Abstract
Trisomy 21 (T21), or Down syndrome (DS), is the most frequent and recognizable cause of intellectual disabilities. The level of disability, as evaluated by the intelligence quotient (IQ) test, varies considerably between patients independent of other factors. To determine the genetic or molecular basis of this difference, a high throughput transcriptomic analysis was performed on twenty T21 patients with high and low IQ, and 10 healthy controls using Digital Gene Expression. More than 90 millions of tags were sequenced in the three libraries. A total of 80 genes of potential interest were selected for the qPCR experiment validation, and three housekeeping genes were used for normalizing purposes. HLA DQA1 and HLA DRB1 were significantly downregulated among the patients with a low IQ, the values found in the healthy controls being intermediate between those noted in the IQ+ and IQ− T21 patients. Interestingly, the intergenic region between these genes contains a binding sequence for the CCCTC-binding factor, or CTCF, and cohesin (a multisubunit complex), both of which are essential for expression of HLA DQA1 and HLA DRB1 and numerous other genes. Our results might lead to the discovery of genes, or genetic markers, that are directly involved in several phenotypes of DS and, eventually, to the identification of potential targets for therapeutic interventions.
Similar content being viewed by others
Introduction
Trisomy 21 (T21) or Down syndrome (DS) is a chromosomal disorder resulting from the triplication of all or part of a chromosome 21. It is a common birth defect, the most frequent and recognizable form of intellectual disabilities (ID), appearing in about one out of every 700 newborns. The average intelligence quotient (IQ) of children with DS is around 50, ranging between 30 and 70. Remarkably, a small number of patients have a profound degree of ID, whereas others have a mild degree despite the absence of any genetic, cultural or familial favoring or disfavoring causes.1
Recent progress in studies of patients with partial T21 and mouse models of T21 suggest that it will be soon possible to link characteristic phenotype changes with differential gene expression of specific genes and help to decipher the molecular basis for these abnormalities, which may lead to treatment of the most distressing aspects of this disorder.2 Such optimism is based on recent success with high-throughput genomic approaches in human medicine, gene expression signatures, and gene profiling studies that have linked specific gene regulation to specific phenotypic abnormalities, and aided the diagnosis, the treatment, or the prevention of diseases in DS.3, 4
The purpose of this paper is to identify a case-series of patients with lower (IQ−) and higher IQ (IQ+) and to describe gene expression or transcriptome differences between them and healthy controls using the Digital Gene Expression (DGE) technique. This pilot description may suggest a genetic indicator of better intellectual prognosis in DS patients.
Materials and methods
Subjects
After the analysis of nearly 1000 clinical files of DS patients at the Jérôme Lejeune Institute, a series of 20 patients with a free and homogeneous T21, aged between 18 and 40 years were enrolled in this study. Patients were classified into two groups: those with a relatively lower IQ (IQ <20 or IQ−; four males and one female), and those with a higher one (IQ >70 or IQ+; six males and nine females). The IQ was measured using the Columbia test, a tool validated and largely used in similar settings in France.
Patients were not taking medications, had no neurological problems (epilepsy, seizures, west syndromes, and so on), no changes suggestive of early dementia, no autism, no endocrinological problems (hypo or hyperthyroidism, diabetes, and so on), no sleep-disordered breathing problems, no hearing impairment or vision impairment, no heart problems, no immune deficiency, and no cancer. In addition, in none of the families of the patients serious events were noted (death, frequent hospitalization, child abuse, and so on).
A third group of 10 individuals without T21 (four males and six females), aged between 26 and 39 years were added as controls.
This study was granted approval from the Institute Jérôme Lejeune Committee on Clinical Investigation and conformed to the tenets of the Declaration of Helsinki.
RNA samples
Blood samples were collected for genetic studies after informed written consent was obtained from all parents or guardians on behalf of the participants because of their inability to provide consent, and from the healthy controls. RNA samples were extracted from lymphoblastoid cell lines. RNA samples were obtained from 4 × 106 cells pellet with RNeasy Plus Mini kit (Qiagen Courtaboeuf, France) and QIAshredder (Qiagen). Control of RNA integrity was performed with the 2100 Bioanalyzer (Agilent Technologies, Lesulis, France) using Eukaryotic Total RNA 6000 Nano Chip (Agilent Technologies). RNA quantity was controlled using NanoDrop ND-1000 spectrophotometer (LABTECH, Palaiseau, France).
DGE library construction and tag-to-gene mapping
Three DGE libraries were constructed from pooled RNA samples of patients IQ+ and IQ−, and healthy controls. The libraries were constructed with Illumina's DGE Tag Profiling kit (ILLUMINA; SanDiego, CA, USA) according to the manufacturer's protocol (version 2.1B), using 5 μg of total RNA (mixing equal amounts of RNA from each individual). Sequencing analysis and base calling were performed using the Illumina Pipeline, and sequence tags were obtained after purity filtering. Data from each DGE library were analyzed with BIOTAG software (Skuldtech, Montpellier, France) for tag detection, tag counting and for assessing DGE library quality.5 Raw and treated data are available on http://www.skuldtech.com/trisomie21.
Tag annotation and selection
A local database compiling Homo sapiens sequences and related information from well-annotated sequences of UniGene clusters (NCBI) was generated. For each sequence of this database, the expected DGE tag (canonical tag) located upstream of the 3′-nearest NlaIII restriction site (CATG) of the sequence (R1), as well as putative tags located in inner positions (labeled as R2, R3 and R4 starting from the 3′ end of the transcript) were extracted.5 Experimental tags obtained from DGE libraries were matched and annotated (exact matches for the 17 bp) using this collection of virtual tags. First, a correspondence for each experimental tag with the virtual canonical tags (R1) was looked for. Then, unmatched experimental tags with the R2 tags, then with R3, and R4 were annotated. Targeted tags were selected using R package DESeq (Anders S, Bioconductor) for processing data without replicates. The analyzed genes were selected according to mathematic filters with the highest differential Fold Change (>1.5), FDR adjusted P-value criterion (<10%, Benjamini-Hochberg Method) based on the type I (α≤5%) error.
cDNA synthesis and real-time PCR
Reverse transcriptions were performed for each of the 30 RNA samples in 20 μl final reaction volume with 1.5 μg of total RNA using 200 units of SuperScript II enzyme (M-MLV RT Type, Invitrogen, St Aubin, France) and 250 ng of random primers according to manufacturer's instructions (25 °C 10 min, 42 °C 50 min, 70 °C 15 min), the same day with the same pipettor set and the same manipulator. qPCR experiments were carried out using SYBR Green chemistry on LightCycler480 qPCR apparatus system I (Roche, Meylan, France). The reaction mix was prepared in a final volume of 10 μl as follows: 1 μl of cDNA matrix (1/30 diluted in H2O) was added to 5 μl of SYBR Green I Master Mix (Roche Applied Science, Meylan, France) and 4 μl of Forward and Reverse primers mix (final concentration in PCR reaction of 0.5 μ M each).
To discriminate specific from non-specific products and primer dimers, a melting curve was obtained by gradual increase in temperature from 65 to 95 °C. PCR efficiency was measured on standard curves performed for each primer pairs using a pool of all cDNA diluted as following: 1/10, 1/100, 1/1000, 1/10000 dilution factor. Primer pairs with PCR efficiency <1.8 were excluded of the analysis. The qPCR data were analyzed using the 2−(ΔΔCT) method.6 Data were normalized using three housekeeping genes: LIMK1 (NM_002314), APEH (BC000362) and TUFM (NM_003321) (Table 1). The Partial Least Squares Discriminant Analysis regression (PLSDA) analysis was performed with the R package mixOmics.7 The PLSDA was used to select the most discriminant genes. According to the mixOmics vignette and from the variance importance plot, a score >1 represent a strong weight in the patient discrimination.7
Results
By using Next-Generation Sequencing, we performed a transcriptomic study using the open method, DGE, to reveal genes differentially expressed in the two different selected phenotypes of DS. More than 90 million tags were sequenced in the two libraries. By taking into consideration the DGE-tags with a minimal occurrence of two times, the two libraries revealed 68 046 unique tags. Raw and treated data were integrated in a database with associated tools for annotation, in silico PCR, tag prediction and data visualization. This database is accessible via a user-friendly website on http://www.skuldtech.com/trisomie21 and can incorporate future data from the Next-Generation Sequencing.
After data were filtered and classified according to the statistical approach by DESeq and the fold induction among the well-annotated tags, 80 genes were selected (Table 2). To explore the individual variability, individual qPCR analysis was performed.
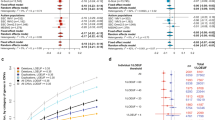
The qPCR data analysis showed that two genes were sufficient enough based on our selection criteria (fold change expression >2.5 and SD<0.1) to discriminate our entire population between IQ− and IQ+. The PLSDA run on these data with the R package mixOmics7 showed that the genes HLA-DQA1 (NM_002122, major histocompatibility complex (MHC), class II, DQ-α 1) and HLA-DRB1 (NM_002124, MHC, class II, DR-β 1) obtained the highest values (HLA-DQA1 got 2.84 and 2.22 for component 1 and 2, respectively, and HLA-DRB1 got 2.56 and 1.90 for component 1 and 2, respectively, of the PLSDA) from the variance importance plot. Both expressions appeared to be less frequent in the IQ− group (Figure 1). Interestingly, the values found in the healthy controls were between both those noted in the IQ− and IQ+ T21 patients. A Monte–Carlo test on a factorial discriminate analysis gave a significant P-value of 0.02.
Discussion
T21 is a direct consequence of either an additional copy of protein-coding genes that are dosage sensitive or an additional copy of non-protein-coding sequences that are regulatory or otherwise functional. The effect of some dosage-sensitive genes on the phenotypes might be allele-specific, and could also depend on the polymorphism of coding and non-coding (regulatory, mRNAs, and so on) nucleotide sequences and on the combination and interaction of all these variants coding for proteins alleles with qualitative (alleles with amino-acid variation) or quantitative (alleles with variation in gene expression level) traits. Dosage sensitive genes could act directly to induce pathogenesis or indirectly by interacting with genes or gene products of either aneuploid or non-aneuploid genes or gene products.8 It is a reasonable speculation that the genetic background of individuals has an important role in the variability of phenotypic severity that is seen in DS.2
In order to understand why some T21 patients present a wide difference in their intelligence despite the absence of any known favoring or disfavoring factors, we initiate the construction of three transcriptomic libraries, in T21 IQ− and IQ+ patients and in healthy controls using the SAGE technique. Indeed, searching for transcriptional alterations is an easier (or more effective) means to find relevant loci than would be complete sequencing. Such approach was already tested in DS and different pathologies.9, 10, 11, 12
The low number of patients, especially in the IQ− group, was secondary to the fact that we wanted to restrict this study only to the T21 patients that we could discriminate by IQ test. Interestingly, the group IQ− was predominantly males, whereas the IQ+ group was frequently females with a sex ratio of 2/3. Few reports already pointed the fact that boys with DS are more affected than girls but without any reason.13
Two genes with major expression differences were found: HLA-DQA1 and HLA-DRB1. Both genes were less expressed in the T21 IQ− population (Figure 1). HLA loci were already reported as leading to significant risk factor for celiac disease in DS patients,14 but never for ID. Nevertheless, HLA has been associated with cognitive ability. For example, Cohen et al.15 showed that a proportion of patients with primary neuronal degeneration of the Alzheimer type have the HLA-B7 antigen, and that patients with HLA-B7 antigens had selective attention span significantly lower than Alzheimer patients without the antigen. Recently, HLA-DRB1 has been associated with cognitive ability in both demented and non-demented individuals,16 and a positive association of HLA-DR4 with attention deficit and hyperactivity disorder and the role of the HLA-DRB1 in the etiology of some types of childhood neuropsychiatric illnesses were reported.17 Future qualitative and quantitative studies at the level of the HLA-DQA1 and HLA-DRB1 proteins on the PBLs of more IQ+ and IQ− DS patients might further validate our results found at the transcriptional level. Likewise, the comparison between the SNPs located within the DS critical region on chromosome 21 in the two groups of DS patients might be informative.
Interestingly, the intergenic region between HLA-DQA1 and HLA-DRB1, is characterized by the presence of a CTCF-binding CCCTC sequence, XL9, of high histone acetylation and of particular importance for the control of the expression of both HLA genes and for the chromatin architecture of the MHC class II locus.18, 19, 20, 21 The fact that CTCF expression is increased and transcription of HLA DQA1 and HLA DRB1 is decreased in the IQ− group could be in favor to a repressor role for CTCF. On the other hand, the RFX1 gene, coding for a protein that binds to the X-boxes of MHC class II genes and which is essential in their expression, shows a three-fold decreased level of expression in the IQ− group in comparison to the IQ+ group. In contrast, in the IQ+ group, the downregulated CTCF gene expression and the increased RFX1 gene expression could explain the upregulation of the HLA-DQA1 and HLA-DRB1 genes. Furthermore, it was found recently that cohesin and CTCF-binding sites have a high degree of overlap, and interacts with each other.20, 22, 23, 24 Moreover, often cohesin and CTCF have been found to colocalize at several thousand sites in non-repetitive sequences in the human genome.22, 24, 25 In addition to the regulation of gene expression, cohesin has also other important functions: it forms a huge tripartite ring, mediates the sister chromatid cohesion, and facilitates the repair of damaged DNA.26, 27 From our analysis, we can postulate that HLA-DQA1 and HLA-DRB1 may represent a genetic biomarker for predicting differences in ID conditions, but also that polymorphisms or mutations of the cohesin subunits might have an important role for the non-disjunction of the chromosomes 21 and/or for the dysregulation of the expression of many genes.
Other genes present a different transcriptional pattern between the IQ− and IQ+ groups. Although this difference is not as obvious as with HLA, it might be interesting to be carefully looked at (Table 2). For instance, APP (Amyloid-β (A4) precursor protein), located on chromosome 21q21.3, was overexpressed in the IQ− group. Duplication of the APP gene was found to lead to early-onset Alzheimer disease (AD) and prominent cerebral amyloid angiopathy.28 Triplication of the APP gene accelerates the APP expression, leading to cerebral accumulation of APP-derived amyloid-β peptides, early-onset AD neuropathology, and age-dependent cognitive sequelae.29 At relatively early ages, DS patients develop progressive formation and extracellular aggregation of amyloid-β peptide, considered as one of the causal factors in the pathogenesis of AD.30 The cystathionine β-synthase (CBS) gene, located on human chromosome 21q22.3 encodes a key enzyme of sulfur-containing amino acid metabolism, a pathway involved in several brain physiological processes. It is overexpressed in the brain of individuals with DS,31 and thus was considered as a good candidate in having a role in the DS cognitive profile.32 Recently, Régnier et al.33 studied the neural consequences of CBS overexpression in a transgenic mouse line expressing the human CBS gene. They observed that the transgenic mice showed normal behavior, and that hippocampal synaptic plasticity was facilitated. Thus, they raised the possibility that CBS overexpression might have an advantageous effect on some cognitive functions in DS. Our results do not confirm the latter hypothesis as we found that CBS was overexpressed in the IQ− group v/s IQ+ group. The glycinamide phosphoribosyltransferase (GART) gene, located on 21q22.11 was found overexpressed in the IQ+ group. GART is an essential enzyme in de novo purine biosynthesis (OMIM 138440). In 1993, Peeters et al.34 studied the variations in mitotic index of lymphocyte cultures to which various metabolites of purine synthesis (inosine, adenosine and guanosine) were added. They unexpectedly found a significant decrease in mitotic index in DS patients without psychiatric complications when compared with normal controls, and opposite reactions in DS patients presenting psychotic features. They concluded that T21 patients may have different purine metabolism depending on whether or not they have associated psychiatric complications. Purine-rich diet or the prescription of exogenous inosine, and serotonine-rich diet were used to treat psychotic T21 patients and some results in reduction of self-injurious behavior were reported.34, 35 Our results tend to give a beginning of explanation to the latter observations. The overexpression of GART in IQ+ patients might prevent the apparition of any abnormal behavior in T21 patients leading to a better IQ. Supplementing such patients with a purine-rich diet might not be beneficial (as seen by the decrease mitotic index) on the contrary of patients with a lower expression of GART.
Interestingly, DYRK1A (Dual-specificity tyrosine-(Y)-phosphorylation-regulated kinase 1A), located on chromosome 21q22.2 and known to have a significant role in developmental brain defects and in early onset neurodegeneration, neuronal loss and dementia in DS36 did not show a significantly different expression between the IQ− and IQ+ groups.
In conclusion, we established a transcriptome of DS patients with IQ− and IQ+ and found that HLA-DQA1 and HLA-DRB1 may discriminate between the two populations. However, the pools of eligible patients available for such studies in particular centers are usually small, and therefore findings remain limited in their power to significantly rule in or out a marker that discriminates early between DS who will be IQ− and who will be IQ+. Larger multicenter series will allow to determine in a valid way the presence of such markers and whether they are sex-specific. Beyond providing evidence to support the hypothesis that has directed much of the work discussed, the importance of determining valid markers would have major consequences for informing pathogenesis and for providing defined targets to combat pathogenesis. This may lead to the discovery of genes that are ‘directly’ involved in several phenotypes of DS and eventually to the identification of potential targets for therapeutic interventions. For example, the genetic association with HLA supports the involvement of the immune system in ID and offers new targets for drug development.
Continued and increasing investments in research on the genetic and molecular basis of T21 promise to transform the lives of these individuals and the communities in which they live.
References
Carr J : Six weeks to 21 Years: A longitudinal study of children with Down’s syndrome and their families. J Child Psychol Psychiatry 1991; 29: 407–431.
Mégarbané A, Ravel A, Mircher C et al: The 50th anniversary of the discovery of trisomy 21: the past, present, and future of research and treatment of Down syndrome. Genet Med 2009; 11: 611–616.
Lima FA, Moreira-Filho CA, Ramos PL et al: Decreased AIRE expression and global thymic hypofunction in Down syndrome. J Immunol 2011; 187: 3422–3430.
Loudin MG, Wang J, Eastwood Leung HC et al: Genomic profiling in Down syndrome acute lymphoblastic leukemia identifies histone gene deletions associated with altered methylation profiles. Leukemia 2011; 25: 1555–1563.
Piquemal D, Commes T, Manchon L et al: Transcriptome analysis of monocytic leukemia cell differentiation. Genomics 2002; 80: 361–371.
Livak KJ, Schmittgen TD : Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods 2001; 25: 402–408.
Lê Cao KA, Boitard S, Besse P, Sparse PLS : discriminant analysis: biologically relevant feature selection and graphical displays for multiclass problems. BMC Bioinformatics 2011; 12: 253.
Antonarakis SE, Lyle R, Dermitzakis ET, Reymond A, Deutsch S : Chromosome 21 and down syndrome: from genomics to pathophysiology. Nat Rev Genet 2004; 5: 725–738.
Booij BB, Lindahl T, Wetterberg P et al: A gene expression pattern in blood for the early detection of Alzheimer's disease. J Alzheimers Dis 2011; 23: 109–119.
Chrast R, Scott HS, Papasavvas MP et al: The mouse brain transcriptome by SAGE: differences in gene expression between P30 brains of the partial trisomy 16 mouse model of Down syndrome (Ts65Dn) and normals. Genome Res 2000; 10: 2006–2021.
Costa V, Angelini C, D'Apice L et al: Massive-scale RNA-Seq analysis of non ribosomal transcriptome in human trisomy 21. PLoS ONE 2011; 6: e18493.
Sommer CA, Pavarino-Bertelli EC, Goloni-Bertollo EM, Henrique-Silva F : Identification of dysregulated genes in lymphocytes from children with Down syndrome. Genome 2008; 51: 19–29.
Kittler P, Krinsky-McHale SJ, Devenny DA : Sex differences in performance over 7 years on the Wechsler Intelligence Scale for Children-Revised among adults with intellectual disability. J Intellect Disabil Res 2004; 48: 114–122.
Book L, Hart A, Black J, Feolo M, Zone JJ, Neuhausen SL : Prevalence and clinical characteristics of celiac disease in Down’s syndrome in a US study. Am J Med Genet 2001; 98: 70–74.
Cohen D, Eisdorfer C, Walford RL : Histocompatibility antigens (HLA) and patterns of cognitive loss in dementia of the Alzheimer type. Neurobiol Aging 1981; 2: 277–280.
Payton A, van den Boogerd E, Davidson Y et al: Influence and interactions of cathepsin D, HLA-DRB1 and APOE on cognitive abilities in an older non-demented population. Genes Brain Behav 2006; 5: 23–31.
Aureli A, Sebastiani P, Del Beato T et al: Investigation on the possible relationship existing between the HLA-DR gene and attention deficit hyperactivity disorder and/or mental retardation. Int J Immunopathol Pharmacol 2008; 21: 985–991.
Majumder P, Gomez JA, Boss JM : The human major histocompatibility complex class II HLA-DRB1 and HLA-DQA1 genes are separated by a CTCF-binding enhancer-blocking element. J Biol Chem 2006; 281: 18435–18443.
Majumder P, Gomez JA, Chadwick BP, Boss JM : The insulator factor CTCF controls MHC class II gene expression and is required for the formation of long-distance chromatin interactions. J Exp Med 2008; 205: 785–798.
Majumder P, Boss JM : CTCF controls expression and chromatin architecture of the human major histocompatibility complex class II locus. Mol Cell Biol 2010; 30: 4211–4223.
Majumder P, Boss JM : DNA methylation dysregulates and silences the HLA-DQ locus by altering chromatin architecture. Genes Immun 2011a; 12: 291–299.
Parelho V, Hadjur S, Spivakov M et al: Cohesins functionally associate with CTCF on mammalian chromosome arms. Cell 2008; 132: 422–433.
Majumder P, Boss JM : Cohesin regulates MHC class II genes through interactions with MHC class II insulators. J Immunol 2011; 187: 4236–4244.
Wendt KS, Yoshida K, Itoh T et al: Cohesin mediates transcriptional insulation by CCCTC-binding factor. Nature 2008; 451: 796–801.
Wendt KS, Peters JM : How cohesin and CTCF cooperate in regulating gene expression. Chromosome Res 2009; 17: 201–214.
Kudo NR, Wassmann K, Anger M et al: Resolution of chiasmata in oocytes requires separase-mediated proteolysis. Cell 2006; 126: 135–146.
Kudo NR, Anger M, Peters AH et al: Role of cleavage by separase of the Rec8 kleisin subunit of cohesin during mammalian meiosis I. J Cell Sci 2009; 122: 2686–2698.
Rovelet-Lecrux A, Hannequin D, Raux G et al: APP locus duplication causes autosomal dominant early-onset Alzheimer disease with cerebral amyloid angiopathy. Nat Genet 2006; 38: 24–26.
Moncaster JA, Pineda R, Moir RD et al: Alzheimer's disease amyloid-beta links lens and brain pathology in Down syndrome. PLoS ONE 2010; 5: e10659.
Azkona G, Amador-Arjona A, Obradors-Tarragó C et al: Characterization of a mouse model overexpressing beta-site APP-cleaving enzyme 2 reveals a new role for BACE2. Genes Brain Behav 2010; 9: 160–172.
Lockstone HE, Harris LW, Swatton JE et al: Gene expression profiling in the adult Down syndrome brain. Genomics 2007; 90: 647–660.
Lejeune J : Pathogenesis of mental deficiency in trisomy 21. Am J Med Genet 2000; 7: 20–30.
Régnier V, Billard JM, Gupta S et al: Brain phenotype of transgenic mice overexpressing cystathionine β-synthase. PLoS ONE 2012; 7: e29056.
Peeters MA, Megarbane A, Cattaneo F, Rethore MO, Lejeune J : Differences in purine metabolism in patients with Down’s syndrome. J Intellect Disabil Res 1993; 37: 491–505.
Gedye A : Dietary increase in serotonine reduces self-injurious behavior in a Down’s syndrome adult. J Ment Defic Res 1990; 34: 247–258.
Wegiel J, Gong CX, Hwang YW : The role of DYRK1A in neurodegenerative diseases. FEBS J 2011; 278: 236–245.
Acknowledgements
This work was supported by the Jérôme Lejeune Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Mégarbané, A., Noguier, F., Stora, S. et al. The intellectual disability of trisomy 21: differences in gene expression in a case series of patients with lower and higher IQ. Eur J Hum Genet 21, 1253–1259 (2013). https://doi.org/10.1038/ejhg.2013.24
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2013.24
Keywords
This article is cited by
-
Methylomic profiling in trisomy 21 identifies cognition- and Alzheimer’s disease-related dysregulation
Clinical Epigenetics (2019)
-
A quantitative transcriptome reference map of the normal human brain
neurogenetics (2014)
-
Performance of Down syndrome subjects during a coincident timing task
International Archives of Medicine (2013)