Abstract
Homozygous variants in PGAP1 (post-GPI attachment to proteins 1) have recently been identified in two families with developmental delay, seizures and/or spasticity. PGAP1 is a member of the glycosylphosphatidylinositol anchor biosynthesis and remodeling pathway and defects in this pathway are a subclass of congenital disorders of glycosylation. Here we performed whole-exome sequencing in an individual with cerebral visual impairment (CVI), intellectual disability (ID), and factor XII deficiency and revealed compound heterozygous variants in PGAP1, c.274_276del (p.(Pro92del)) and c.921_925del (p.(Lys308Asnfs*25)). Subsequently, PGAP1-deficient Chinese hamster ovary (CHO)-cell lines were transfected with either mutant or wild-type constructs and their sensitivity to phosphatidylinositol-specific phospholipase C (PI-PLC) treatment was measured. The mutant constructs could not rescue the PGAP1-deficient CHO cell lines resistance to PI-PLC treatment. In addition, lymphoblastoid cell lines (LCLs) of the affected individual showed no sensitivity to PI-PLC treatment, whereas the LCLs of the heterozygous carrier parents were partially resistant. In conclusion, we report novel PGAP1 variants in a boy with CVI and ID and a proven functional loss of PGAP1 and show, to our knowledge, for the first time this genetic association with CVI.
Similar content being viewed by others
Introduction
Cerebral visual impairment (CVI) accounts for 27% of the visual impairment in children, and is due to a disorder in projection and/or interpretation of the visual input in the brain.1, 2, 3 Acquired damages, for example, due to preterm birth, are well known causes of CVI, whereas genetic factors are largely unidentified.4 Several chromosomal aberrations have been associated with CVI, such as Phelan–McDermid syndrome (22q13.3 deletion) and 1p36 microdeletion syndrome.5 Moreover, single-gene disorders can be implicated in CVI, and, recently, we identified de novo variants in NR2F1 as a cause of CVI.6 In addition, in several congenital disorders of glycosylation (CDG type 1a, type 1q and type 1v) CVI has been reported.4, 7, 8, 9 Glycosylation disorders are caused by a defect in the glycosylation of glycoproteins and glycolipids, and one subclass is the defect in glycosylphosphatidylinositol (GPI) anchor glycosylation.10
GPI anchor many cell-surface proteins with various functions, so called GPI-anchored proteins (GPI-APs), to the membrane of eukaryotic cells.11, 12, 13 GPI is synthesized in the endoplasmic reticulum, transferred to the proteins, and remodeled. During the biosynthesis of GPI anchors, an acyl chain is linked to the inositol. This acyl chain is a transient structure and is necessary for efficient completion of later steps in GPI biosynthesis. After the attachment of GPI to the protein, this acyl chain is removed by PGAP1 (post-GPI attachment to proteins 1), the first step of the remodeling phase.14 Subsequently, the GPI anchor is transported from the endoplasmic reticulum to the plasma membrane through the Golgi apparatus and further remodeled. When the inositol-linked acyl chain is not removed, this transport is delayed,14 but the presence of this acyl chain does not affect the cell-surface expression of GPI-APs.15
Several genes known to be involved in the GPI anchor biosynthesis (PIGA, PIGL, PIGN, PIGT, PIGV, PIGO, PIGW and PIGQ) and modeling (PGAP2 and PGAP3) are implicated in X-linked and autosomal recessive intellectual disability disorders.16, 17, 18, 19, 20, 21, 22, 23, 24, 25, 26 Additional features such as seizures, congenital abnormalities and facial dysmorphisms are commonly present. Moreover, CVI has been reported in individuals with PIGA, PIGN and PIGT variants, suggesting that the GPI anchor biosynthesis is important in the development of the visual areas of the brain.18, 19, 27, 28
Recently, two families with developmental delay, seizures and/or spasticity were reported with homozygous variants in PGAP1.15, 29 However, PGAP1 variants have not to date been associated with CVI.
Here we report an individual with CVI and intellectual disability (ID) and variants in PGAP1, thereby showing the association between PGAP1 and CVI.
Materials and methods
Case report
The boy was the second child of healthy unrelated parents. He was born after a normal pregnancy by a planned caesarian section because of a difficult delivery of the previous child. His birth weight was 3550 g (70th centile) at a gestational age of 38 weeks and Apgar scores were 9 and 10, after 1 and 5 min, respectively. During the neonatal period, he was hypotonic and there were feeding difficulties. His development was delayed as he started walking and speaking around the age of 2–2½ years. At the age of 8 years 3 months, his total IQ was 49 (Wechsler Intelligence Scale for Children-III). He was very social in his behavior. There were no signs of epilepsy and an MRI of the brain was normal at age 5 years. Hearing was normal, but ophthalmological examination at 7 years revealed CVI with a visual acuity of 0.3 (Landolt C). He had strabismus divergence and nystagmus. Slit lamp examination and funduscopy were normal. He looked only shortly at a target and he had more difficulties with recognition of objects in clutter (crowding) than was expected for his age.
He had mild asthma and his APTT was prolonged due to a factor XII deficiency (factor XII activity <1%). His mother and sister also had decreased factor XII activity (40% and 22%, respectively), with a normal APTT. There were no signs of immunodeficiency.
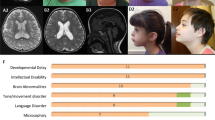
Physical examination at 7 years 10 months showed a height of 131 cm (50th centile) and head circumference of 52.5 cm (50th centile). Facial dysmorphisms consisted of upward slanting palpebral fissures, deep-set eyes, large ear lobes and prominent helices and antihelices (Figure 1). His teeth showed extra mamelons, and diminished enamel was noted. Previous genetic studies, consisting of FMR1 gene, AIP gene, array CGH and metabolic studies (including transferrin glycosylation assay) showed normal results. This study was approved by the Ethics Committee of the Radboud University Medical Center (Commissie Mensgebonden Onderzoek, regio Arnhem-Nijmegen), and written informed consent was obtained.
Phenotype of the affected individual and the functional analysis of the variants. (a) Photographs of the affected individual, note the upward slanting palpebral fissures, deep-set eyes, large ear lobes and prominent helices and antihelices. (b) Schematic representation of PGAP1-containing transmembrane domains (in green). The positions of the variants identified in this study and in the previous reports are indicated. †For this variant the effect on amino acid sequence was not studied. ††Considered as a variant without functional impact. (c) CHO cells deficient for PGAP1 were used to investigate the expected structural abnormalities of the GPI anchors by testing the sensitivity of GPI-APs to PI-PLC. Wild-type PGAP1 and the construct containing the variant c.914C>T (p.(Ala305Val)) rescued the sensitivity for PI-PLC strongly suggesting that this variant is benign. The PGAP1 constructs containing the c.274_276del (p.(Pro92del)) and c.921_925del (p.(Lys308Asnfs*25)) variants did not increase the sensitivity for PI-PLC, suggesting that both are causal.
Whole-exome sequencing (WES)
WES in this individual (CVI10) was performed as part of a larger study to identify the genetic causes of CVI.6 WES was performed by the Baylor-Hopkins Center for Mendelian Genomics, using methods reported previously (Supplementary Material and Methods, whole-exome sequencing).6, 30 The results were analyzed for de novo variants, homozygous variants, compound heterozygous variants and hemizygous variants as described in de Ligt et al.31 Truncating variants, splice site variants and missense variants predicted to be pathogenic were validated by Sanger sequencing in the affected subject and his parents. The variants identified have been submitted to the Leiden Open Variation Database LOVD database (http://databases.lovd.nl/, individual #00025011).
Functional analysis using CHO cells
PGAP1-deficient CHO cells (C10)14 were transiently transfected with wild-type or mutant pMEFLAG-hPGAP1 by electroporation as reported previously (Supplementary Material and Methods, Functional analysis using CHO cells).15 Four days after transfection, cells were treated with or without PI-PLC. Surface expression of GPI-APs was determined by staining cells with mouse anti-human CD59 (5H8), -human DAF (IA10), -hamster uPAR (5D6) antibodies and analyzed by flow cytometry using Flowjo software (Tommy Digital Inc., Tokyo, Japan).
PI-PLC treatment and FACS analysis
Heparin blood samples were collected from the affected individual and his unaffected parents and lymphoblastoid cell lines (LCLs) were generated (Supplementary Material and Methods, PI-PLC treatment and FACS analysis). Four days after transfection, cells were treated with or without PI-PLC as reported previously.15 Surface expression of GPI-APs was determined by staining cells with mouse anti-human CD59 (5H8), -human DAF (IA10), -human CD48 (BJ40) antibodies and analyzed by flow cytometry using Flowjo software.
Results
WES
WES in the affected individual and the parents was performed with an average coverage of 126 ×. After applying the above mentioned prioritizations step only two variants in PGAP1 remained, a paternal inherited chr2.hg19:g.197784746_197784748del; c.274_276del; p.(Pro92del) and a maternal inherited chr2.hg19:g.197761857_197761861del; c.921_925del; p.(Lys308Asnfs*25) (MIM 611655, RefSeq accession number NM_024989.3; Supplementary Material,Supplementary Table S1). The variants were validated by Sanger sequencing, and an additional maternal inherited variant was identified, chr2.hg19:g.197761868C>T; c.914C>T; p.(Ala305Val), which was previously filtered out due to quality settings (Supplementary Material, Supplementary Figure S1). In addition, the exome results were screened for rare variants (<1% occurrence) in the coding regions of F12 (MIM 610619, RefSeq accession number NM_000505.3). A paternal inherited heterozygous variant was identified and validated with Sanger sequencing chr5.hg19:g.176831388G>A; c.827G>A; p.(Trp276*).
Functional analysis using CHO cells
Under the presence of PI-PLC, structurally normal GPI-APs, without inositol-linked acyl chain, are cleaved from the cell membrane.32 CHO cells deficient for PGAP1 were used to investigate the expected structural abnormalities of the GPI anchors by testing the sensitivity of GPI-APs to PI-PLC. Wild-type PGAP1 and the p.(Ala305Val) rescued the sensitivity for PI-PLC (Figure 1 and Supplementary Figure S2). However, transfection with empty vector, the p.(Pro92del) mutant or p.(Lys308Asnfs*25) mutant, did not increase the sensitivity for PI-PLC (Supplementary Figure S3).
PI-PLC treatment and FACS analysis
In addition, the LCLs of the affected individual and parents were analyzed for PI-PLC resistance. The LCLs of the affected individual were completely resistant to PI-PLC treatment, whereas the LCLs of the parents were partially resistant (Supplementary Figure S4).
Discussion
Here we present an individual with CVI, ID, minor facial dysmorphisms and factor XII deficiency. WES revealed compound heterozygous variants in PGAP1, c.274_276del (p.(Pro92del)), c.914C>T (p.(Ala305Val)) and c.921_925del (p.(Lys308Asnfs*25)). To investigate the functional impact of the variants, PI-PLC sensitivity assay was performed in PGAP1-deficient CHO cells. The c.914C>T (p.(Ala305Val)) mutant rescued the PI-PLC sensitivity in PGAP1-deficient CHO cells, similar to wild-type PGAP1. Therefore, this variant was considered to have no functional impact. The c.274_276del (p.(Pro92del)) and c.921_925del (p.(Lys308Asnfs*25)) mutant constructs, however, were not able to rescue the sensitivity, indicating a functional loss of the cDNA constructs. In addition, the PI-PLC sensitivity was investigated in LCLs derived from the affected individual and parents. The LCLs of the affected individual were resistant to PI-PLC, indicating that the GPI-APs are structurally abnormal, whereas the parents showed a partial resistance. These results are in line with the reported family with PGAP1 variants by Murakami et al.15 The other family reported by Novarino et al.29 was not tested. Besides a developmental delay, the phenotype is variable. The reported siblings by Murakami et al. had severe developmental delay with seizures, stereotypic movements and brain atrophy (Table 1). The siblings reported by Novarino et al. had spasticity and structural brain abnormalities. In the individual described in this study, no seizures or brain abnormalities were present, but he had impaired vision and CVI. No ophthalmological examination was performed in the Murakami siblings. In the Novarino siblings, no ‘ophthalmological signs’ were present, but whether complete ophthalmological examination had been performed remained unclear. Depending on the genetic background of the mouse strain used, PGAP1 deficiency in mice could lead to various phenotypes, ranging from a normal structural brain to forebrain abnormalities, including holoprosencephaly.33, 34, 35, 36 In the mutant phenotypically abnormal mice, Wnt signaling and Nodal signaling were reported to be affected.35, 36 However, the precise role of PGAP1 in the function and development of the brain and more specific of the visual system remains to be elucidated.
One marker of CDG syndromes can be the abnormal glycosylation of transferrin.37 However, in the subclass of GPI anchor glycosylation defects, no abnormal transferrin is detectable, making this marker unreliable in diagnosing GPI anchor glycosylation defects, such as those caused by variants in PGAP1.
One other feature of CDG syndromes is the deficiency of coagulation factors.37 However, factor XII deficiency has not been reported so far, neither has a direct functional link between factor XII and GPI-APs been made. Factor XII deficiency can be caused by variants in F12; a heterozygous variant can lead to an intermediated level of factor XII activity, whereas homozygous or compound heterozygous variants lead to almost no activity (<1%).38, 39 The current individual had a factor XII activity <1%, and a paternal inherited loss of function variant in F12, c.827G>A (p.(Trp276*)), but his father had a normal factor XII activity. The mother had an intermediate factor XII activity, but no variant could be identified in her exome results. Whether the variants in PGAP1 are related to the factor XII deficiency is yet unclear.
Two other GPI anchor-modeling proteins, PGAP2 and PGAP3, are implicated in hyperphosphatasia with mental retardation syndrome, also named Mabry syndrome.22, 23, 24 These individuals showed, in addition to ID, seizures, typical facial dysmorphisms and an increased alkaline phosphatase (ALP). This increase is due to diminished GPI-APs on the cell surface, resulting in less binding of ALP to the cell membrane and more ALP in the plasma.22, 23, 24, 40, 41 ALP was not measured in the PGAP1-affected individuals, but in PGAP1-deficient cells, no diminished cell-surface expression of GPI-APs was measured, making elevated ALP levels less likely.15 In addition, the typical facial dysmorphisms of Mabry syndrome, consisting of apparent hypertelorism, long palpebral fissures, short nose with broad nasal bridge and tip and tented upper lip vermillion, were not present in the here presented individual.
In conclusion, we identified novel PGAP1 variants in an intellectual disabled boy with a proven functional loss of PGAP1 and showed for the first time the genetic association with cerebral visual impairment. Additional affected individuals will be required to gain better insights into pathophysiology and acquire knowledge of the clinical spectrum of this novel disorder.
References
Boonstra N, Limburg H, Tijmes N, van Genderen M, Schuil J, van Nispen R : Changes in causes of low vision between 1988 and 2009 in a Dutch population of children. Acta Ophthalmol 2012; 90: 277–286.
Rahi JS, Cable N : Severe visual impairment and blindness in children in the UK. Lancet 2003; 362: 1359–1365.
Dutton GN, Jacobson LK : Cerebral visual impairment in children. Semin Neonatol 2001; 6: 477–485.
Bosch DG, Boonstra FN, Willemsen MA, Cremers FP, de Vries BB : Low vision due to cerebral visual impairment: differentiating between acquired and genetic causes. BMC Ophthalmol 2014; 14: 59.
Bosch DG, Boonstra FN, Reijnders MR, Pfundt R, Cremers FP, de Vries BB : Chromosomal aberrations in cerebral visual impairment. Eur J Paediatr Neurol 2014; 18: 677–684.
Bosch DG, Boonstra FN, Gonzaga-Jauregui C et al: NR2F1 mutations cause optic atrophy with intellectual disability. Am J Hum Genet 2014; 94: 303–309.
Jensen H, Kjaergaard S, Klie F, Moller HU : Ophthalmic manifestations of congenital disorder of glycosylation type 1a. Ophthalmic Genet 2003; 24: 81–88.
Morava E, Wevers RA, Cantagrel V et al: A novel cerebello-ocular syndrome with abnormal glycosylation due to abnormalities in dolichol metabolism. Brain 2010; 133: 3210–3220.
Enns GM, Shashi V, Bainbridge M et al: Mutations in NGLY1 cause an inherited disorder of the endoplasmic reticulum-associated degradation pathway. Genet Med 2014; 16: 751–758.
Jaeken J : Congenital disorders of glycosylation (CDG): it's (nearly) all in it!. J Inherit Metab Dis 2011; 34: 853–858.
Fujita M, Kinoshita T : GPI-anchor remodeling: potential functions of GPI-anchors in intracellular trafficking and membrane dynamics. Biochim Biophys Acta 2012; 1821: 1050–1058.
Tiede A, Bastisch I, Schubert J, Orlean P, Schmidt RE : Biosynthesis of glycosylphosphatidylinositols in mammals and unicellular microbes. Biol Chem 1999; 380: 503–523.
McConville MJ, Menon AK : Recent developments in the cell biology and biochemistry of glycosylphosphatidylinositol lipids (review). Mol Membr Biol 2000; 17: 1–16.
Tanaka S, Maeda Y, Tashima Y, Kinoshita T : Inositol deacylation of glycosylphosphatidylinositol-anchored proteins is mediated by mammalian PGAP1 and yeast Bst1p. J Biol Chem 2004; 279: 14256–14263.
Murakami Y, Tawamie H, Maeda Y et al: Null mutation in PGAP1 impairing Gpi-anchor maturation in patients with intellectual disability and encephalopathy. PLoS Genet 2014; 10: e1004320.
Johnston JJ, Gropman AL, Sapp JC et al: The phenotype of a germline mutation in PIGA: the gene somatically mutated in paroxysmal nocturnal hemoglobinuria. Am J Hum Genet 2012; 90: 295–300.
Ng BG, Hackmann K, Jones MA et al: Mutations in the glycosylphosphatidylinositol gene PIGL cause CHIME syndrome. Am J Hum Genet 2012; 90: 685–688.
Maydan G, Noyman I, Har-Zahav A et al: Multiple congenital anomalies-hypotonia-seizures syndrome is caused by a mutation in PIGN. J Med Genet 2011; 48: 383–389.
Kvarnung M, Nilsson D, Lindstrand A et al: A novel intellectual disability syndrome caused by GPI anchor deficiency due to homozygous mutations in PIGT. J Med Genet 2013; 50: 521–528.
Krawitz PM, Schweiger MR, Rodelsperger C et al: Identity-by-descent filtering of exome sequence data identifies PIGV mutations in hyperphosphatasia mental retardation syndrome. Nat Genet 2010; 42: 827–829.
Krawitz PM, Murakami Y, Hecht J et al: Mutations in PIGO, a member of the GPI-anchor-synthesis pathway, cause hyperphosphatasia with mental retardation. Am J Hum Genet 2012; 91: 146–151.
Krawitz PM, Murakami Y, Riess A et al: PGAP2 mutations, affecting the GPI-anchor-synthesis pathway, cause hyperphosphatasia with mental retardation syndrome. Am J Hum Genet 2013; 92: 584–589.
Hansen L, Tawamie H, Murakami Y et al: Hypomorphic mutations in PGAP2, encoding a GPI-anchor-remodeling protein, cause autosomal-recessive intellectual disability. Am J Hum Genet 2013; 92: 575–583.
Howard MF, Murakami Y, Pagnamenta AT et al: Mutations in PGAP3 impair GPI-anchor maturation, causing a subtype of hyperphosphatasia with mental retardation. Am J Hum Genet 2014; 94: 278–287.
Chiyonobu T, Inoue N, Morimoto M, Kinoshita T, Murakami Y : Glycosylphosphatidylinositol (GPI) anchor deficiency caused by mutations in PIGW is associated with West syndrome and hyperphosphatasia with mental retardation syndrome. J Med Genet 2014; 51: 203–207.
Martin HC, Kim GE, Pagnamenta AT et al: Clinical whole-genome sequencing in severe early-onset epilepsy reveals new genes and improves molecular diagnosis. Hum Mol Genet 2014; 23: 3200–3211.
Swoboda KJ, Margraf RL, Carey JC et al: A novel germline PIGA mutation in Ferro-Cerebro-Cutaneous syndrome: a neurodegenerative X-linked epileptic encephalopathy with systemic iron-overload. Am J Med Genet A 2014; 164A: 17–28.
Nakashima M, Kashii H, Murakami Y et al: Novel compound heterozygous PIGT mutations caused multiple congenital anomalies-hypotonia-seizures syndrome 3. Neurogenetics 2014; 15: 193–200.
Novarino G, Fenstermaker AG, Zaki MS et al: Exome sequencing links corticospinal motor neuron disease to common neurodegenerative disorders. Science 2014; 343: 506–511.
Lupski JR, Gonzaga-Jauregui C, Yang Y et al: Exome sequencing resolves apparent incidental findings and reveals further complexity of SH3TC2 variant alleles causing Charcot-Marie-Tooth neuropathy. Genome Med 2013; 5: 57.
de Ligt J, Willemsen MH, van Bon BW et al: Diagnostic exome sequencing in persons with severe intellectual disability. N Engl J Med 2012; 367: 1921–1929.
Roberts WL, Myher JJ, Kuksis A, Low MG, Rosenberry TL : Lipid analysis of the glycoinositol phospholipid membrane anchor of human erythrocyte acetylcholinesterase. Palmitoylation of inositol results in resistance to phosphatidylinositol-specific phospholipase C. J Biol Chem 1988; 263: 18766–18775.
Zoltewicz JS, Plummer NW, Lin MI, Peterson AS : oto is a homeotic locus with a role in anteroposterior development that is partially redundant with Lim1. Development 1999; 126: 5085–5095.
Ueda Y, Yamaguchi R, Ikawa M et al: PGAP1 knock-out mice show otocephaly and male infertility. J Biol Chem 2007; 282: 30373–30380.
Zoltewicz JS, Ashique AM, Choe Y et al: Wnt signaling is regulated by endoplasmic reticulum retention. PloS One 2009; 4: e6191.
McKean DM, Niswander L : Defects in GPI biosynthesis perturb Cripto signaling during forebrain development in two new mouse models of holoprosencephaly. Biol Open 2012; 1: 874–883.
Sparks SE, Krasnewich DM et al: Congenital disorders of n-linked glycosylation pathway overview; in Pagon RA, Adam MP, Ardinger HH (eds): GeneReviews. Seattle, WA, USA: University of Washington, 1993.
Schloesser M, Hofferbert S, Bartz U, Lutze G, Lammle B, Engel W : The novel acceptor splice site mutation 11396(G—>A) in the factor XII gene causes a truncated transcript in cross-reacting material negative patients. Hum Mol Genet 1995; 4: 1235–1237.
Schloesser M, Zeerleder S, Lutze G et al: Mutations in the human factor XII gene. Blood 1997; 90: 3967–3977.
Tashima Y, Taguchi R, Murata C, Ashida H, Kinoshita T, Maeda Y : PGAP2 is essential for correct processing and stable expression of GPI-anchored proteins. Mol Biol Cell 2006; 17: 1410–1420.
Murakami Y, Kanzawa N, Saito K et al: Mechanism for release of alkaline phosphatase caused by glycosylphosphatidylinositol deficiency in patients with hyperphosphatasia mental retardation syndrome. J Biol Chem 2012; 287: 6318–6325.
Acknowledgements
We are grateful to the individual involved and his family for their support and cooperation. We thank the technical ophthalmological assistant P. Rison for his assistance during the ophthalmological examination. This work has been supported by grants from Stichting ODAS (to FNB and FPMC), Vereniging Bartiméus-Sonneheerdt (5781251 to FNB and FPMC), and the Dutch Organization for Health Research and Development (917-86-319 and 912-12-109 to BBAdV). In addition, this study was accomplished in part through the Center for Mendelian Genomics research effort funded by the National Institutes of Health and supported by the National Human Genome Research Institute grant U54HG006542 to the Baylor-Hopkins Center for Mendelian Genomics.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Bosch, D., Boonstra, F., Kinoshita, T. et al. Cerebral visual impairment and intellectual disability caused by PGAP1 variants. Eur J Hum Genet 23, 1689–1693 (2015). https://doi.org/10.1038/ejhg.2015.42
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2015.42
This article is cited by
-
Compound heterozygous variants in PGAP1 causing severe psychomotor retardation, brain atrophy, recurrent apneas and delayed myelination: a case report and literature review
BMC Neurology (2016)
-
Molecular subtyping and improved treatment of neurodevelopmental disease
Genome Medicine (2016)
-
Novel genetic causes for cerebral visual impairment
European Journal of Human Genetics (2016)