Abstract
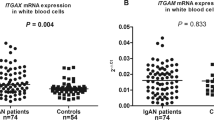
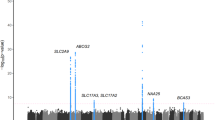
IgA nephropathy (IgAN) is a complex syndrome with high genetic heterogeneity. More recently, a genome-wide association study (GWAS) from Southern Han population revealed that variants within 8p23.1, where the DEFA genes encoding a-defensins assembled, were associated with susceptibility to IgAN. To replicate the association and fine-map the genetic variants, a case–control genetic study from an independent Northern Han cohort was conducted. A total of 60 single-nucleotide polymorphisms in a region spanning 350 kb encompassing the DEFA genes cluster were analyzed in 2096 individuals. Copy number variations of DEFA1A3 within the loci were also checked for the independent association. Functional significance of the associated variants was further examined by the in silico method as well as by cis-acting expression quantitative trait loci analysis with mRNA. It showed that 17 out of 60 (28.3%) variants were associated with susceptibility to IgAN. Two independent signals with functional potentials were discovered (rs2738058, P=4.64 × 10−5, odds ratio (OR)=0.76, 95% confidence interval (CI) 0.66–0.87 and rs9644778, P=4.78 × 10−3, OR=1.21, 95% CI 1.06–1.39). Besides, marginally significant association of rs9644778 risk genotype with lower proportion of gross hematuria (CC+CA vs AA 35.2% vs 30.2%, P=0.073) was observed. In conclusion, DEFA gene polymorphisms have potentially pathogenic roles in IgAN, and the role of mucosal immunity in the pathogenesis of IgAN has to be emphasized.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Tsukamoto Y, Wang H, Becker G, Chen HC, Han DS, Harris D et al. Report of the Asian Forum of Chronic Kidney Disease Initiative (AFCKDI) 2007. "Current status and perspective of CKD in Asia": diversity and specificity among Asian countries. Clin Exp Nephrol 2009; 13: 249–256.
Kiryluk K, Novak J, Gharavi AG . Pathogenesis of immunoglobulin A nephropathy: recent insight from genetic studies. Annu Rev Med 2013; 64: 339–356.
Kiryluk K, Julian BA, Wyatt RJ, Scolari F, Zhang H, Novak J et al. Genetic studies of IgA nephropathy: past, present, and future. Pediatr Nephrol 2010; 25: 2257–2268.
Feehally J, Farrall M, Boland A, Gale DP, Gut I, Heath S et al. HLA has strongest association with IgA nephropathy in genome-wide analysis. J Am Soc Nephrol 2010; 21: 1791–1797.
Gharavi AG, Kiryluk K, Choi M, Li Y, Hou P, Xie J et al. Genome-wide association study identifies susceptibility loci for IgA nephropathy. Nat Genet 2011; 43: 321–327.
Yu XQ, Li M, Zhang H, Low HQ, Wei X, Wang JQ et al. A genome-wide association study in Han Chinese identifies multiple susceptibility loci for IgA nephropathy. Nat Genet 2011; 44: 178–182.
Julian BA, Quiggins PA, Thompson JS, Woodford SY, Gleason K, Wyatt RJ . Familial IgA nephropathy. Evidence of an inherited mechanism of disease. N Engl J Med 1985; 312: 202–208.
Kiryluk K, Li Y, Sanna-Cherchi S, Rohanizadegan M, Suzuki H, Eitner F et al. Geographic differences in genetic susceptibility to IgA nephropathy: GWAS replication study and geospatial risk analysis. PLoS Genet 2012; 8: e1002765.
Kiryluk K, Li Y, Scolari F, Sanna-Cherchi S, Choi M, Verbitsky M et al. Discovery of new risk loci for IgA nephropathy implicates genes involved in immunity against intestinal pathogens. Nat Genet 2014; 46: 1187–1196.
Lehrer RI, Lu W . α-Defensins in human innate immunity. Immunol Rev 2012; 245: 84–112.
Saraheimo M, Forsblom C, Pettersson-Fernholm K, Flyvbjerg A, Groop PH, Frystyk J . Increased levels of alpha-defensin (-1, -2 and -3) in type 1 diabetic patients with nephropathy. Nephrol Dial Transplant 2008; 23: 914–918.
Bokarewa MI, Jin T, Tarkowski A . Intraarticular release and accumulation of defensins and bactericidal/permeability-increasing protein in patients with rheumatoid arthritis. J Rheumatol 2003; 30: 1719–1724.
Sthoeger ZM, Bezalel S, Chapnik N, Asher I, Froy O . High alpha-defensin levels in patients with systemic lupus erythematosus. Immunology 2009; 127: 116–122.
Peluso G, De Santis M, Inzitari R, Fanali C, Cabras T, Messana I et al. Proteomic study of salivary peptides and proteins in patients with Sjögren’s syndrome before and after pilocarpine treatment. Arthritis Rheum 2007; 56: 2216–2222.
Ahn JK, Cha HS, Lee J, Jeon CH, Koh EM . Correlation of DEFA1 gene copy number variation with intestinal involvement in Behcet’s disease. J Korean Med Sci 2012; 27: 107–109.
Chen Q, Hakimi M, Wu S, Jin Y, Cheng B, Wang H et al. Increased genomic copy number of DEFA1/DEFA3 is associated with susceptibility to severe sepsis in Chinese Han population. Anesthesiology 2010; 112: 1428–1434.
Jespersgaard C, Fode P, Dybdahl M, Vind I, Nielsen OH, Csillag C et al. Alpha-defensin DEFA1A3 gene copy number elevation in Danish Crohn’s disease patients. Dig Dis Sci 2011; 56: 3517–3524.
Zhou XJ, Cheng FJ, Lv JC, Luo H, Yu F, Chen M et al. Higher DEFB4 genomic copy number in SLE and ANCA-associated small vasculitis. Rheumatology (Oxford) 2012; 51: 992–995.
Guo L, Du Y, Chang S, Zhang K, Wang J . rSNPBase: a database for curated regulatory SNPs. Nucleic Acids Res 2014; 42: D1033–D1039.
Gerstein MB, Kundaje A, Hariharan M, Landt SG, Yan KK, Cheng C et al. Architecture of the human regulatory network derived from ENCODE data. Nature 2012; 489: 91–100.
Schaub MA, Boyle AP, Kundaje A, Batzoglou S, Snyder M . Linking disease associations with regulatory information in the human genome. Genome Res 2012; 22: 1748–1759.
Matys V, Kel-Margoulis OV, Fricke E, Liebich I, Land S, Barre-Dirrie A et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res 2006; 34: D108–D110.
Pique-Regi R, Degner JF, Pai AA, Gaffney DJ, Gilad Y, Pritchard JK . Accurate inference of transcription factor binding from DNA sequence and chromatin accessibility data. Genome Res 2011; 21: 447–455.
Palii CG, Perez-Iratxeta C, Yao Z, Cao Y, Dai F, Davison J et al. Differential genomic targeting of the transcription factor TAL1 in alternate haematopoietic lineages. Embo J 2011; 30: 494–509.
Fanciulli M, Norsworthy PJ, Petretto E, Dong R, Harper L, Kamesh L et al. FCGR3B copy number variation is associated with susceptibility to systemic, but not organ-specific, autoimmunity. Nat Genet 2007; 39: 721–723.
Yang Y, Chung EK, Wu YL, Savelli SL, Nagaraja HN, Zhou B et al. Gene copy-number variation and associated polymorphisms of complement component C4 in human systemic lupus erythematosus (SLE): low copy number is a risk factor for and high copy number is a protective factor against SLE susceptibility in European Americans. Am J Hum Genet 2007; 80: 1037–1054.
Sebat J, Lakshmi B, Malhotra D, Troge J, Lese-Martin C, Walsh T et al. Strong association of de novo copy number mutations with autism. Science 2007; 316: 445–449.
Black HA, Khan FF, Tyson J, Al AJ . Inferring mechanisms of copy number change from haplotype structures at the human DEFA1A3 locus. BMC Genomics 2014; 15: 614.
Teumer A, Holtfreter B, Volker U, Petersmann A, Nauck M, Biffar R et al. Genome-wide association study of chronic periodontitis in a general German population. J Clin Periodontol 2013; 40: 977–985.
Sanders JT, Wyatt RJ . IgA nephropathy and Henoch-Schonlein purpura nephritis. Curr Opin Pediatr 2008; 20: 163–170.
Wyatt RJ, Kritchevsky SB, Woodford SY, Miller PM, Roy SR, Holland NH et al. IgA nephropathy: long-term prognosis for pediatric patients. J Pediatr 1995; 127: 913–919.
Rodriguez-Garcia M, Oliva H, Climent N, Escribese MM, Garcia F, Moran TM et al. Impact of alpha-defensins1-3 on the maturation and differentiation of human monocyte-derived DCs. Concentration-dependent opposite dual effects. Clin Immunol 2009; 131: 374–384.
Barratt J, Feehally J . IgA nephropathy. J Am Soc Nephrol 2005; 16: 2088–2097.
Cheng FJ, Zhou XJ, Zhao YF, Zhao MH, Zhang H . Alpha-defensin DEFA1A3 gene copy number variation in Asians and its genetic association study in Chinese systemic lupus erythematosus patients. Gene 2013; 517: 158–163.
Drmanac R, Sparks AB, Callow MJ, Halpern AL, Burns NL, Kermani BG et al. Human genome sequencing using unchained base reads on self-assembling DNA nanoarrays. Science 2010; 327: 78–81.
Zhou XJ, Lu XL, Lv JC, Yang HZ, Qin LX, Zhao MH et al. Genetic association of PRDM1-ATG5 intergenic region and autophagy with systemic lupus erythematosus in a Chinese population. Ann Rheum Dis 2011; 70: 1330–1337.
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 2012; 489: 57–74.
Gabriel SB, Schaffner SF, Nguyen H, Moore JM, Roy J, Blumenstiel B et al. The structure of haplotype blocks in the human genome. Science 2002; 296: 2225–2229.
Acknowledgements
We thank our collaborators of Ali G Gharavi from the Department of Medicine, Columbia University College of Physicians and Surgeons, New York City, NY, USA, for genotyping sorting and assistance. We also thank all the members of the laboratory for technical assistance and the patients and their families for their cooperation and for giving consent to participate in this study. This work was supported by grants from the National Natural Science Foundation of China (No. 81200524), The Foundation of Ministry of Education of China (20120001120008), Major State Basic Research Development Program of China (973 program, No. 2012CB517700), Research Fund of Beijing Municipal Science and Technology for the Outstanding PhD Program (20121000110) and Natural Science Fund of China to the Innovation Research Group (81321064).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Disclaimer
The funders had no role in study design, data collection and analysis, decision to publish or preparation of the manuscript.
Supplementary Information accompanies this paper on Genes and Immunity website
Supplementary information
Rights and permissions
About this article
Cite this article
Qi, Y., Zhou, X., Cheng, F. et al. DEFA gene variants associated with IgA nephropathy in a Chinese population. Genes Immun 16, 231–237 (2015). https://doi.org/10.1038/gene.2015.1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2015.1