Abstract
The objective of this study was to evaluate the relationship between blood mRNA, disease activity and treatment effects in a longitudinal study of patients with dermatomyositis (DM) or polymyositis (PM). In all, 24 patients with DM or PM were followed for up to 6 years (mean of 1.9 years) at 2–7 follow-up visits while receiving standard clinical care. Clinical data and blood samples collected at 80 patient visits were used for the analysis of cytokine-induced gene expression for the signaling pathways of type 1 interferon (IFN), tumor necrosis factor-α, IL-1β, granulocyte-monocyte colony-stimulating factor, IL-10 and IL-13. A type 1 IFN signature score, but not other cytokine signature scores in the blood of patients with DM or PM, correlated highly with disease activity, decreased significantly with immunomodulatory therapies and showed concordant changes with major changes in disease activity. Type 1 IFN signature score in the blood correlates with disease activity in longitudinal follow-up of individual patients with DM or PM. The type 1 IFN-inducible gene transcripts in the blood have potential utility for monitoring disease activity in patients with DM or PM.
Similar content being viewed by others
Introduction
The inflammatory myopathies (IMs) are rare acquired autoimmune diseases of skeletal muscle.1, 2 Clinical and pathological features distinguish subtypes including dermatomyositis (DM), polymyositis (PM), inclusion body myositis (IBM), non-specific myositis and necrotizing myopathy.3 The incidence of DM/PM has been estimated at 5.5 cases per million people. Pharmacological therapy for DM and PM consists of corticosteroids and other immunomodulatory medications, while IBM responds poorly to treatment.1, 2, 4, 5 Some patients with DM or PM respond poorly to treatment and many experience morbidity from side effects. There is a need for safer and more effective treatment in these diseases.
One potential therapeutic approach is to target individual cytokine pathway that might contribute to the pathogenesis of the disease. Recognition that cytokines are present in IM muscle biopsy samples began with immunohistochemical studies for cytokine proteins.6 This approach is confounded by a number of technical and biological limitations,7 including non-specific immunoreactivity, transient expression and low concentration. For these reasons, some investigators turned to examining cytokine mRNA transcripts in IM muscle homogenates.8 Initial PCR-based studies of 12 cytokine transcripts including tumor necrosis factor (TNF)-α, IL-1β (IL-1β), granulocyte-monocyte colony-stimulating factor (GM-CSF), interferon (IFN)-α and IFN-γ generally found no IM subtype-specific differences from non-IM muscle, except for GM-CSF, which was detected in 12 of 15 IM muscle samples but none in 10 controls.8 Many subsequent studies of cytokine transcripts and proteins (reviewed in Figarella-Branger et al.9 and Salomonsson and Lundberg10) have reported variable and sometimes conflicting results.
Because of the potential for transient expression and low concentration of cytokines, their downstream persisting effects have been considered as an alternative and potentially more precise way to evaluate cytokine pathway activation. More than 25 years ago, tubuloreticular inclusions (also known as lupus inclusions), a unique pathological lesion in DM muscle endothelial cells,11, 12 were recognized as a downstream marker of type 1 IFN signaling. Tubuloreticular inclusions develop in patients treated with IFN-α13, 14 and in cultured endothelial and other cells directly in response to IFN-α and IFN-β15, 16, 17, but not IFN-γ.14 More recently, cytokine-inducible gene transcripts detected by microarrays18, 19, 20, 21 have been utilized as downstream markers of cytokine stimulation. These studies have shown marked overexpression of type 1 IFN-inducible genes in DM but not PM or IBM muscle.22
Marked overexpression of type 1 IFN-inducible genes has also been observed in the blood of patients with DM or PM,23 and in patients with systemic lupus erythematosus (SLE).24, 25 A previous cross-sectional and paired-visit study of 28 patients with DM, PM and IBM showed that type 1 IFN-inducible genes were the highest differentially expressed genes in DM and PM blood samples compared with normal controls.23 In this study, we report on studies that extended this cross-sectional study of type 1 IFN-inducible gene expression to a longitudinal study that also assessed other cytokine-inducible signaling pathways to evaluate whether activation of individual cytokine pathway correlates with the disease activities in patients with DM or PM. Such approach could provide further scientific rationale on individual cytokine as therapeutic target for patients with DM or PM along with other preclinical data.
Results
Positive correlation of type 1 IFN-inducible but not other cytokine-inducible blood gene expression with disease activity in individual patients during longitudinal follow-up
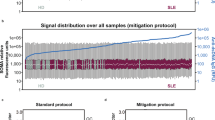
The blood gene expression profile of 136 type 1 IFN-inducible genes overexpressed in 42 patients with DM or PM at single visits (either the first visit from the longitudinal study or at the prescreening visit for patients enrolled in a clinical trial evaluating anti-IFNα therapy in DM or PM) is shown in Figure 1a. Of the 42 patients with DM or PM, 21 were DM and the other 21 were PM. The type 1 IFN signature was significantly overexpressed in the blood of both DM and PM patients, but not between DM and PM (Figure 1b). The fact that DM and PM have similar type 1 IFN signature level, along with the observation that the other cytokine signatures evaluated in the study (GM-CSF, IL-10, IL-13, IL-1β or TNF-α) also show no statistically significant difference between DM and PM (data not shown), provides scientific rationale to combine DM and PM patients to evaluate activation of cytokine pathways at molecular level in these patients. This is likely an effective way to improve statistical power of the study due to limit of sample size. Previous cross-sectional and paired-visit studies of blood gene expression profiling for 15 patients with DM or PM at 32 visits showed that type 1 IFN-inducible genes were as a group the highest differentially expressed genes compared with normal in DM and PM. In the current longitudinal study of 24 patients with DM or PM over 80 visits, 21 patients showed elevation of the type 1 IFN-inducible gene composite score (compared with healthy normal controls), as defined by a value of at least four at some time during the course of the study. These patients were carried forward in subsequent analyses. Three patients with PM (two receiving prednisone plus methotrexate, one receiving prednisone plus mycophenolate throughout the study) had low type 1 IFN-inducible gene composite scores (defined by a value <4) that never differed from scores of normal controls throughout their courses. Among these three patients, two had blood collection in three visits and the other patient had two blood collections during the whole study.
Overexpression of type 1 IFN-inducible genes in the blood of patients with DM or PM. (a) Heatmap representing expression of 136 type 1 IFN-inducible genes significantly overexpressed (q-value < 0.05 and fold change >2) in DM (N=21) and PM subjects (N=21) compared with normal healthy donors (N=24) The horizontal bar at the bottom of the figure identifies the following: normal healthy donors (black), weak type 1 IFN-inducible gene signature score (fold change < 4; blue), moderate type 1 IFN-inducible gene signature score (4⩽fold change < 10; green), and high type 1 IFN-inducible gene signature score (fold change⩾10; red). (b) Type 1 IFN gene signature scores for the same set of subjects as in (a), grouped by normal healthy donors (black), DM (green) and PM (blue) subjects. Horizontal bars represent the median value for each group. Significance is defined as follows: **P<0.01; NS, not significant.
The 21 patients with an elevated type 1 IFN-inducible gene composite score were stratified into a two-group comparison with low (modified Myositis Intention to Treat scale (MITAX) score ⩽6; nine patients) and high disease activity (modified MITAX score >6; 12 patients) categories. Only the type 1 IFN signature was significantly different between all three comparisons of: normal healthy donors and patients with low or moderate disease activity, normal healthy donors and patients with high disease activity, and patients with low or moderate disease activity and patients with high disease activity (Figure 2a). For the rest of the cytokine signatures studied (GM-CSF, IL-10, IL-13, IL-1β or TNF-α), there was no significant difference in the cytokine signature scores between patients with low or moderate disease activity, and patients with high disease activity (Figure 2a; See Supplementary Table S4 for P-value).
Correlation of type 1 IFN-inducible gene signature score in the blood (but not other cytokine-inducible gene signatures) with disease activity of individual patients with DM or PM during longitudinal follow-up. (a) Grouped analysis of various cytokine-inducible composite gene signature scores for 21 type 1 IFN signature-positive patients at visit 1 and for 36 normal healthy donors. Patients are grouped by relatively low or moderate clinically measured disease activity (modified MITAX⩽6) and high disease activity (modified MITAX>6). Horizontal bars represent the median value for each group. Only the type 1 IFN signature is significantly different between all three comparisons of: normal healthy donors and patients with low or moderate disease activity, normal healthy donors and patients with high disease activity and patients with low or moderate disease activity and patients with high disease activity. (b) Patient-specific analysis of the cytokine-inducible gene signatures for 18 patients with any variation in disease activity and (c) three patients that demonstrated no change in disease activity throughout the study. For panels b, c each patient is represented by a line. Significance is defined as follows: *P<0.05; **P<0.01; ***P<0.0001; NS, not significant.
The type 1 IFN-inducible 13-gene signature score furthermore showed high correlation with clinical disease activity within individual patients during longitudinal follow-up. Patients (N=18) with any variation in disease activity among their visits showed strong correlation between modified MITAX score changes and changes in the 13-gene type 1 IFN-inducible gene composite score (P=0.0002; Figure 4b). The 13-gene score increased in every pair of lowest-to-highest disease activity visits in each of the 18 patients. Other cytokine-inducible gene composite scores showed no such correlation with disease activity (Figure 2b). Patients (N=3) with no variation in disease activity (as defined in the Materials and methods) showed no changes in expression levels of any cytokine-inducible composite scores (Figures 2c and 4c).
Comparison of type 1 IFN-inducible gene score and single gene transcript measurements with disease activity. (a) Grouped analysis of transcripts IFI44L and RSAD2 measurements for 21 type 1 IFN signature-positive patients at visit 1 and for 36 normal healthy donors. Patients are grouped by relatively low or moderate clinically measured disease activity (modified MITAX⩽6) and high disease activity (modified MITAX>6). Horizontal bars represent the median value for each group. (b) Patient-specific analysis of the type 1 IFN-inducible gene score, IFI44L and RSAD2 for 18 patients with any variation in disease activity during the longitudinal follow-up. (c) Three patients that showed no change in disease activity throughout the study. For panels b, c, each patient is represented by a line. Significance is defined as follows: *P<0.05; **P<0.01; ***P<0.0001; NS, not significant.
Major clinical disease activity changes were associated with major changes in the type 1 IFN-inducible gene signature score. Among the 80 DM and PM patient visits, there were 50 pairs of consecutive visits, with 16 major changes in disease activity (defined as a change in modified MITAX score of >3; 11 improvements and five worsening events). All 16 changes were associated with major changes in the type 1 IFN-inducible gene composite score in the concordant direction (Table 1). Furthermore, for the 12 patients with at least three follow-up visits (total of 53 visits), the gene composite score was significantly correlated with disease clinical activity (P=0.0007) in a repeated measures analysis after controlling for treatment status. Other cytokine-inducible gene scores did not significantly predict disease activity (P>0.05 for all). For the seven patients with four or more visits, correlation between changes in disease activity and type 1 IFN-inducible composite scores persisted (Figure 3a), particularly for those patients with enough visits (at least five) to quantify the Pearson's correlation (range 0.73–0.90). For one PM refractory patient (BGE92), a limitation of the modified MITAX score (it reaches a maximum and cannot increase with further increases in disease activity) limited its use, but the serum creatine kinase (increasing from 3369–32 877 IU l−1) reflected the changes in disease activity and correlated strongly with the changes in the type 1 IFN signature score (Figure 3b).
DM and PM patients demonstrate a correlation between modified MITAX score and type 1 IFN gene composite score in the blood. (a) Comparisons between modified MITAX score and the type 1 IFN-inducible gene composite score over time for seven patients (BGE15, BGE92, BGE99, BGE10, BGE106, BGE119 and BGE147) with at least four total visits. (b) An additional correlation is seen for a PM patient, BGE92 between the type 1 IFN-inducible gene composite score and creatine kinase levels across time. Even though the disease activity of the patient increased substantially, the muscle disease MITAX subscore was already at a maximal value (9 points).
Several single type 1 IFN-inducible gene transcripts have potential clinical utility to monitor disease activity
In both the previous cross-sectional study23 and the current longitudinal study, among approximately 20 000 unique genes studied, two of the highest differentially expressed type 1 IFN-inducible transcripts in DM and PM blood were IFI44L (mean of 117-fold of normal) and RSAD2 (58-fold). Quantitative real-time PCR transcript measurements correlated highly with the microarray results in whole-blood samples from 15 patients (nine DM and six PM) for IFI44L (r=0.98) and RSAD2 (r=0.98).
These single gene transcript biomarkers correlated with disease activity comparably to the 13-gene composite score (Figure 4), suggesting that they have potential clinical utility as individual blood biomarkers to monitor disease activity in patients with DM or PM.
Discussion
Cytokines have a significant role in regulating immune responses in inflammatory and autoimmune diseases. Targeting cytokines has clinical efficacy in patients with psoriasis and rheumatoid arthritis.26, 27, 28, 29 Cytokine gene signatures have been utilized to address various issues from disease pathogenesis to drug effect in multiple diseases from various studies reported in the literature. The type 1 IFN-inducible blood gene signature is neutralized in SLE patients by an anti-IFN-α monoclonal antibody, and such neutralization is related to improvement in disease activity (that is, clearance of skin rash).30 An IL-17-inducible gene signature has been used to evaluate the effect of cyclosporin A in psoriasis.31 An IL-13-inducible gene signature is used to evaluate the activation of IL-13 pathway in the airway epithelial cells of asthma patients and also treatment effect of corticosteroids in these same patients.32 Gene expression profiling using whole-genome arrays in longitudinal studies allows tracking of multiple cytokine activities and their signaling pathways at the molecular level, and study of their correlation with disease activities during treatment. These studies may lead to the identification of new therapeutic targets in patients with systemic autoimmune diseases, as demonstrated by the discovery of type 1 IFN as a potential therapeutic target for SLE.24, 25
In this study, we developed a 13-gene composite score to capture the activation of the type 1 IFN signaling pathway in the blood of patients with DM or PM, and found it strongly correlated with disease activity as measured by modified MITAX score in a longitudinal study. First, only the type 1 IFN signature is statistically different in all comparisons of: normal healthy donors and patients with low and moderate disease activity; normal healthy donors and patients with high disease activity; patients with low/moderate and high disease activity. Secondly, for the 18 type 1 IFN-signature-positive patients, a strong correlation was observed between changes in disease activity and changes in the type 1 IFN signature score. Finally, of all 16 major changes in disease activity in individual patients as defined by a change in modified MITAX score of >3 observed in the 80 visits, there are concordant changes in disease activity and type 1 IFN signature score for all events. We also evaluated TNF-α, IL-1β, IL-10, IL-13 and GM-CSF signaling pathways in these patients during the longitudinal study. Although some of these pathways (GM-CSF, TNF-α and IL1-β) are activated in the blood of patients with DM or PM, (1) their cytokine-inducible gene signature scores did not show any major correlation with disease activity; and (2) the changes of their cytokine signature scores and changes in the disease activity of these patients also showed no correlation. These findings provide strong support to the hypotheses that type 1 IFN has a mechanistic role in the pathogenesis of DM and PM.22, 33, 34 Although the sample size is modest (24 patients are enrolled in the longitudinal study, and 21 from the 24 patients had elevation of type 1 IFN signature during the course of the study) in the study, due to the low incidence of patients with DM or PM, it still represents a longitudinal study of such nature with most patients enrolled where data are available publicly. That said, the conclusions will be strengthened in another independent study or ideally in a clinical trial that evaluates anti-type 1 IFN therapy in patients with DM or PM.
These findings also support the use of gene transcript measurements to provide value in monitoring disease activity in patients with DM or PM. These measurements are analogous to serum creatine kinase, in that although they may not reflect absolute levels of disease activity across patients, they appear to reflect changes in disease activity within individual patients. They may be of particular value in helping to distinguish IBM from PM, for which it is frequently initially misdiagnosed.
There are several implications of these data for drug development. The significant overexpression of type 1 IFN-inducible genes in the blood of patients with DM or PM is likely to provide an effective pharmacodynamic marker for assessing anti-type 1 IFN monoclonal antibody therapy in these patients, as in the case of SLE.27 High correlation of significant expression changes in type 1 IFN-inducible genes in peripheral blood with major changes in disease activity additionally suggests that their measurement has potential utility to monitor disease activity of patients with DM or PM during treatment courses.
An important limitation of this study is its performance in the context of routine clinical care. Patients were seen for routine clinical visits, and disease activity was measured retrospectively from clinical notes, blinded to gene expression data. Blood collection proceeded as clinically allowed, not at equal interval time-points according to a predetermined protocol. Future studies using prospective collection of blood samples and disease activity measures would be preferred.
Materials and methods
Patients
In all, 24 patients with inflammatory myopathies (15 with DM and 9 with PM) were enrolled from a single hospital center into a longitudinal exploratory study of the association between gene expression in blood samples and clinical features including disease activity. Clinical and gene expression data from 80 patient-visits were collected over a period of up to 6 years (mean period of participation was 1.9 years). In detail, 2, 2, 3, 7 and 10 patients had blood samples collected for seven, six, four, three and two visits, respectively. Clinical features are summarized in Supplementary Table 1. Early cross-sectional studies of some of these patients after one or two visits were previously reported;23 the current study increased the number of enrolled patients and performed a longitudinal analysis of up to seven visits per patient (mean of 3.3 visits). All patients satisfied widely used descriptive criteria for DM or PM26 as well as more stringent research criteria.3 Normal blood samples were studied from 36 healthy donors.
Disease activity assessment
At each visit, whole-blood samples were collected for gene expression studies and clinical data were collected that allowed for subsequent assessment of disease activity using three individual system subsections (constitutional, cutaneous and muscle disease activities) of the MITAX. Using the MITAX subsection scores as described,23 0 points were given for inactive, 1 point for stable mild disease, 3 points for moderately active disease and 9 points for active disease requiring disease-modifying treatment, for each of the three subsections, and the values were summed for a total score. Disease activity variation was defined as a change in this modified MITAX score, or in cases where the modified MITAX score was already maximally saturated, a change in serum creatine kinase (for example, an increase in muscle disease activity is not captured by MITAX for a patient with 9 points for muscle disease activity whose muscle disease worsens further).
Total RNA extraction, microarray assays and TaqMan quantitative real-time PCR assays
RNA was isolated from whole blood using PAXgene (PreAnalytix GmbH, Hombrechtikon, Switzerland) tubes or buffy coat samples. Affymetrix U133 Plus 2.0 microarray assays (Affymetrix, Santa Clara, CA, USA) were carried out as previously described.23 Fluidigm's Biomark system dynamic array platform (Fluidigm Corp., South San Francisco, CA, USA) was used to measure expression levels of a panel of selected immune response genes in the blood of 15 patients with DM or PM.
Construction of cytokine-inducible gene expression sets and gene scores
We constructed gene sets reflective of cytokine-inducible transcripts for type 1 IFN and the five other cytokines GM-CSF, IL-10, IL-13, IL-1β or TNF-α. Type 1 IFN-inducible genes are well characterized, and we defined this gene signature empirically from patient used in the study following procedures similarly to previously described for SLE.30, 35 In short, transcript profiling data were generated from the first visit of the 24 DM or PM patients enrolled in the longitudinal study in combination with a second cohort of 18 DM or PM patients screened for enrollment in a clinical trial of anti-IFN-α therapy for myositis.36 All 42 patients satisfied widely used descriptive criteria for DM or PM.37 In all, 13 overexpressed type 1 IFN-inducible genes (from the 100 most overexpressed genes; Supplementary Table 2) were selected to construct a type 1 IFN signature as follows: the six most overexpressed type 1 IFN-inducible genes in the blood of DM and PM patients (IFI27, RSAD2, IFI44L, IFI44, OAS1 and IFIT1), plus ISG15, one of the most overexpressed type 1 IFN-inducible genes in DM muscle specimens,22, 23 and six well-characterized type 1 IFN-inducible genes (OAS3, HERC5, MX1, ESPTI1, IFIT3 and IFI6). Gene signature scores with this 13-gene panel (a subset of the 21-gene panel for SLE) correlated well with scores used to characterize type 1 IFN signaling pathway activation in SLE, using both the 21-gene panel (r=0.98) and a dynamic top 25 most overexpressed type 1 IFN-inducible gene approach in individual patients (r=0.82).30, 35
Because there are no generally accepted gene expression signatures for the other cytokines used in this study, we generated empirically defined 13-gene set signatures for them as follows: whole blood collected from four healthy donors was exposed to either GM-CSF, IL-10, IL-13, IL-1β or TNF-α (R&D Systems, Minneapolis, MN, USA) at concentrations of 3 × EC50, vehicle (1 × phosphate-buffered saline) control, cytokine+control antibody (R347, 10 μg ml−1) or cytokine+cytokine-specific neutralizing monoclonal antibody (R&D Systems, 10 μg ml−1), as described.38 Microarray experiments performed on extracted RNA were used to identify for each cytokine the 13 most highly overexpressed (average fold change >2) and cytokine-specific antibody-neutralized uniquely annotated genes (Supplementary Table 3). An average signal intensity neutralization requirement of greater than 75% was implemented for all cytokine+cytokine-specific antibody conditions compared with cytokine stimulation alone, with the exception of IL-13, for inclusion in the final 13-gene signature panels. Due to the relatively weaker neutralization by the anti-IL-13 antibody, the gene selection methodology was modified to allow for inclusion of genes displaying neutralization less than 50% for compiling an IL-13 gene signature.
For each of the 24 patients and for each cytokine, we computed a gene expression score as the median fold change compared with the reference pool of 36 healthy donors for the 13-gene cytokine-specific-inducible gene set.
Microarray data analysis
Microarray data analysis was performed as previously described,30, 35 using GC-Content Robust Multichip Analysis (GCRMA), which was implemented in the Bioconductor (http://www.bioconductor.org/) GCRMA package. All two-group comparisons involving the gene signature scores were calculated using a two-sample Wilcoxon–Mann–Whitney test, while paired comparisons were calculated using a Wilcoxon signed-rank test. All P-values reported are unadjusted for multiple testing. For identification of patients with a positive type 1 IFN-inducible gene signature score, we calculated significance analysis of microarrays with false discovery rate adjustment in R (R Development Core Team, University of Auckland, New Zealand). Pearson's correlation coefficients are provided for all tests of association between continuous variables with adequate sample sizes. The Spearman's rank rho statistic was also calculated to provide any advantage in a non-parametric summary, and similar results of association were observed. The small sample sizes in these correlation tests prohibit accurate interpretation of P-values, so they were not calculated in these cases
A linear mixed model for a longitudinal analysis
A repeated measures analysis was conducted for only those patients with at least three follow-up visits (total N=53 follow-up visits for 12 patients). Six different regression models were fit where a binary coding of modified MITAX score (low and high disease activity as defined previously) was regressed on each of the six cytokine-inducible gene signature scores, all modeled as random effects. Treatment status at the initial visit was used as a binary covariate (that is, any treatment/no treatment) and modeled as a fixed effect in each model in order to control for the influence of medication on gene expression changes.
References
Mastaglia FL, Garlepp MJ, Phillips BA, Zilko PJ . Inflammatory myopathies: clinical, diagnostic and therapeutic aspects. Muscle Nerve 2003; 27: 407–425.
Christopher-Stine L, Plotz PH . Adult inflammatory myopathies. Best Pract Res Clin Rheumatol 2004; 18: 331–344.
Hoogendijk JE, Amato AA, Lecky BR, Choy EH, Lundberg IE, Rose MR et al. 119th ENMC international workshop: trial design in adult idiopathic inflammatory myopathies, with the exception of inclusion body myositis, 10–12 October 2003, Naarden, The Netherlands. Neuromuscul Disord 2004; 14: 337–345.
Amato AA, Griggs RC . Treatment of idiopathic inflammatory myopathies. Curr Opin Neurol 2003; 16: 569–575.
Choy EH, Hoogendijk JE, Lecky B, Winer JB, Gordon P . Withdrawn: immunosuppressant and immunomodulatory treatment for dermatomyositis and polymyositis. Cochrane Database Syst Rev 2009; 4: CD003643.
Isenberg DA, Rowe D, Shearer M, Novick D, Beverley PC . Localization of interferons and interleukin 2 in polymyositis and muscular dystrophy. Clin Exp Immunol 1986; 63: 450–458.
Emslie-Smith AM, Arahata K, Engel AG . Major histocompatibility complex class I antigen expression, immunolocalization of interferon subtypes, and T cell-mediated cytotoxicity in myopathies. Hum Pathol 1989; 20: 224–231.
Lundberg I, Brengman JM, Engel AG . Analysis of cytokine expression in muscle in inflammatory myopathies, Duchenne dystrophy, and non-weak controls. J Neuroimmunol 1995; 63: 9–16.
Figarella-Branger D, Civatte M, Bartoli C, Pellissier JF . Cytokines, chemokines, and cell adhesion molecules in inflammatory myopathies. Muscle Nerve 2003; 28: 659–682.
Salomonsson S, Lundberg IE . Cytokines in idiopathic inflammatory myopathies. Autoimmunity 2006; 39: 177–190.
Banker BQ . Dermatomyostis of childhood, ultrastructural alteratious of muscle and intramuscular blood vessels. J Neuropathol Exp Neurol 1975; 34: 46–75.
Norton WL, Velayos E, Robison L . Endothelial inclusions in dermatomyositis. Ann Rheum Dis 1970; 29: 67–72.
Grimley PM, Davis GL, Kang YH, Dooley JS, Strohmaier J, Hoofnagle JH . Tubuloreticular inclusions in peripheral blood mononuclear cells related to systemic therapy with alpha-interferon. Lab Invest 1985; 52: 638–649.
Rich SA, Owens TR, Bartholomew LE, Gutterman JU . Immune interferon does not stimulate formation of alpha and beta interferon induced human lupus-type inclusions. Lancet 1983; 1: 127–128.
Grimley PM, Rutherford MN, Kang YH, Williams T, Woody JN, Silverman RH . Formation of tubuloreticular inclusions in human lymphoma cells compared to the induction of 2′-5′-oligoadenylate synthetase by leucocyte interferon in dose-effect and kinetic studies. Cancer Res 1984; 44: 3480–3488.
Kuyama J, Kanayama Y, Mizutani H, Katagiri S, Tamaki T, Yonezawa T et al. Formation of tubuloreticular inclusions in mitogen-stimulated human lymphocyte cultures by endogenous or exogenous alpha-interferon. Ultrastruct Pathol 1986; 10: 77–85.
Feldman D, Goldstein AL, Cox DC, Grimley PM . Cultured human endothelial cells treated with recombinant leukocyte A interferon. Tubuloreticular inclusion formation, antiproliferative effect, and 2′,5′ oligoadenylate synthetase induction. Lab Invest 1988; 58: 584–589.
Greenberg SA, Sanoudou D, Haslett JN, Kohane IS, Kunkel LM, Beggs AH et al. Molecular profiles of inflammatory myopathies. Neurology 2002; 59: 1170–1182.
Raju R, Dalakas MC . Gene expression profile in the muscles of patients with inflammatory myopathies: effect of therapy with IVIg and biological validation of clinically relevant genes. Brain 2005; 128 (Part 8): 1887–1896.
Tezak Z, Hoffman EP, Lutz JL, Fedczyna TO, Stephan D, Bremer EG et al. Gene expression profiling in DQA1*0501+ children with untreated dermatomyositis: a novel model of pathogenesis. J Immunol 2002; 168: 4154–4163.
Greenberg SA . A gene expression approach to study perturbed pathways in myositis. Curr Opin Rheumatol 2007; 19: 536–541.
Greenberg SA, Pinkus JL, Pinkus GS, Burleson T, Sanoudou D, Tawil R et al. Interferon-alpha/beta-mediated innate immune mechanisms in dermatomyositis. Ann Neurol 2005; 57: 664–678.
Walsh RJ, Kong SW, Yao Y, Jallal B, Kiener PA, Pinkus JL et al. Type I interferon-inducible gene expression in blood is present and reflects disease activity in dermatomyositis and polymyositis. Arthritis Rheum 2007; 56: 3784–3792.
Baechler EC, Batliwalla FM, Karypis G, Gaffney PM, Ortmann WA, Espe KJ et al. Interferon-inducible gene expression signature in peripheral blood cells of patients with severe lupus. Proc Natl Acad Sci USA 2003; 100: 2610–2615.
Bennett L, Palucka AK, Arce E, Cantrell V, Borvak J, Banchereau J et al. Interferon and granulopoiesis signatures in systemic lupus erythematosus blood. J Exp Med 2003; 197: 711–723.
Bongartz T, Sutton AJ, Sweeting MJ, Buchan I, Matteson EL, Montori V . Anti-TNF antibody therapy in rheumatoid arthritis and the risk of serious infections and malignancies: systematic review and meta-analysis of rare harmful effects in randomized controlled trials. Jama 2006; 295: 2275–2285.
Brimhall AK, King LN, Licciardone JC, Jacobe H, Menter A . Safety and efficacy of alefacept, efalizumab, etanercept and infliximab in treating moderate to severe plaque psoriasis: a meta-analysis of randomized controlled trials. Br J Dermatol 2008; 159: 274–285.
Ding C, Xu J, Li J . ABT-874, a fully human monoclonal anti-IL-12/IL-23 antibody for the potential treatment of autoimmune diseases. Curr Opin Investig Drugs 2008; 9: 515–522.
Kavanaugh A . Interleukin-6 inhibition and clinical efficacy in rheumatoid arthritis treatment—data from randomized clinical trials. Bull NYU Hosp Jt Dis 2007; 65 (Suppl 1): S16–S20.
Yao Y, Richman L, Higgs BW, Morehouse C, De los Reyes M, Brohawn P et al. Neutralization of IFN-alpha/beta-inducible genes and downstream effect in a phase I trial of an anti-IFN-alpa monoclonal antibody in SLE. Arthritis Rheum 2009; 60: 1785–1796.
Haider AS, Lowes MA, Suarez-Farinas M, Zaba LC, Cardinale I, Khatcherian A et al. Identification of cellular pathways of ‘type 1,’ Th17T cells, and TNF- and inducible nitric oxide synthase-producing dendritic cells in autoimmune inflammation through pharmacogenomic study of cyclosporine A in psoriasis. J Immunol 2008; 180: 1913–1920.
Woodruff PG, Boushey HA, Dolganov GM, Barker CS, Yang YH, Donnelly S et al. Genome-wide profiling identifies epithelial cell genes associated with asthma and with treatment response to corticosteroids. Proc Natl Acad Sci USA 2007; 104: 15858–15863.
Greenberg SA . Type 1 interferons and myositis. Arthritis Res Ther 2010; 12 (Suppl 1): S4.
Salajegheh M, Kong SW, Pinkus JL, Walsh RJ, Liao A, Nazareno R et al. Interferon-stimulated gene 15 (ISG15) conjugates proteins in dermatomyositis muscle with perifascicular atrophy. Ann Neurol 2010; 67: 53–63.
Yao Y, Higgs BW, Morehouse C, de Los Reyes M, Trigona W, Brohawn P et al. Development of potential pharmacodynamic and diagnostic markers for anti-IFN-α monoclonal antibody trials in systemic lupus erythematosus. Hum Genomics Proteomics 2009; 2009: pii: 374312.
A phase 1b, randomized, double-blind, placebo-controlled, multicenter study to evaluate safety of multiple-dose, intravenously administered MEDI-545, a fully human anti Interferon-alpha monoclonal antibody, in adult patients with dermatomyositis or polymyositis. http://clinicaltrialsgov/ct2/show/NCT0053309.
Bohan A, Peter JB . Polymyositis and dermatomyositis (first of two parts). N Engl J Med 1975; 292: 344–347.
Yao Y, Richman L, Morehouse C, de los Reyes M, Higgs BW, Boutrin A et al. Type I interferon: potential therapeutic target for psoriasis? PLoS One 2008; 3: e2737.
Acknowledgements
We would like to thank Melissa de los Reyes, Jiaqi Huang and Jianliang Zhang for providing technical assistance; Robert Georgantas III and Tom Hollon for providing editing and critical review of the manuscript.
SAG was supported by Muscular Dystrophy Association MDA3523. Some microarray-based gene expression profiling was performed by the Harvard Neuromuscular Disease Project Core Microarray laboratory supported by NIAMS P50NS40828, and some profiling was performed at MedImmune laboratories, supported by MedImmune, LLC.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The following authors are full-time employees of MedImmune: Brandon W. Higgs, Chris Morehouse, Philip Brohawn, Wei Zhu, Peter A. Kiener, Bahija Jallal, and Yihong Yao. MedImmune is developing anti-type 1 interferon therapy for autoimmune diseases:
Steven A. Greenberg and Anthony A Amato have served as consultants for MedImmune. The Brigham and Women's Hospital manages research sponsored by MedImmune and intellectual property pertaining to myositis diagnostics.
Additional information
Supplementary Information accompanies the paper on Genes and Immunity website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Greenberg, S., Higgs, B., Morehouse, C. et al. Relationship between disease activity and type 1 interferon- and other cytokine-inducible gene expression in blood in dermatomyositis and polymyositis. Genes Immun 13, 207–213 (2012). https://doi.org/10.1038/gene.2011.61
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2011.61
Keywords
This article is cited by
-
A case of autoimmune pulmonary alveolar proteinosis during the course of treatment of rapidly progressive interstitial pneumonia associated with anti-MDA5 antibody-positive dermatomyositis
BMC Pulmonary Medicine (2024)
-
Stromal vascular fraction in the treatment of myositis
Cell Death Discovery (2023)
-
Long-term safety of COVID vaccination in individuals with idiopathic inflammatory myopathies: results from the COVAD study
Rheumatology International (2023)
-
Update on Biomarkers of Vasculopathy in Juvenile and Adult Myositis
Current Rheumatology Reports (2022)
-
JAK inhibitors: a potential treatment for JDM in the context of the role of interferon-driven pathology
Pediatric Rheumatology (2021)