Abstract
Mucopolysaccharidosis type I (MPS I) is an autosomal recessive genetic disorder that results in a wide range of clinical symptoms from mild somatic complications and a normal lifespan to severe central nervous system involvement and a significantly shortened lifespan. An extensive review of the literature was performed to pool data from studies that have identified mutations in patients with mucopolysaccharidosis type I (MPS I) and have reported clinical information about disease severity in an attempt to make correlations between a patient’s genotype and phenotype. To date, all patients with a nonsense mutation identified on both alleles have developed the severe form of MPS I. The phenotypes of patients with missense, insertion, deletion, or splice site mutations are much more variable. Missense mutations are the most likely to allow for some residual enzyme activity, and in particular, the R89Q mutation has been identified in several mild patients even when in combination with a nonsense mutation. Conversely, most splice site and insertion/deletion mutations result in the severe phenotype unless in combination with a less severe missense mutation. Currently, genotype-phenotype correlations cannot be confidently made unless the patient has 2 nonsense mutations. Although most families have private mutations, some insight into phenotypic expression may be obtained by observing the clinical severity of other patients with the same genotype. This review also confirms that MPS I allele frequencies vary between different ethnic populations, and that W402X and Q70X are the most common mutations and are present in over 50% of Caucasian alleles.
Similar content being viewed by others
Main
Mucopolysaccharidosis type I (MPS I) is an autosomal recessive genetic disorder that is caused by the deficiency or absence of α-l-iduronidase, a lysosomal enzyme involved in the degradation of the glycosaminoglycans heparan sulfate and dermatan sulfate. Impaired degradation of these glycosaminoglycans leads to a wide range of clinical manifestations, including hepatosplenomegaly, dysostosis multiplex, coarse facial features, severe arthropathy, hearing loss, visual impairment, restrictive lung disease, upper airway obstruction, valvular heart disease, communicating hydrocephalus, and spinal cord compression. Progressive mental retardation develops in patients at the most severe end of the disease spectrum. Historically, MPS I has been classified into 3 distinct phenotypes, severe (Hurler), mild (Scheie), and intermediate (Hurler/Scheie), but in reality, these 3 phenotypes merely represent different points on a continuous spectrum of disease severity. The 3 phenotypes are usually defined as (1) severe, when onset of symptoms are before 12 months of age, survival is < 10 years, and mental retardation manifests before the age of 3 years; (2) mild, when onset of symptoms are after 5 years of age, survival is normal, and mental retardation is absent; and (3) intermediate, when onset of symptoms is between 1 and 6 years, survival is variable, and mental retardation is absent or mild but not present before 3 years of age.1
Biochemical and immunological techniques have been only partially successful in predicting clinical severity. Although it is reasonable to assume that there must be some residual enzyme activity in milder phenotypes, it is very difficult to demonstrate such residual activity in cultured cells.2,3 Isolation and characterization of the human α-l-iduronidase gene has made it possible to identify primary disease-causing mutations and to attempt genotype-phenotype correlations.
The α-l-iduronidase gene (IDUA) is situated on chromosome 4p16.3 and contains 14 exons.4 To date, approximately 100 mutations, including missense, nonsense, splice site, deletions, and insertions, have been identified throughout the gene. The 3 phenotypes appear to be caused primarily by different combinations of mutant alleles at the IDUA gene locus. Severe patients are predicted to have mutations on both alleles that prevent the production of any functional enzyme, whereas mild patients are predicted to have at least one allele that allows for some residual enzyme activity. Patients with the intermediate phenotype have at least one allele that allows for a trace amount of residual enzyme activity to prevent severe disease.5 As an additional level of complexity, 30 nonpathogenic alleles have been reported that may influence the levels of α-l-iduronidase activity in normal individuals and may be responsible for the variable disease severity reported in some patients with the same pair of disease-causing mutations.4,5
METHODS
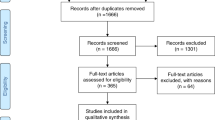
A search of the medical literature was performed in the biomedical database MEDLINE for the years 1970 to 2002. Search terms used to identify citations that provided information on patients’ genotypes and resultant phenotypes included the subject headings “mucopolysaccharidosis I/genetics,” “DNA mutational analysis,” and “mutation.” The search identified 30 relevant citations.1–30
The online Human Gene Mutation Database (http://archive.uwcm.ac.uk/uwcm/mg/hgmd0.html) was also searched for MPS I mutations, and one additional mutation was found that was not cited in MEDLINE.31
Every effort was made to prevent counting patients more than one time. This was accomplished by noting the authors and their affiliations in addition to the patient’s ethnicity, date of birth, and sex, where indicated. This information was cross-referenced among all the citations and consideration was taken to avoid duplications. Several articles specifically mentioned which patients’ genotype/phenotype correlations had been previously published.
Search terms used to identify reports of allele frequency included the subject headings “mucopolysaccharidosis I” and “gene frequency.” This literature search also included any mention of the words “common,” “commonly,” “frequent,” “frequently,” or “frequency” if present within 3 words of “mutation” or “mutations.” The search identified 11 relevant citations.4,5,7,9,10,12–14,16,18,20
RESULTS
This review identified 18 nonsense, 45 missense, 9 splice site (intron), and 25 insertion/deletion mutations in MPS I patients (Tables 1–4). Most of these mutations are associated with the severe phenotype.
Nonsense mutations
All nonsense mutations reported to date are believed to result in the total lack of enzyme activity (null mutation). Accordingly, patients who are homozygous for a particular nonsense allele or heterozygous for two different nonsense alleles have presented with the severe phenotype (Table 1).
Patients who are heterozygous for a nonsense allele and another type of mutation (missense, splice site, or insertion/deletion) have presented with a wide range of clinical phenotypes depending on the severity of the second allele. If a patient presented with the severe phenotype and had one allele that was a nonsense mutation, then the other allele was deemed also to be severe. Likewise, if a patient presented with a less severe phenotype and had one nonsense allele, then the second allele was assumed to allow for some residual enzyme function.
Missense mutations
Although the substitution of a single amino acid can severely impair enzyme function, missense mutations are most likely to be compatible with some residual IDUA activity.4 The majority of patients with mild or intermediate phenotypes have had at least one missense mutation. R89Q is the most common mild mutation, and in general, patients who have a least one allele with R89Q will present with the mild form of the disease. However, there are many reports of missense mutations in patients with severe disease. Table 2 lists known missense mutations along with the identified second allele and resulting phenotype. The missense mutation, P533R, is unusual in that it has been identified in the homozygous state in patients with mild, intermediate, intermediate-severe, and severe phenotypes. Although there is no explanation for this wide variability, MPS I disease progression appears to be less severe in P533R/W402X heterozygotes and P533R homozygotes compared to W402X or Q70X homozygotes or W402X/Q70X heterozygotes.27
Splice-site mutations
Most splice-site mutations profoundly affect normal splicing, leading to a very unstable mRNA and thus, the severe phenotype when associated with a second null mutation.5,8,20,22,25 The exception is the 678-7g>a (IVS 5-7g>a) mutation, which produces a small amount of normal IDUA mRNA and protein that prevents significant storage of substrate. This mutation has been identified in association with a null mutation in several mild patients.5,25 Table 3 lists known splice site mutations with the identified second allele and resulting phenotype.
Deletions/insertions
Twenty-three small deletions or insertions have been detected, with most causing severe MPS I. Only two insertions (396insAC, 974ins12) and one deletion (deltaD444/445) have been identified in mild patients in association with a severe allele.17 Table 4 lists known insertion and deletion mutations with the identified second allele and resulting phenotype.
Mutation frequency
Overall, the most commonly reported mutations have been the nonsense mutations, W402X and Q70X, although their frequencies vary in MPS I patients from different ethnic backgrounds. For instance, the W402X allele is frequent (45%–60%) in North America, Australia, Spain, and the United Kingdom, but is less common in Scandinavia (17%) and Italy (11%).1,4,5,20 On the other hand, Q70X has a much higher frequency in Scandinavia (62%) versus other European countries (16%), North America (17%), and Australia (17%).4,5,17,18 In Japan, no patient has been shown to carry either the W402X or Q70X mutation, whereas the R89Q and 704ins5 mutations are common.16 There is a high frequency of the homozygous P533R mutation in Moroccan patients, most likely as a result of the high occurrence of consanguineous unions.10 Table 5 includes reports from the literature of the frequency of mutations in different parts of the world.
Polymorphisms and other nonpathogenic alleles
Polymorphisms, defined as benign genetic variations present in > 1% of the normal (unaffected) population as well as other rare nonpathogenic alleles, have been described in the IDUA gene on the same allele (cis) as a known disease causing mutation. To date, 30 nonpathogenic sequence variants have been detected in the IDUA gene.5 It is not known what effect any of these sequence variants may have on the stability and processing of the IDUA mRNA, or the activity, stability, or transport of α-l-iduronidase. However, these polymorphisms likely contribute to the variability in α-l-iduronidase activity seen in healthy individuals, and it is also likely that they modify the severity of MPS I disease when present in combination with known MPS I mutations.21 For example, the A361T polymorphism, which was present in a patient with the mild allele R89Q, was thought to potentiate the deleterious effect of the R89Q mutation by decreasing the activity of the mutant α-l-iduronidase, thus altering the clinical phenotype from mild to intermediate.22 A second example is of a severe patient with 3 IDUA sequence changes: 974ins12, P496R, and the polymorphism A591T.5 As stated, the combination of the 974ins12 mutation with null mutations has been reported in patients with the mild to intermediate phenotype, implying that 974ins12 mutation allows for some residual enzyme activity. The severe phenotype in this patient was suggested to be caused by the A591T polymorphism acting in cis with the 974ins12 mutation.5
Mutation location
Attempts to predict the clinical phenotype based on the type of amino acid change or the location of the molecular lesion have been largely unsuccessful. Two mutations, R89Q and R89W, are associated with a mild phenotype, suggesting that the arginine (R) at position 89 is not vital for enzyme function.3 These mutations are in close proximity to H82P and A75T, which have resulted in severe phenotypes. On the other hand, the missense mutation A75P (with Q70X) has been reported in patients with a mild phenotype, suggesting that the replacement of alanine (A) by proline (P), rather than threonine (T), allows some functional enzyme to be produced.5
Another example is the close proximity of the 3 mild to intermediate mutations L490P, R492P, and P496L, which suggests that they lie in a location that can tolerate a nonconservative substitution to or from proline (P). However, the mutations, R489P and P496R, have been associated with the severe phenotype.2,5 Therefore, it appears that the substitution of a proline (P) for an arginine (R) at codon 496 causes greater functional disruption than replacement by a leucine (L) residue.
A final example involves the missense mutations, X654G and X654C, both of which alter the normal stop codon at the end of the coding sequence and result in a frameshift. X654G, when associated with a null mutation, results in an intermediate phenotype, whereas X654C, when present in the homozygous state, results in the severe phenotype.2,4
DISCUSSION
One of the main purposes of mutation identification is to be able to establish genotype-phenotype correlations that will allow for a patient’s clinical phenotype to be predicted from the genotype. Currently, we can only predict that nonsense mutations will invariably cause severe MPS I disease, if present on both IDUA alleles. The clinical consequences of amino acid substitutions or of other mutations that are not clearly null can be predicted only by looking at the phenotype of the patients in which these mutations have previously been identified. Even then, predictions must be made with caution because the same mutation may lead to varying severity due to the combination of IDUA alleles, attenuating polymorphisms, and other rare sequence variants, genetic background, and environmental effects.20 In addition, many patients have at least one private mutation making phenotype prediction difficult. Collection of genotype and phenotype data in a central registry will be helpful, and toward this end, Genzyme Corporation has established an MPS I Disease Registry (http://www.MPSIRegistry.com).
At this time, determination of genotype of MPS I patients is not a routine procedure, and if performed, is usually limited to a panel of nine recurrent mutations (W402X, Q70X, A327P, L218P, 474-2a>g, R89Q, P533R, A75T, and 678-7g>a). Future techniques need to be pursued to enable prediction of disease progression in patients with unique, rare, partially defined, and unknown genotypes. By further examining the stability of the IDUA mRNA and protein, better insight into the effect of a mutation on enzyme activity may be gained. A direct correlation between residual α-l-iduronidase activity in cultured MPS I fibroblasts and phenotypic severity appears promising, but the methodology is sophisticated and has been performed on only a small number of patients on a research basis.3
The accurate prediction of genotype-phenotype correlations in MPS I has significant implications given the wide spectrum of disease severity possible and the choice of treatment options. The only two specific treatments, bone marrow transplantation32–35 and the recently approved enzyme replacement therapy, recombinant human α-l-iduronidase (Aldurazyme [laronidase]),36–38 have very different risk-benefit profiles. Bone marrow transplantation has been shown to improve many of the somatic symptoms and stabilize cognitive functioning in patients with severe disease.32–35 However, because of its associated high morbidity and mortality, bone marrow transplantation is generally reserved for MPS I patients with severe disease who are under 2 years of age and have preserved cognitive function. Aldurazyme has demonstrated efficacy in improving the noncentral nervous system features of MPS I disease, specifically pulmonary function and walking capacity, but it is not expected to cross the blood-brain barrier and have an impact on cognitive function. Combination bone marrow transplantation and enzyme replacement therapy, particularly in young patients, is a third treatment option that warrants further clinical investigation. With the prospects of newborn screening for MPS I in the future, it will become all the more important to have the best predictive information available for families and caregivers to make the most rational choice for treatment.
References
Venturi N, Rovelli A, Parini R, Menni F, Brambillasca F, Bertagnolio F, et al. Molecular analysis of 30 mucopolysaccharidosis type I patients: evaluation of the mutational spectrum in Italian population and identification of 13 novel mutations. Hum Mutat 2002; 20: 231–231.
Tieu P, Bach G, Matynia A, Hwang M, Neufeld E . Four novel mutations underlying mild or intermediate forms of alpha-L-iduronidase deficiency (MPS IS and MPS IH/S). Hum Mutat 1995; 6: 55–59.
Bunge S, Clements PR, Byers S, Kleijer WJ, Brooks DA, Hopwood JJ . Genotype-phenotype correlations in mucopolysaccharidosis type I using enzyme kinetics, immunoquantification and in vitro turnover studies. Biochim Biophys Acta 1998; 1407: 249–256.
Scott HS, Bunge S, Gal A, Clarke LA, Morris CP, Hopwood JJ . Molecular genetics of mucopolysaccharidosis type I: diagnostic, clinical, and biological implications. Hum Mutat 1995; 6: 288–302.
Beesley CE, Meaney CA, Greenland GA, Adams VA, Vellodi AA, Young EP, et al. Mutational analysis of 85 mucopolysaccharidosis type I families: frequency of known mutations, identification of 17 novel mutations and in vitro expression of missense mutations. Hum Genet 2001; 109: 503–511.
Lee-Chen GJ, Lin SP, Chen IS, Chang JH, Yang CW, Chin YW . Mucopolysaccharidosis type I: identification and characterization of mutations affecting alpha-L-iduronidase activity. J Formos Med Assoc 2002; 101: 425–428.
Kroepfl T, Milos I, Paul K, Plecko B, Paschke E . The frequency of common mutations among patients with mucopolysaccharidosis types I, II and IIIA diagnosed in Austria over the last 17 years. Clin Genet 2001; 60: 393–394.
Teng YN, Wang TR, Hwu WL, Lin SP, Lee-Chen GJ . Identification and characterization of -3c-g acceptor splice site mutation in human α-l-iduronidase associated with mucopolysaccharidosis type IH/S. Clin Genet 2000; 57: 131–136.
Matte U, Leistner S, Lima L, Schwartz I, Giugliani R . Unique frequency of known mutations in Brazilian MPS I patients. Am J Med Genet 2000; 90: 108–109.
Alif N, Hess K, Straczek J, Sebbar S, N’Bou A, Nabet P, et al. Mucopolysaccharidosis type I: characterization of a common mutation that causes Hurler syndrome in Moroccan subjects. Ann Hum Genet 1999; 63: 9–16.
Lee-Chen GJ, Lin SP, Tang YF, Chin YW . Mucopolysaccharidosis type I: characterization of novel mutations affecting alpha-L-iduronidase activity. Clin Genet 1999; 56: 66–70.
Gort L, Chabas A, Coll MJ . Analysis of five mutations in 20 mucopolysaccharidosis type 1 patients: high prevalence of the W402X mutation. Hum Mutat 1998; 11: 332–333.
Voskoboeva E, Krasnopolskaya X, Mirenburg T, Weber B, Hopwood J . Molecular genetics of mucopolysaccharidosis type I: mutation analysis among the patients of the former Soviet Union. Mol Genet Metab 1998; 65: 174–180.
Gatti R, DiNatale P, Villani G, Filocamo M, Muller V, Guo X, et al. Mutations among Italian mucopolysaccharidosis type I patients. J Inherit Metab Dis 1997; 20: 803–806.
Lee-Chen GJ, Wang TR . Mucopolysaccharidosis type I: identification of novel mutations that cause Hurler/Scheie syndrome in Chinese families. J Med Genet 1997; 34: 939–941.
Yamagishi A, Tomatsu S, Fukuda S, Uchiyama A, Shimozawa N, Suzuki Y, et al. Mucopolysaccharidosis type I: identification of common mutations that cause Hurler and Scheie syndromes in Japanese populations. Hum Mutat 1996; 7: 23–29.
Bunge S, Kleijer WJ, Steglich C, Beck M, Schwinger E, Gal A . Mucopolysaccharidosis type I: identification of 13 novel mutations of the α-l-iduronidase gene. Hum Mutat 1995; 6: 91–94.
Bunge S, Kleijer WJ, Steglich C, Beck M, Zuther C, Morris CP, et al. Mucopolysaccharidosis type I: identification of 8 novel mutations and determination of the frequency of the two common α-l-iduronidase mutations (W402X and Q70X) among European patients. Hum Mol Genet 1994; 3: 861–866.
Tieu PT, Menon K, Neufeld EF . A mutant stop codon (TAG) in the IDUA gene is used as an acceptor splice site in a patient with Hurler syndrome (MPS IH). Hum Mutat 1994; 3: 333–336.
Clarke LA, Nelson PV, Warrington CL, Morris CP, Hopwood JJ, Scott HS . Mutation analysis of 19 North American mucopolysaccharidosis type I patients: identification of two additional frequent mutations. Hum Mutat 1994; 3: 275–282.
Scott HS, Nelson PV, Litjens T, Hopwood JJ, Morris CP . Multiple polymorphisms within the α-l-iduronidase gene (IDUA): implications for a role in modification of MPS-I disease phenotype. Hum Mol Genet 1993; 2: 1471–1473.
Scott H, Litjens T, Nelson P, Thompson P, Brooks D, Hopwood J, et al. Identification of mutations in the α-l-iduronidase gene (IDUA) that cause Hurler and Scheie syndromes. Am J Hum Genet 1993; 53: 973–986.
Clarke LA, Scott HS . Two novel mutations causing mucopolysaccharidosis type I detected by single strand conformational analysis of the α-l-iduronidase gene. Hum Mol Genet 1993; 2: 1311–1312.
Bach G, Moskowitz SM, Tieu PT, Matynia A, Neufeld EF . Molecular analysis of Hurler syndrome in Druze and Muslim Arab patients in Israel: multiple allelic mutations of the IDUA gene in a small geographic area. Am J Hum Genet 1993; 53: 330–338.
Moskowitz SM, Tieu PT, Neufeld EF . Mutation in Scheie syndrome (MPS IS): a G–>A transition creates new splice site in intron 5 of one IDUA allele. Hum Mutat 1993; 2: 141–144.
Moskowitz SM, Tieu PT, Neufeld EF . A deletion/insertion mutation in the IDUA gene in a Libyan Jewish patient with Hurler syndrome (mucopolysaccharidosis IH). Hum Mutat 1993; 2: 71–73.
Scott HS, Litjens T, Nelson PV, Brooks DA, Hopwood JJ, Morris CP . α-l-iduronidase mutations (Q70X, and P533R) associate with a severe Hurler phenotype. Hum Mutat 1992; 1: 333–339.
Scott HS, Litjens T, Hopwood JJ, Morris CP . A common mutation for mucopolysaccharidosis type I associated with a severe Hurler syndrome phenotype. Hum Mutat 1992; 1: 103–108.
Li P, Wood T, Thompson J . Diversity of mutations and distribution of single nucleotide polymorphic alleles in the human α-l-iduronidase (IDUA) gene. Genet Med 2002; 4: 420–426.
Matte U, Yogalingam G, Brooks D, Leistner S, Schwartz I, Lima L, et al. Identification and characterization of 13 new mutations in mucopolysaccharidosis type I patients. Mol Genet Metab 2003; 78: 37–43.
Bartholomew D, McClellan J . A novel missense mutation in the human IDUA gene associated with a severe Hurler’s phenotype. Hum Mutat 1998; 12: 291–291.
Peters C, Balthazor M, Shapiro EG, King RJ, Kollman C, Hegland JD, et al. Outcome of unrelated donor bone marrow transplantation in 40 children with Hurler syndrome. Blood 1996; 87: 4894–4902.
Peters C, Shapiro EG, Anderson J, et al. the Storage Disease Collaborative Study Group. Hurler syndrome, II: outcome of HLA-genotypically identical sibling and HLA-haploidentical related donor bone marrow transplantation in fifty-four children. Blood 1998; 91: 2601–2608.
Guffon N, Souillet G, Maire I, Straczek J, Guibaud P . Follow-up of nine patients with Hurler syndrome after bone marrow transplantation. J Pediatr 1998; 133: 119–125.
Vellodi A, Young EP, Cooper A, Wraith JE, Winchester B, Meaney C, et al. Bone marrow transplantation for mucopolysaccharidosis type I: experience of two British centres. Arch Dis Child 1997; 76: 92–99.
Kakkis ED, Muenzer J, Tiller G, Waber L, Belmont J, Passage M, et al. Enzyme replacement therapy in mucopolysaccharidosis I. N Engl J Med 2001; 344: 182–188.
Beck M, Wraith J, Clarke L, Kolodny E, Pastores G, Muenzer J . A phase 3 study of rhIDUA enzyme therapy for MPS I [abstract]. J Inherit Metab Dis 2002; 25 (suppl 1): 120.
Clarke L, Muenzer J, Kolodny E, Pastores G, Beck M, Wraith J . RhIDU enzyme replacement therapy for MPS I: 24-extension study [abstract]. Am J Hum Genet 2002; 71 (suppl 1): 581.
Antonarakis SE Nomenclature Working Group. Recommendations for a nomenclature system for human gene mutations. Hum Mutat 1998; 11: 1–3.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Terlato, N., Cox, G. Can mucopolysaccharidosis type I disease severity be predicted based on a patient’s genotype? A comprehensive review of the literature. Genet Med 5, 286–294 (2003). https://doi.org/10.1097/00125817-200307000-00004
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1097/00125817-200307000-00004