Abstract
The abnormal expansion of a ≥36 CAG unit tract in the Huntingtin gene (HTT) leads to Huntington's disease (HD), but has also been associated with cancer: the incidence of cancer is lower in HD patients than in age-matched controls, but HD-causing variants of HTT accelerate the progression of breast tumors and the development of metastases in mouse models of breast cancer. To investigate the relationship between HTT CAGs and cancer, data concerning 2407 women with BRCA1 or BRCA2 mutations that predispose to breast and ovarian cancers and 431 patients with breast cancer without family histories were studied; the size of the CAG expansions on both HTT alleles was determined in each subject. The proportion of individuals carrying a CAG expansion in a pathological range for HD was 10 times more frequent than previously reported in the literature. In carriers of BRCA2 mutations, the length of the HTT CAG tract was correlated with lower incidence of ovarian cancer. Among carriers of BRCA1 mutations who developed a breast cancer, its onset occurred 2.4 years earlier in individuals with intermediate HTT alleles (≥27) than in those with a CAG tract <27. Finally, in patients with sporadic HER2 breast cancer, metastasis increased by a factor of 11.10 per 10 additional CAG repeats in HTT. We concluded that whereas long CAG length could be associated with lower cancer incidence, it could also be paradoxically associated with cancer severity (age of apparition and metastasis development).
Similar content being viewed by others
Introduction
The Huntingtin (HTT) gene has mostly been studied in the context of the neurodegenerative disorder Huntington’s disease (HD). HTT carries a polymorphic trinucleotide repeat of CAG in exon 1, encoding a polyglutamine stretch in the huntingtin protein (HTT).1 In the general population, the CAG expansion ranges from 9 to 35 repeats with an average between 17 and 20 repeats. An expansion exceeding 35 results in HD.2, 3, 4 Age at symptomatic disease is negatively correlated with the CAG tract size, the longer the repeat, the earlier the disease onset. Although for individuals with 40 or more CAG repeats, the expansion is necessary and sufficient to cause HD, carriers of 36–39 CAG repeats may have incomplete penetrance or later onset. Intermediate alleles between 27 and 35 CAG repeats were originally thought to have no clinical implications. However, this view has been recently challenged5 and evidence shows that patients with an intermediate number of repeats could manifest some aspects of the disease.6
Two studies reported that the incidence of cancer is lower in patients with HD than in aged-matched controls.7, 8 On the other hand, HD-causing variants of HTT accelerate the progression of breast tumors and the development of metastasis in mouse models of breast cancer.9 Thus, whereas the incidence of cancer may be low among HD patients, a large number of CAG repeats might be an aggravating factor when a cancer is already established. However, it is not known whether the size of the CAG tract <36 repeats influences breast cancer incidence and progression in the non-HD population.
Most of breast cancer are sporadic and involved a multitude of risk factors.10 A small proportion of breast cancers are caused by one of several transmitted mutations, including germline mutations in BRCA1 and BRCA2, which confer a high risk of breast and ovarian cancer.11 In this study, we first investigated if HTT CAG repeats length is a BRCA1/2 cancer modifier by analyzing the association between the length of HTT CAG repeat and breast and ovarian cancer incidence or age of cancer diagnosis. Furthermore, in a second cohort of patients with breast cancer of sporadic origin, we explored the associations between the length of CAG tract and breast cancer prognosis.
Subjects and methods
Patients
This study used information from 2838 individuals, including 2407 women with hereditary BRCA1/2 mutations (cohort 1) and 431 women with sporadic breast cancer (cohort 2).
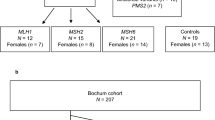
Cohort 1 (Figure 1): individuals with BRCA1 and BRCA2 mutations were recruited through Genetic Modifiers of Cancer Risk in BRCA1/2 Mutation Carriers (GEMO), a cohort of women carrying BRCA1 or BRCA2 germline mutations. GEMO is the French branch of the international initiative CIMBA (Consortium of Investigators of Modifiers of BRCA1 and BRCA2), which aims at identifying modifiers of breast cancer risk through the collection of DNA and clinical data from a large number of BRCA1 and BRCA2 mutation carriers.12, 13, 14 Participation in the GEMO cohort was proposed during the cancer clinic in which patients were informed of a positive BRCA1/2 test result. Their written informed consent was obtained. In total, 2407 women with hereditary BRCA1 (1608, 67%) or BRCA2 (779, 33%) mutations were identified. Among the 1746 patients who developed a cancer (1138 BRCA1, 71%; 608 BRCA2, 76%), 1353 had breast cancer (807 BRCA1, 71%; 546 BRCA2, 90%), 254 had ovarian cancer (207 BRCA1, 18%; 47 BRCA2, 8%) and 139 had both breast and ovarian cancer (124 BRCA1, 11%; 15 BRCA2, 2%) (Figure 1).
Cohort 2: informations about women with breast cancer of sporadic origin were collected (Supplementary Table S1). Four hundred and thirty-one consecutive primary unilateral invasive primary breast tumors were excised from women at the Institut Curie/Hôpital René Huguenin (France) between 1978 and 2008.15, 16 Immediately following surgery, the samples were stored in liquid nitrogen until RNA extraction. Samples were considered suitable if the proportion of tumor cells, determined histologically, was >70%. The patients met the following criteria: unilateral non-metastatic primary breast carcinoma for which complete clinical, histological and biological data were available; no radiotherapy or chemotherapy before surgery; and full follow-up at the Institut Curie/Hôpital René Huguenin. Patients underwent physical examinations and routine chest radiography every 3 months for 2 years, and then annually. Mammograms were performed annually. The histological type and the number of positive axillary nodes were established at the time of surgery. The malignancy of infiltrating carcinomas was scored according to the Scarff–Bloom–Richardson (SBR) histoprognostic system. Estrogen receptor (ER), progesterone receptor (PR) and HER2 status were determined from protein content by biochemical methods (dextran-coated charcoal method, enzymatic immuno-assay or immunohistochemistry) and confirmed by real-time quantitative RT-PCR assays of ERα, PR and HER2. We subdivided our total population (n=431) into four sub-groups: the ‘luminal’ subtype expressed hormone receptors (HR; ER+ or PR+) and showed no amplification of HER2 (n=275); the ‘luminal B/HER2+’ subtype expressed HR (ER+ or PR+) and overexpressed the HER2 receptor (n=50); the ‘HER2+’ subtype did not express HR but was positive for HER2 (ER− and PR−/HER2+, n=42); and the ‘triple negative’ subtype was negative for HR and for HER2 overexpression (ER− and PR−/HER2−, n=64). Standard prognostic factors for these tumors are presented in Supplementary Table S1. The median of follow-up was 8.4 years (range 4 months to 29 years). All patients admitted before 2007 were informed that their tumor samples might be used for breast cancer progression studies and they were given the opportunity to refuse the use of their samples. Since 2007, patients have also given their approval by signing an informed consent form. This study was approved by the local ethic committee (Breast Group of René Huguenin Hospital).
For both cohorts, only women with available genotyping information about HTT CAG length were included in the analysis. All samples and corresponding data were encoded for anonymization.
Determination of HTT CAG length
Cohort 1: individuals provided blood samples for BRCA1/2 genetic testing. DNA was extracted with classical protocols that may have differed among diagnostic laboratories. Cohort 2: total RNA was extracted from biopsies with the acid–phenol guanidinium method. All samples were sent to GenoScreen, France (www.genoscreen.com). PCR was performed in 25 μl reactions containing 20 ng of template DNA or cDNA, 1 × reaction buffer, 37.5 pmol MgCl2, 6 pmol dNTP, 10 pmol fluorescent primer, 10 pmol primer and 1 U Taq polymerase (FastStart–Roche Diagnostics, Pleasanton, CA, USA).
Primers sequences were: 5′-TGGCCCGGTGCTGAG3-′ (forward) and 5′-CGGTGGCGGCTGTTG3-′ (reverse). The reverse primer was located immediately after the CAG expansion: the fragments analyzed did not contain the variable flanking CCG repeats. The PCR cycle consisted of an initial denaturation step at 95 °C for 10 min, followed by 40 cycles of denaturation at 95 °C for 30 s, annealing at 62 °C for 30 s and extension at 72 °C for 1 min, and a final extension at 72 °C for 10 min. Each amplification product was mixed with Hi-Di Formamide and GeneScan 500 LIZ Size Standard (Applied Biosystems, Foster City, CA, USA). Fragments were separated on an Applied Biosystems 3730XL DNA Analyzer. Alleles were scored with GeneMapper v4.0 software (Applied Biosystems).
Statistical analysis
The allele containing the most CAG repeats was designated the ‘long’ allele, the other was termed the ‘short’ allele. The two alleles were analyzed individually in each patients. Intermediate alleles exceeding 26 CAG repeats and pathologically expanded alleles exceeding 35 CAG repeats were distinguished.
The association between the size of HTT CAG tract and breast and/or ovarian cancer incidence in BRCA1 and BRCA2 mutation carriers was investigated with a retrospective likelihood approach of the observed genotypes conditional on the disease phenotype. This method17, 18 adjusts for the fact that BRCA1/2 mutation carriers were not randomly sampled with respect to their disease phenotype. These models, primarily developed for categorical covariates, were used when the size of the HTT allele was considered as a continuous covariate (short and long alleles). Both single disease risk and competing risk models were implemented in a Fortran program.
We first analyzed the association of HTT CAG repeat length with either breast or ovarian cancer. For the breast cancer risk association analysis, patients were censored at their age when the first ovarian cancer was detected, bilateral prophylactic mastectomy performed or age at last observation. Only individuals with breast cancer were considered as affected; the remaining BRCA1/2 mutation carriers were assumed to be unaffected by the disease. For the ovarian cancer risk association analysis, patients were censored at the age when bilateral oophorectomy was performed or at the age of last observation. Only individuals with ovarian cancer were considered as affected; the remaining BRCA1/2 mutation carriers were assumed to be unaffected by the disease.
We then analyzed the association of HTT CAG length with both breast and ovarian cancer, because BRCA1/2 mutation carriers are at risk of developing both of these diseases. We used a competing risk analysis and estimated simultaneously hazard ratios for both breast and ovarian cancer. In this method,17 patients were followed up to the age when breast or ovarian cancer was diagnosed, whichever occurred first, and were assumed to be affected with the corresponding disease. Patients were censored at the age when bilateral prophylactic mastectomy was performed for breast cancer, bilateral oophorectomy for ovarian cancer or at the age of last observation and were considered to be unaffected by both diseases. When patients developed breast and ovarian cancers at the same age, they were considered to be ovarian cancer cases. ANOVA was used with an empirical sandwich estimator to take into account the effect of family to analyze the association between the age when BRCA1 or BRCA2 cancer was diagnosed and HTT CAG length.
Pearson correlation tests were used to investigate the association between HTT CAG tract size and age at onset of sporadic breast cancer. ANOVA was used, followed by pairwise comparisons with Tukey–Kramer adjustment of P-values for significant differences to compare HTT CAG tract sizes between categories of qualitative variables. The following variables were tested: macroscopic tumor size (≤25 or >25 mm), ER, PR and HER2 receptor status (positive or negative), SBR histological grade (I, II or III), lymph node status (0, 1–3 or >3) and molecular subtypes. χ2-tests were performed to evaluate whether specific clinico-pathological characteristics were more frequent in patients with ≥27 CAG in one HTT allele than in patients with <27 CAG in both HTT alleles.
Metastasis-free survival (MFS) was defined as the interval of time between surgery and detection of the first metastasis in patients who developed a metastatic disease and the interval between surgery and last checkup for patients without metastases. Cox proportional hazard univariate analysis followed by multivariate models with a stepwise procedure were used to evaluate risk factors for the development of metastases in the whole cohort and in each molecular subtype of breast cancer. The following variables were included in the whole cohort analysis: age when cancer was diagnosed, macroscopic tumor size, tumor grade, lymph node status, ER, PR and HER2 status and consequently, molecular subtype, length of the CAG tract on both HTT alleles and the presence of ≥27 CAG on one HTT allele. For the analysis of each molecular subtype, all variables were included except for ER, PR, HER2 status and molecular subtype. All the variables with a P-value <0.10 in univariate analysis were included in multivariate analysis.
All analyses were performed with SAS version 9.3. P-values of 0.05 and below were considered significant. All data are available in the data set file.
Results
HTT CAG length among cancer patients
The distribution of the HTT CAG repeat number in both cohorts 1 and 2 was comparable to that described in the general population worldwide19 with a mean number of repetitions of 20 on the long allele and 17 on the short allele (Figure 2).
Among the 2838 individuals in the two cohorts, five patients carried a pathological CAG expansion of 36 repeats or more on one allele of the HTT gene (Table 1). The overall prevalence of patients carrying a HTT allele with a CAG repeat equal or over 36 was therefore high at ~1:568 (0.18%); higher than previous estimates (0.008–0.017% reported in ref. 20, 21, 22, 23). Furthermore, we found a similar proportion of carriers of HD-causing CAG expansions in an independent cohort affected with inherited ataxia (1:541; EUROSCA cohort; data not shown), suggesting that this unexpected high prevalence is not specific to cancer population.
Association between HTT CAG length and BRCA1/2 cancer
We then analyzed the association between HTT CAG tract size and breast and ovarian cancer incidence in cohort 1. In BRCA1 mutation carriers, HTT CAG repeat length was not associated with the incidence of breast or ovarian cancer, whether the analyses were performed separately or simultaneously for these cancers (Table 2).
A similar result was obtained for breast cancer in BRCA2 mutation carriers. However, a large number of CAG repeats in HTT was correlated with lower BRCA2 ovarian cancer incidence in single (long allele: HR 0.35 95% CI 0.14–0.87; P=0.0245) and competing analyses (short allele: HR 0.24 95% CI 0.06–0.92; P=0.037).
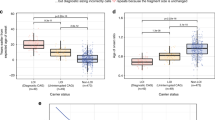
Next, we investigated whether HTT CAG length was associated with age at onset among BRCA1 or BRCA2 carriers who developed a cancer. BRCA1 carriers developed breast cancer at 41.0±9.5 years of age on average and BRCA2 at 43.3±10.1. BRCA1 carriers with a CAG repeat number ≥27 on the long HTT allele developed cancer 2.43±1.06 years earlier than patients with HTT CAG lengths under 27 (P=0.0235; Table 3). In the BRCA2 population, we observed the same trend although the relationship was not significant (P=0.0900; Table 3). These effects were specific to breast cancer as we did not observed associations between HTT CAG length and the age at ovarian cancer onset in BRCA1/2 carriers. Similar results were obtained when the four patients carrying a pathological HD CAG repeat ≥36 in this cohort were excluded (data not shown).
Associations between HTT CAG length and sporadic breast cancer
In cohort 2, there were no significant associations between the size of the HTT CAG expansion and any of the clinic-pathological variables studied (Supplementary Table S1). Results obtained in patients carrying expansions with ≥27 CAGs were comparable to those obtained for patients with <27 CAG repeats (Supplementary Table S1).
We then analyzed patient MFS in the population of cohort 2 as a whole (Supplementary Table S2) as well as in each breast cancer molecular subtype (Table 4 and Supplementary Table S3) with univariate and multivariate Cox proportional hazards models. At last follow-up, 160 patients had developed metastasis (37%) and classical prognostic factors influenced MFS progression (Supplementary Tables S2 and S3 and Table 4). In the whole population, the CAG size in the HTT short allele emerged as a predictor of MFS with tumor size, SBR grade II and III and lymph node invasion after multivariate analysis (HR 2.19, 95% CI 1.01–4.73; P=0.0469; Supplementary Table S2). We therefore analyzed patient MFS in each breast cancer molecular subtype (Table 4 for HER2+ and Supplementary Table S3 for other molecular subtypes). In the HER2+ subtype specifically, univariate analysis revealed a positive correlation between the length of HTT CAGs on the long allele and the risk of metastasis, which increased by a factor of 11.10 for every 10 additional CAG repeats (95% CI 2.09–58.64; P=0.0046; Table 4). Furthermore, the length of the CAG repeat in the long HTT allele was the only prognostic factor of MFS after adjustment for lymph node invasion by multivariate analysis.
Discussion
Two studies have reported a low incidence of cancer in polyglutamine disorders without taking into account the number of CAG repeats or cancer outcome.7, 8 Our data reveal an association between the length of the polymorphic CAG tract in wild-type HTT and cancer. The lower incidence of cancer in HD patients7, 8 is consistent with our results showing that long CAG repeats in HTT are associated with protection against ovarian cancer. In sporadic breast cancer patients, CAG repeat size was a strong independent factor of metastases, specifically in the HER2-positive subtype. Whereas the incidence of cancer may be inversely correlated to the CAG length, a large number of CAG repeats might enhance the progression of breast cancer once tumorigenesis has been initiated.
Our observations may be related to the biology of HTT, which is regulated according to the length of the polyglutamine sequence encoded by the CAG tract. Although HTT is widely expressed, the amount of the protein differs between tissues and cell types within the same tissue;24, 25 therefore, the length of the CAG expansion in HTT may differentially influence various types of cancer. Furthermore, HTT may be involved in particular cancer subtypes depending on the signaling pathways engaged in the oncogenic processes. For instance, the regulatory effect of HTT could differ in cancers of BRCA1 and BRCA2 origin, as HTT was reported to interact with BRCA1 -and not BRCA2- through the BRCA1-associated RING domain protein, BARD1.26 Loss of wild-type HTT in breast cancer promotes metastasis by altering tight junctions.27 Furthermore, HTT with a CAG repeat ≥36 leads to abnormalities in HER2 endocytosis in breast cancer cells, which affect cell motility and metastatic behavior thus promoting tumorigenesis and metastasis in HD mice.9 CAG tracts below 36 repeats may also regulate tight junctions maintenance and/or HTT-mediated endocytosis and affect breast cancer development.7, 8, 9, 27
This study has a number of limitations that may influence the generalizability and translational potential of this research. Individuals from cohort 1 were recruited as carriers of a BRCA1/2 mutation. This testing strategy may lead to underreporting of non-founder mutations. As such, some bias in the ascertainment of the full spectrum of mutations could have occurred. The retrospective likelihood approach allows correcting for this bias.17 For cohort 1, carriers of BRCA2 mutations composed a smaller sample set; in particular, the number of women with BRCA2-associated ovarian cancers was relatively small. The associations in cohort 1 did not remain statistically significant after corrections for multiple testing; therefore, we cannot exclude false positive and further studies are needed to investigate these findings. A further limitation of our study is based on the retrospective likelihood approach that relies on the availability of external disease incidences for the mutation carriers. A specific selection of control subjects within the same population of BRCA1/2 mutation carriers sampled in similar conditions as BRCA1 and BRCA2 mutation carriers would allow further clinicopathological comparison based on tumor characteristics. In addition, the sample size in the cohort 2 is limited regarding the sample size of the cohort 1 leading to a smaller power for this cohort. Furthermore, cohort 1 is a ‘prognostic’ cohort as BRCA1/2 mutation carriers are enrolled before the development of breast or/and ovarian cancer, whereas cohort 2 is a ‘prognosis’ cohort established >30 years ago that allows the investigation of long-term prognosis that is not possible in cohort 1 due to shorter follow-up.
Besides, we found a high prevalence of pathological HD CAG repeats in HTT compared to previous estimates of 8 and 17.3 per 100 000 worldwide.20, 21, 22, 23 Here, we estimated CAG repeat length in a cancer population with a priori no HD bias. Even if these allele sizes are associated with low penetrance, they are pathological and risk to expand further upon transmission.28 As we found a similar proportion of carriers of HD-causing CAG expansions in an independent cohort affected with inherited ataxia (1:541; EUROSCA cohort; data not shown), it is less expected that the high prevalence is specific to cancer population. Recent evidence indeed suggests that the prevalence of HD is underestimated: in British Columbia it was estimated at 13.7 per 100 000 in the general population and 17.3 per 100 000 in the Caucasian population.23 Studies investigating the prevalence of HD have used estimation models to extend observations from subpopulations of individuals at risk or from patients diagnosed with HD to the general population. Thus, there is a crucial need to decipher the prevalence of HD-causing variants in the general population without using estimation models.
We have described here associations between the size of the length of CAG tracts in HTT and features of breast and ovary cancer with potential implications for the follow-up of cancer patients. Future studies should address the molecular mechanisms underlying the specific regulatory effects of HTT CAG expansion on various types of cancer.
References
Group THsDCR: A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes: The Huntington's Disease Collaborative Research Group. Cell 1993; 72: 971–983.
Rubinsztein DC, Leggo J, Coles R, et al: Phenotypic characterization of individuals with 30-40 CAG repeats in the Huntington disease (HD) gene reveals HD cases with 36 repeats and apparently normal elderly individuals with 36-39 repeats. Am J Hum Genet 1996; 59: 16–22.
Andrew SE, Goldberg YP, Kremer B, et al: The relationship between trinucleotide (Cag) repeat length and clinical-features of Huntingtons-disease. Nat Genet 1993; 4: 398–403.
Snell RG, MacMillan JC, Cheadle JP, et al: Relationship between trinucleotide repeat expansion and phenotypic variation in Huntington's disease. Nat Genet 1993; 4: 393–397.
Semaka A, Hayden M : Evidence-based genetic counselling implications for Huntington disease intermediate allele predictive test results. Clin Genet 2013.
Killoran A, Biglan KM, Jankovic J, et al: Characterization of the Huntington intermediate CAG repeat expansion phenotype in PHAROS. Neurology 2013; 80: 2022–2027.
Ji J, Sundquist K, Sundquist J : Cancer incidence in patients with polyglutamine diseases: a population-based study in Sweden. Lancet Oncol 2012; 13: 642–648.
Sorensen SA, Fenger K, Olsen JH : Significantly lower incidence of cancer among patients with Huntington disease: an apoptotic effect of an expanded polyglutamine tract? Cancer 1999; 86: 1342–1346.
Moreira Sousa C, McGuire JR, Thion MS, et al: The Huntington disease protein accelerates breast tumour development and metastasis through ErbB2/HER2 signalling. EMBO Mol Med 2013; 5: 309–325.
Kelsey JL, Gammon MD, John EM : Reproductive factors and breast cancer. Epidemiol Rev 1993; 15: 36–47.
Narod SA, Foulkes WD : BRCA1 and BRCA2: 1994 and beyond. Nat Rev Cancer 2004; 4: 665–676.
Bojesen SE, Pooley KA, Johnatty SE, et al: Multiple independent variants at the TERT locus are associated with telomere length and risks of breast and ovarian cancer. Nat Genet 2013; 45: 384e371–372.
Antoniou AC, Wang X, Fredericksen ZS, et al: A locus on 19p13 modifies risk of breast cancer in BRCA1 mutation carriers and is associated with hormone receptor-negative breast cancer in the general population. Nat Genet 2010; 42: 885–892.
Chenevix-Trench G, Milne RL, Antoniou AC, et al: An international initiative to identify genetic modifiers of cancer risk in BRCA1 and BRCA2 mutation carriers: the Consortium of Investigators of Modifiers of BRCA1 and BRCA2 (CIMBA). Breast Cancer Res 2007; 9: 104.
Bieche I, Parfait B, Laurendeau I, et al: Quantification of estrogen receptor alpha and beta expression in sporadic breast cancer. Oncogene 2001; 20: 8109–8115.
Awadelkarim KD, Callens C, Rosse C, et al: Quantification of PKC family genes in sporadic breast cancer by qRT-PCR: evidence that PKCiota/lambda overexpression is an independent prognostic factor. Int J Cancer 2012; 131: 2852–2862.
Barnes DR, Lee A, Investigators E et al: Evaluation of association methods for analysing modifiers of disease risk in carriers of high-risk mutations. Genet Epidemiol 2012; 36: 274–291.
Antoniou AC, Sinilnikova OM, Simard J, et al: RAD51 135G—>C modifies breast cancer risk among BRCA2 mutation carriers: results from a combined analysis of 19 studies. Am J Hum Genet 2007; 81: 1186–1200.
Lee JM, Ramos EM, Lee JH, et al: CAG repeat expansion in Huntington disease determines age at onset in a fully dominant fashion. Neurology 2012; 78: 690–695.
Tassicker RJ, Teltscher B, Trembath MK, et al: Problems assessing uptake of Huntington disease predictive testing and a proposed solution. Eur J Hum Genet 2009; 17: 66–70.
Evans SJ, Douglas I, Rawlins MD, et al: Prevalence of adult Huntington's disease in the UK based on diagnoses recorded in general practice records. J Neurol Neurosurg Psychiatry 2013; 84: 1156–1160.
Morrison PJ, Harding-Lester S, Bradley A : Uptake of Huntington disease predictive testing in a complete population. Clin Genet 2011; 80: 281–286.
Fisher ER, Hayden MR : Multisource ascertainment of Huntington disease in Canada: prevalence and population at risk. Mov Disord 2013; 29: 105–114.
Moreira Sousa C, Humbert S : Huntingtin: here, there, everywhere!. J Huntingtons Dis 2013; 2: 395–403.
Elias S, Thion MS, Yu H, et al: Huntingtin regulates mammary stem cell division and differentiation. Stem Cell Rep 2014; 2: 491–506.
Goehler H, Lalowski M, Stelzl U, et al: A protein interaction network links GIT1, an enhancer of huntingtin aggregation, to Huntington's disease. Mol Cell 2004; 15: 853–865.
Thion MS, McGuire JR, Sousa CM, et al: Unraveling the role of Huntingtin in breast cancer metastasis. J Natl Cancer Inst 2015; 107: djv208.
Semaka A, Kay C, Doty C, et al: CAG size-specific risk estimates for intermediate allele repeat instability in Huntington disease. J Med Genet 2013; 50: 696–703.
Acknowledgements
We thank Frédéric Saudou, Anne Vincent-Salomon, the members of the Humbert and Saudou laboratories for discussions and the GEMO study: National Cancer Genetics Network (UNICANCER Genetic Group, France) and the patients for the biological samples. We also thank all the collaborating GEMO groups for their contribution to this study (a list of collaborating GEMO Centers and their members follows). The coordinating centers, Unité Mixte de Génétique Constitutionnelle des Cancers Fréquents, Hospices Civils de Lyon–Centre Léon Bérard and Equipe (Génétique du cancer du sein), Centre de Recherche en Cancérologie de Lyon: Olga Sinilnikova, Sylvie Mazoyer, Francesca Damiola, Laure Barjhoux, Carole Verny-Pierre, Alain Calender, Sophie Giraud, Mélanie Léone; and Service de Génétique Oncologique, Institut Curie, Paris: Dominique Stoppa-Lyonnet, Marion Gauthier-Villars, Bruno Buecher, Claude Houdayer, Virginie Moncoutier, Muriel Belotti, Carole Tirapo, Antoine de Pauw. Institut Gustave Roussy, Villejuif: Brigitte Bressac-de-Paillerets, Olivier Caron. Centre Jean Perrin, Clermont–Ferrand: Yves-Jean Bignon, Nancy Uhrhammer. Centre Léon Bérard, Lyon: Christine Lasset, Valérie Bonadona, Sandrine Handallou. Centre François Baclesse, Caen: Agnès Hardouin, Pascaline Berthet. Institut Paoli Calmettes, Marseille: Hagay Sobol, Violaine Bourdon, Tetsuro Noguchi, Audrey Remenieras, François Eisinger. CHU Arnaud-de-Villeneuve, Montpellier: Isabelle Coupier, Pascal Pujol. Centre Oscar Lambret, Lille: Jean-Philippe Peyrat, Joëlle Fournier, Françoise Révillion, Philippe Vennin, Claude Adenis. Hôpital René Huguenin/Institut Curie, St Cloud: Etienne Rouleau, Rosette Lidereau, Liliane Demange, Catherine Nogues. Centre Paul Strauss, Strasbourg: Danièle Muller, Jean-Pierre Fricker. Institut Bergonié, Bordeaux: Emmanuelle Barouk-Simonet, Françoise Bonnet, Virginie Bubien, Nicolas Sevenet, Michel Longy. Institut Claudius Regaud, Toulouse: Christine Toulas, Rosine Guimbaud, Laurence Gladieff, Viviane Feillel. CHU Grenoble: Dominique Leroux, Hélène Dreyfus, Christine Rebischung, Magalie Peysselon. CHU Dijon: Fanny Coron, Laurence Faivre. CHU St-Etienne: Fabienne Prieur, Marine Lebrun, Caroline Kientz. Hôtel Dieu Centre Hospitalier, Chambéry: Sandra Fert Ferrer. Centre Antoine Lacassagne, Nice: Marc Frénay. CHU Limoges: Laurence Vénat-Bouvet. CHU Nantes: Capucine Delnatte. CHU Bretonneau, Tours: Isabelle Mortemousque. Groupe Hospitalier Pitié-Salpétrière, Paris: Florence Coulet, Chrystelle Colas, Florent Soubrier. CHU Vandoeuvre-les-Nancy: Johanna Sokolowska, Myriam Bronner. CHU Besançon: Marie-Agnès Collonge-Rame, Alexandre Damette. Creighton University, Omaha, USA: Henry T Lynch, Carrie L Snyder. The study was supported by the Ligue National Contre le Cancer, the Association ‘Le cancer du sein, parlons-en!’ Award, and the Canadian Institutes of Health Research for the 'CIHR Team in Familial Risks of Breast Cancer' program. The EUROSCA Clinical study (EUROSCA/LSHM-CT-2004-503304) was supported in part by the European Union 6th framework program, and included the following principle investigators: J Berciano (Santander), Sylvia Boesch (Innsbruck), Alexis Brice (Paris), Alessandro Filla (Naples), Paola Giunti (London), Thomas Klockgether (Bonn), Ludger Schols and Olaf Riess (Tübingen), Maria Rakowicz (Warsaw), Caterina Mariotti (Milano), Bela Melegh (Budapest), Massimo Pandolfo (Brussels), Bart van de Warrenburg (Nijmegen).
Author information
Authors and Affiliations
Consortia
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Thion, M., Tézenas du Montcel, S., Golmard, JL. et al. CAG repeat size in Huntingtin alleles is associated with cancer prognosis. Eur J Hum Genet 24, 1310–1315 (2016). https://doi.org/10.1038/ejhg.2016.13
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2016.13
This article is cited by
-
Informing about genetic risk in families with Huntington disease: comparison of attitudes across two decades
European Journal of Human Genetics (2021)
-
How the variability between computer-assisted analysis procedures evaluating immune markers can influence patients’ outcome prediction
Histochemistry and Cell Biology (2021)
-
Antagonistic pleiotropy in mice carrying a CAG repeat expansion in the range causing Huntington’s disease
Scientific Reports (2019)