyMOL Molecular Graphics System (Schrödinger L. version 1.3r1; 2010). Data Bank: PDB ID code 1PU0, http://www.pdb.org. (b) FALS-associated mutations with stop codons are mapped to the SOD1 diagram. The positions of eight different premature stop codons (details in Supplementary Table S4 online) are shown in β3, the active site loop, and the β8 strand. AA, amino acid residues from 1 to 153. Cys57 and Cys146 form a disulfide bond. FALS, familial amyotrophic lateral sclerosis; SOD1, superoxide dismutase 1 gene.
Discussion
Next-generation sequencing has already shown promise as a diagnostic tool for patients with enigmatic disorders and features that suggest a primary genetic etiology, such as a strong family history, developmental anomalies, or unusual presentations of common diseases.9,16 The FALS phenotype has significant genetic heterogeneity, likely caused by mutations in more than a dozen of genes. Using whole-exome sequencing as a diagnostic tool may mitigate the need for extensive phenotypic assessments by selecting a handful of candidate genes that should be sequenced. In the case of all known FALS-causative genes being excluded, which only explain 25–35% of patients with FALS, extensive analysis of whole-exome data may identify novel causative genes for selected patients.
SOD1 mutation is rare in Chinese FALS. Of 163 different SOD1 mutations (ALS Online Genetics Database: http://alsod.iop.kcl.ac.uk) identified thus far, only 10 (6.1 %) were found in 10 different Chinese FALS families ( Supplementary Table S5 online).17 The Cys146X mutation, identified by this new strategy, is a unique SOD1 mutation associated with FALS. In these patients, the loss of the Cys57–Cys146 disulfide bond and the C-terminal dimer contact region was associated with an earlier onset of FALS at a mean age of 36.3 years. However, the disruption of the Cys57–Cys146 formation alone, such as in the case of the Cys146Arg mutation,18 was associated with a late onset of FALS in the ages between 50 and 61 years. This difference suggests the importance of the C-terminal dimer contact region for SOD1 protein stability and shows that this region may not be required for novel cytotoxic functions of mutant SOD1.
Of note, patients with shorter polypeptides, due to premature stop codons, did not show shorter duration and/or earlier onset of ALS. The fact that no null mutants have been reported suggests that the mutant SOD1 polypeptide subunit is essential for cytotoxic action. This notion is also supported by the observation that longer durations in patients were associated with a short polypeptide of only 33 residues in the Val29insA mutation, whereas shorter durations in patients were linked with longer polypeptides between 120–146 residues due to the mutations in the active-site loop. Indeed, the truncation of the polypeptide chain may dramatically open the active site, causing a decrease in substrate specificity and an increase in the catalysis of harmful chemical reactions such as peroxidation. Therefore, the increased cytotoxicity likely arises from the extreme structural and functional changes in the active site, β strands, and the dimer interface observed in the mutant enzymes.
Currently, there are a good number of studies demonstrating that all the truncating mutations would abolish or interfere with the integrity of the intrachain Cys57–Cys146 disulfide bond and thus, weaken its dimer interface and increase the propensity for aggregation.19 However, variable clinical features and courses, due to the loss of disulfide bonds of SOD1,20 suggest that other factors may be involved in the pathogenesis of the disease.
Taken together, the identification of the Cys146X mutation revealed its severe effects on the onset and duration of the disease and also suggested the important role of the disulfide bonds and the C-terminal dimer contact region in SOD1 stability. The implementation of whole-exome sequencing in ALS genomic analysis, for the first time at the clinical level, challenges the conventional model of testing and analyzing “one gene at a time.”
Disclosure
The authors declare no conflict of interest.
References
Donkervoort S, Siddique T . Amyotrophic lateral sclerosis overview. In: Pagon RA, Bird TD, Dolan CR, et al.(eds). GeneReviews. University of Washington: Seattle, WA, 1993.
Traynor BJ, Codd MB, Corr B, Forde C, Frost E, Hardiman O . Incidence and prevalence of ALS in Ireland, 1995-1997: a population-based study. Neurology 1999;52:504–509.
Siddique T, Ajroud-Driss S . Familial amyotrophic lateral sclerosis, a historical perspective. Acta Myol 2011;30:117–120.
Andersen PM, Al-Chalabi A . Clinical genetics of amyotrophic lateral sclerosis: what do we really know? Nat Rev Neurol 2011;7:603–615.
Ticozzi N, Tiloca C, Morelli C, et al. Genetics of familial Amyotrophic lateral sclerosis. Arch Ital Biol 2011;149:65–82.
Deng HX, Chen W, Hong ST, et al. Mutations in UBQLN2 cause dominant X-linked juvenile and adult-onset ALS and ALS/dementia. Nature 2011;477:211–215.
Rosen DR, Siddique T, Patterson D, et al. Mutations in Cu/Zn superoxide dismutase gene are associated with familial amyotrophic lateral sclerosis. Nature 1993;362:59–62.
Johnson JO, Mandrioli J, Benatar M, et al.; ITALSGEN Consortium. Exome sequencing reveals VCP mutations as a cause of familial ALS. Neuron 2010;68:857–864.
Berg JS, Evans JP, Leigh MW, et al. Next generation massively parallel sequencing of targeted exomes to identify genetic mutations in primary ciliary dyskinesia: implications for application to clinical testing. Genet Med 2011;13:218–229.
Evans JP, Berg JS . Next-generation DNA sequencing, regulation, and the limits of paternalism: the next challenge. JAMA 2011;306:2376–2377.
Worthey EA, Mayer AN, Syverson GD, et al. Making a definitive diagnosis: successful clinical application of whole exome sequencing in a child with intractable inflammatory bowel disease. Genet Med 2011;13: 255–262.
Hou H, Zhao F, Zhou L, et al. MagicViewer: integrated solution for next-generation sequencing data visualization and genetic variation detection and annotation. Nucleic Acids Res 2010;38(Web Server issue): W732–W736.
Li R, Li Y, Kristiansen K, Wang J . SOAP: short oligonucleotide alignment program. Bioinformatics 2008;24:713–714.
Deng HX, Hentati A, Tainer JA, et al. Amyotrophic lateral sclerosis and structural defects in Cu,Zn superoxide dismutase. Science 1993; 261:1047–1051.
Ng PC, Henikoff S . SIFT: Predicting amino acid changes that affect protein function. Nucleic Acids Res 2003;31:3812–3814.
Gahl WA, Markello TC, Toro C, et al. The National Institutes of Health Undiagnosed Diseases Program: insights into rare diseases. Genet Med 2012;14:51–59.
Zhang H, Zhao H, Lu M, et al. A rare Cu/Zn superoxide dismutase mutation causing familial amyotrophic lateral sclerosis with variable age of onset and incomplete penetrance in China. Amyotroph Lateral Scler Other Motor Neuron Disord 2005;6:234–238.
Ito K, Uchiyama T, Fukutake T, Arai K, Kanesaka T, Hattori T . [Different clinical phenotypes of siblings with familial amyotrophic lateral sclerosis showing Cys146Arg point mutation of superoxide dismutase 1 gene]. Rinsho Shinkeigaku 2002;42:175–177.
Lindberg MJ, Normark J, Holmgren A, Oliveberg M . Folding of human superoxide dismutase: disulfide reduction prevents dimerization and produces marginally stable monomers. Proc Natl Acad Sci USA 2004;101: 15893–15898.
Vassall KA, Stubbs HR, Primmer HA, et al. Decreased stability and increased formation of soluble aggregates by immature superoxide dismutase do not account for disease severity in ALS. Proc Natl Acad Sci USA 2011; 108:2210–2215.
Acknowledgements
We thank the patients and their families for their participation in this study. We thank the National Key Laboratory of Medical Genetics, at Changsha, for assisting in DNA sample collection in this study. This work was supported by the National Natural Science Foundation of China (81110114) and the Natural Science Foundation of Zhejiang Province (Z2110521).
As a disclaimer, T.C. represented his own perspective in the article, not that of the National Institute of Dental and Craniofacial Research or the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Table S1.
(DOC 41 kb)
Supplementary Table S2.
(DOC 39 kb)
Supplementary Table S3.
(DOC 38 kb)
Supplementary Table S4.
(DOC 84 kb)
Supplementary Table S5.
(DOC 89 kb)
Rights and permissions
About this article
Cite this article
Wu, J., Shen, E., Shi, D. et al. Identification of a novel Cys146X mutation of SOD1 in familial amyotrophic lateral sclerosis by whole-exome sequencing. Genet Med 14, 823–826 (2012). https://doi.org/10.1038/gim.2012.50
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2012.50
Keywords
This article is cited by
-
Strain heterogeneity, cooccurrence network, taxonomic composition and functional profile of the healthy ocular surface microbiome
Eye and Vision (2021)
-
Premature termination codons in SOD1 causing Amyotrophic Lateral Sclerosis are predicted to escape the nonsense-mediated mRNA decay
Scientific Reports (2020)
-
Quantum chemical and molecular mechanics studies on the assessment of interactions between resveratrol and mutant SOD1 (G93A) protein
Journal of Computer-Aided Molecular Design (2018)
-
Exome-assistant: a rapid and easy detection of disease-related genes and genetic variations from exome sequencing
BMC Genomics (2012)