Abstract
Genetic and nongenetic factors contribute to development of pseudoexfoliation (PEX) syndrome, a complex, age-related, generalized matrix process frequently associated with glaucoma. To identify specific genetic variants underlying its etiology, we performed a genome-wide association study (GWAS) using a DNA-pooling approach. Therefore, equimolar amounts of DNA samples of 80 subjects with PEX syndrome, 80 with PEX glaucoma (PEXG) and 80 controls were combined into separate pools and hybridized to 500K SNP arrays (Affymetrix). Array probe intensity data were analyzed and visualized with expressly developed software tools GPFrontend and GPGraphics in combination with GenePool software. For replication, independent German cohorts of 610 unrelated patients with PEX/PEXG and 364 controls as well as Italian cohorts of 249 patients and 190 controls were used. Of 19, 17 SNPs showing significant allele frequency difference in DNA pools were confirmed by individual genotyping. Further single genotyping at CNTNAP2 locus revealed association between PEX/PEXG for two SNPs, which was confirmed in an independent German but not the Italian cohort. Both SNPs remained significant in the combined German cohorts even after Bonferroni correction (rs2107856: Pc=0.0108, rs2141388: Pc=0.0072). CNTNAP2 was found to be ubiquitously expressed in all human ocular tissues, particularly in retina, and localized to cell membranes of epithelial, endothelial, smooth muscle, glial and neuronal cells. Confirming efficiency of GWAS with DNA-pooling approach by detection of the known LOXL1 locus, our study data show evidence for association of CNTNAP2 with PEX syndrome and PEXG in German patients.
Similar content being viewed by others
Introduction
Pseudoexfoliation (PEX) syndrome, a late onset, generalized disorder of the extracellular matrix, is characterized by production and accumulation of an abnormal extracellular material in many intra- and extraocular tissues.1 Clinically, it manifests with marked intraocular changes as it involves all tissues of the anterior segment of the eye. Whitish fibrillogranular material is visible by slit-lamp microscopy on the anterior lens capsule and at the pupillary border.2 Furthermore PEX is one of the most common causes of open-angle glaucoma accounting for about 25% of all open-angle glaucoma worldwide.2 The etiology of PEX syndrome is still poorly understood but increased prevalence of the disease in certain populations and observed familial aggregation are compatible with PEX being a complex genetic disorder.3, 4
Proteomic approaches identified the presence of elastic fiber components (such as fibrillin-1, vitronectin and fibulin-2), clusterin as well as apolipoprotein E (APOE) within PEX material deposits, suggesting them as possible functional candidate genes for PEX disease.5, 6 To date however, among these candidates only polymorphisms in the gene encoding for clusterin (CLU) have been shown to be associated both with PEX syndrome and PEX glaucoma (PEXG) in different populations.7, 8 Association with alleles of the gene encoding APOE however is still controversial9, 10 but could be excluded in our population of PEX patients.11 A unique genome-wide scan carried out with 75 PEX patients and 14 474 population-based control subjects allowed the detection of a strong association between two nonsynonymous single nucleotide polymorphisms (SNPs) in the lysyl oxidase-like 1 (LOXL1) gene and both PEX syndrome and PEXG in populations from Iceland and Sweden.12 This association has been subsequently confirmed in many cohorts worldwide,13, 14, 15, 16, 17, 18, 19, 20, 21, 22 including our present group of PEX patients.23
Aiming to reduce experimental costs, DNA-pooling analysis has gained interest as a potential alternative to individual genotyping.24, 25 So far risk factors for a number of complex traits such as alcohol addiction,26 mental impairment,27, 28 multiple sclerosis29 and abnormal cholesterol levels30 have been identified. Furthermore, these studies have been greatly facilitated by development of specialized analysis software such as GenePool31 and additions such as GPFrontend and GPGraphics (Uebe et al., submitted) focused on interpretation and graphical evaluation of the pooled hybridization intensities at the single-array SNP.
To discover new genetic variants associated with PEX disease, we applied these developments to a genome-wide association study (GWAS) using pooled DNA samples of patients with PEX and control subjects.
Materials and methods
Study populations
The study was approved by the ethical review boards of the medical faculty of the Universities of Erlangen-Nuremberg, Tuebingen, Wuerzburg (Germany) and that of the hospital in Monfalcone-Gorizia (Italy) and was in accordance with the Declaration of Helsinki Principles. All subjects gave informed consent before entering the study. Our discovery group consisted of 160 patients of German origin (80 PEX syndrome and 80 PEXG patients randomly selected for DNA-pooling experiments) and 80 healthy German subjects. Our German replication cohort was composed of 610 PEX patients and 364 control individuals; the Italian replication cohort of 249 PEX patients and 190 healthy subjects. Recruitment and clinical evaluation were performed as previously described.8, 11, 23
DNA extraction, quantification and pool construction
Genomic DNAs were extracted from peripheral blood leukocytes of patients and control individuals with automated techniques (AutoGenFlex 3000; Autogen, Holliston, MA, USA) using FlexiGene chemistry (Qiagen, Hilden, Germany). DNA samples were quantified using a spectrophotometer (260 nm, Ultrospec III; Pharmacia Biotech, Freiburg, Germany) and diluted to a concentration of 50 ng/μl. Afterwards each DNA was quantified using fluorimetry (Quant-iT PicoGreen dsDNA Assay; Invitrogen, Karlsruhe, Germany) and checked for quality using 1% agarose gel electrophoresis. Degraded DNA samples were excluded from pooling analysis. Three DNA pools were created from randomly selected samples from our cohorts: 100 ng DNA aliquots of 80 German PEX patients, 80 PEXG patients and 80 healthy individuals were each combined into three DNA pools.
SNP microarray allelotyping
Each of the three DNA pools was allelotyped using the GeneChip Mapping 500K Array Set (Affymetrix, Santa Clara, CA, USA) in accordance with the standard protocol for individual DNA samples.
GPFrontend and GPGraphics
Using the Microsoft .NET 2.0 platform and Visual Studio 2005, we have created a graphical software package to analyze data from DNA pooling based microarray GWA studies. This package includes parts of GenePool 0.8.2, existing software suitable for GNU/Linux systems, which we have modified to run in a Microsoft Windows environment and to generate some additional data, enabling it to interact with the .NET parts created by us. The created programs are GPFrontend, a GUI frontend to use the GenePool programs and to generate metadata files, and GPGraphics, a program to further analyze and graphically evaluate GenePool's output (Uebe et al., submitted). The software package reads one of the output files of Gene Pool and creates a bitmap file for every chromosome, in which the score or rank of every SNP is encoded to the height of a peak. Differences of relative allele signals (RAS) between cases and controls were calculated with GenePool using t-test statistics. In GPGraphics, a logarithmic filter was used to visually identify chromosomal regions of high-ranking SNPs, that is, those with highest RAS differences between groups.
Individual genotyping
To validate high-ranking single-nucleotide polymorphisms (SNPs) from the arrays, we selected up to 2 SNPs from eleven clusters containing at least 2 SNPs positioned among the first 1000 SNPs and in high linkage disequilibrium (LD) each other for individual genotyping. Altogether 19 SNPs capturing >95% of overall genetic variance observed in Europeans were genotyped in the 80 PEX syndrome, 80 PEXG and 80 healthy individuals used for DNA-pooling performing predeveloped TaqMan assays (Applied Biosystems, Foster City, CA, USA). A genetic association study was performed.
Next, we selected the candidate locus among the 11 clusters showing the largest odds ratio (OR) and genotyped additional 17 SNPs in our pooling cohort (80 PEX, 80 PEXG and 80 healthy individuals) using TaqMan assays (Applied Biosystems). To replicate association findings, we genotyped these SNPs in further 610 unrelated German PEX/PEXG patients and 364 controls as well as in 249 Italian patients and 190 Italian controls. Genotype data for four arbitrarily selected SNPs was verified by direct sequencing of 24 randomly chosen samples. The average genotyping rate was 97.5%. Hardy–Weinberg equilibrium for all SNPs was confirmed in the control group using the program Haploview.32
CNTNAP2 expression analyses
For quantitative real-time PCR and immunohistochemistry, ocular tissues were obtained from six donor eyes with PEX syndrome without glaucoma (mean age, 81.3±5.3 years; two women, four men) and six normal-appearing donor eyes (mean age, 77.3±6.6 years; three women, three men) without any known ocular disease. The presence of PEX syndrome was confirmed by electron microscopic analysis of small tissue sectors and the absence of glaucoma was confirmed by microscopic analysis of optic nerve cross sections. These eyes were obtained at autopsy and were processed within 8 h after death. In addition, tissues of six eyes with PEX-associated glaucoma (mean age, 81.6±3.1 years; four women, two men) and six glaucomatous eyes without evidence of PEX (mean age, 78.4±7.8 years; four women, two men) were used. These eyes had to be surgically enucleated because of painful absolute glaucoma and were processed immediately after enucleation; their medical history was documented previously.33 Informed consent of tissue donation was obtained from the patients or, in case of autopsy eyes, from their relatives.
For quantitative real-time PCR total RNA was extracted from various ocular tissues using the RNeasy kit (Qiagen) including an on-column DNase I digestion step. cDNA synthesis and quantitative real-time PCR, using the MyiQ Thermal Cycler and software (Bio-Rad, Munich, Germany), was performed as previously described.34 In brief, real-time PCR reactions (25 μl) contained 2 μl of the 1:5 diluted first-strand cDNA, 0.4 μ M each of upstream and downstream primers, 3.5 mM MgCl2 (CNTNAP2) or 3.0 mM MgCl2 (β-actin) and IQ SYBR Green Supermix (Bio-Rad). All samples were analyzed in duplicates. Gene-specific exon-spanning primers (MWG Biotech, Anzing, Germany) for CNTNAP2 (upstream: CTACTGTAACTGCGACGCGGACT, downstream: CGGTCAGTATCTCCAACCACCAC; GenBank accession no. NM_014141) and β-actin (upstream: TGAGGCACTCTTCCAGCCTTC, downstream: GCGTACAGGTCTTTGCGGATG; GenBank accession no. NM_001101) were designed by Primer 3 software. For quantification, standard curves of plasmid-cloned cDNA were run in parallel, and amplification specificity was checked using melt curve and sequence analyses using the Prism 3100 DNA sequencer (Applied Biosystems). For normalization of gene expression levels, mRNA ratios relative to the housekeeping gene β-actin were calculated.
Light microscopic indirect immunofluorescence and electron microscopic immunogold labeling were performed on sections of ocular tissues of PEX and control eyes, as previously described,35 using two antibodies against CNTNAP2 produced in rabbit (ab33994; Abcam, Cambridge, UK and HPA002739; Sigma, St Louis, MO, USA). Antibody binding was detected by Alexa 488-conjugated secondary antibodies (Invitrogen-Molecular Probes, Eugene, OR, USA) or by 10 nm gold-conjugated secondary antibodies (BioCell, Cardiff, UK). Nuclear counterstaining was performed with propidium iodide (Sigma). In negative control experiments, the primary antibodies were replaced by phosphate-buffered saline or equimolar concentrations of irrelevant antibodies.
Statistical analysis
Statistical analysis was performed with the program Haploview (version 4.0)32 which uses χ2-statistics for assessing single SNP and haplotype association. A P-value of <0.05 was considered statistically significant. Obtained P-values were Bonferroni corrected.
Results
Genome-wide association study
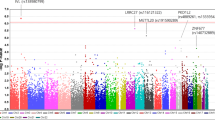
Two genomic DNA pools of PEX patients without and with glaucoma and one of control subjects, prepared as described in Materials and methods, were hybridized to the Affymetrix 500K chip array. For the analysis of pooling-based array data, we used GPFrontend and GPGraphics together with the GenePool software to rank all 500 568 array SNPs according to differences of RAS between cases and controls (ΔRasMean). As expected, a large number of chromosomal regions presented SNPs with marked differences in hybridization intensities including the LOXL1 locus (although SNPs contained in Affymetrix chip are different from those of the Illumina chip in particular for LOXL1 region; Supplementary Figure 1), which has been shown to be associated with the disease.12, 23 The identification of LOXL1 locus among others loci proved the validity of data.
For further analysis we then selected 11 novel clusters of high-ranking SNPs containing at least two SNPs positioned among the first 1000 SNPs and in high LD with each other. We performed individual genotyping with 19 SNPs selected from the clusters. As shown in Table 1, 17 of the 19 SNPs tested showed the expected significant differences (P<0.05) in allelic frequency between patients and control groups, thus confirming that the hybridization intensities of array data were based on true differences in genotype frequency between both groups.
Cluster 8 presenting the highest OR (2.35) was selected for more detailed association analysis. This cluster is located on chromosome 7 within the CNTNAP2 gene locus (Figure 1). A total of 17 additional SNPs from this gene region were genotyped using the DNA samples of the pooled groups (Table 2). For two SNPs the difference in allele frequency between patients and control subjects remained significant after Bonferroni correction (rs2107856 and rs2141388; Bonferroni-corrected Pc=0.0180, OR=2.35, 95% CI: 1.44–3.83; Table 2).
To confirm these results, we genotyped all 19 selected SNPs in a first independent German replication cohort of 610 patients and 364 control subjects. The results of the first replication cohort alone as well as the results of the combined German cohorts are reported in Table 2. Also in the combined groups, the P-values for the two SNPs, rs2107856 and rs2141388, remained significant after Bonferroni correction (rs2107856: Pc=0.0108, OR=1.42, 95% CI: 1.17–1.72; rs2141388: Pc=0.0072, OR=1.43, 95% CI: 1.18–1.74; Table 2), thus suggesting a possible association both with PEX syndrome and PEXG.
A second replication cohort of 249 patients and 190 healthy individuals from Northern Italy was also genotyped for all 19 SNPs (Supplementary Table). In this second population group no evidence for association was detected.
Because a strong association of variants in LOXL1 in PEX and PEXG was reported,12 we compared the allele frequencies at CNTNAP2 locus conditioned on the presence of the identified LOXL1 common risk haplotype G-G. We stratified our German cohorts (discovery, replication and combined cohorts) for carriers and noncarriers of the risk haplotype G-G, which is composed of the major alleles G of the two coding SNPs rs1048661 and rs3825942 (Table 3). Although the subgroups of non-G-G carriers were small sized, we observed significant association to allele T of CNTNAP2 SNPs rs2107856 and rs2141388 in carriers of LOXL1 nonrisk haplotypes (discovery group: P=0.0043, both and combined groups: P=0.0098 and 0.0085; Table 3).
Expression of CNTNAP2
To further investigate the functional role of CNTNAP2 in the human eye and in the pathogenesis of PEX syndrome/glaucoma, we performed a comprehensive mRNA and protein expression analysis in ocular tissues from eyes with PEX syndrome/glaucoma and normal and glaucomatous control eyes. Quantitative real-time PCR showed a ubiquitous expression of CNTNAP2 in virtually all ocular tissues (Figure 2a). No statistically significant differences in expression levels were detected between PEX and control tissues. However, there was a tendency toward reduced CNTNAP2 expression in glaucomatous retinae of both PEXG eyes compared with PEX eyes (3.5±1.8 × 104 vs 6.2±5.1 × 104 copies per μg RNA) and of non-PEXG eyes compared with normal control eyes (1.8±1.1 × 104 vs 3.8±1.3 × 104 copies per μg RNA) (Figure 2b).
Quantitative determination of CNTNAP2 mRNA expression levels in human ocular tissues using real-time PCR technology. The expression levels were normalized against β-actin and the results are expressed as copy number CNTNAP2/μg total RNA. (a) CNTNAP2 mRNA expression in various ocular tissues of normal human donor eyes (n=3). (b) CNTNAP2 mRNA expression in ciliary processes (n=6) and retina (n=3) comparing normal human donor eyes to eyes from patients with glaucoma, PEX syndrome (PEXS) and PEX glaucoma late (PEXGL).
By immunohistochemistry, CNTNAP2 could be localized mainly to cell membranes of epithelial and endothelial cells (Figure 3a), the corneal endothelium (Figure 3b), trabecular endothelial cells and endothelial cells lining Schlemm's canal (Figure 3c), iris pigment epithelium as well as the ciliary epithelium of both PEX and control eyes (Figure 3e). CNTNAP2 was also found to be expressed by vascular endothelial and smooth muscle cells, for example, in the conjunctival stroma, iris stroma (Figure 3d) and choroidal stroma (Figure 3f). In the retina, a marked labeling of retinal ganglion cells and retinal nerve fibers was present (Figure 3g). Moreover, positive staining of nerve fibers and glial cells was observed in the retrolaminar portion of the optic nerve (Figure 3h). On the electron microscopic level, immunogold labeling confirmed a distinct reaction of cell membranes, for example the basal membrane infoldings of nonpigmented ciliary epithelial cells, with both antibodies to CNTNAP2 (Figure 4a). In PEX tissues, immunogold labeling of cell membranes appeared to be reduced close to cellular surface compartments from which the PEX fibrils appeared to emerge (Figure 4b).
Immunofluorescence labeling of CNTNAP2 in ocular tissues of normal human donor eyes. CNTNAP2 immunopositivity (green fluorescence, arrows; red fluorescence, propidium iodide) is observed along basal cell membranes of the limbal epithelium (a), apical cell membranes of the corneal endothelium (b), endothelial cells of trabecular meshwork and Schlemm's canal (c), vascular endothelia and dilator muscle cells in the iris (d), basal and apical membranes of the ciliary epithelium (e), vascular endothelia and smooth muscle cells of the choroid (f), retinal ganglion cells, nerve fibers and bipolar cells in the retina (g), and glial cells (arrows) and nerve fiber bundles in the optic nerve (h). BV, blood vessel; CE, ciliary epithelium; DIL, dilator muscle; DM, Descemet's membrane; GCL, retinal ganglion cell layer; INL, inner nuclear layer; IPE, iris pigment epithelium; LE, limbal epithelium; NFB, nerve fiber bundles; NFL, nerve fiber layer; ONL, outer nuclear layer; PRL, photoreceptor layer; RPE, retinal pigment epithelium; SC, Schlemm's canal; ST, stroma; TM, trabecular meshwork; original magnification, × 100.
Immunogold localization of CNTNAP2 along the basolateral membrane infoldings of the nonpigmented ciliary epithelium of a normal donor eye (a) and an eye with pseudoexfoliation syndrome (b). BM, basement membrane; M, mitochondria; PEX, pseudoexfoliation fibrils; LY, lysosomes; magnification bar=0.5 μm).
In negative control experiments, immunoreactions could be eliminated by replacement of primary antibodies (data not shown).
Discussion
In this study we combined the strengths of microarrays and pooled DNA for a GWAS to identify loci associated with PEX disease. This approach enabled us to investigate a large number of SNPs and individual DNA samples (160 patients and 80 controls) reducing both time and cost normally connected with GWAS of a large number of individual DNA samples. With the help of the newly developed software tools, we were able to detect several loci that showed marked differences in allelic signal frequencies between patients with PEX syndrome/glaucoma and control subjects, including the LOXL1 locus. In the Affymetrix 500K Array Set the best SNP of the LOXL1 locus, rs893818, ranked only at position 2419 with a difference in RAS frequencies (ΔRasMean) of 11.9% between case and control pools (Supplementary Figure 1). Single genotyping of the known associated LOXL1 SNPs, rs1048661 and rs3825942 (not present on the Affymetrix 500K Array Set) with the same DNA samples of the pools revealed a genotypic difference of 15.6% for SNP rs1048661 and 7.2% for SNP rs3825942 (data not shown). Thus the difference at SNP rs893818 observed in the pooling experiment is comparable to that of the two known LOXL1 associated SNPs. Patient selection for pooling was carried out independently of LOXL1 genotype. In addition, single genotyping of the 19 high-ranking SNPs at the 11 selected regions confirmed the differences in allelic signal frequencies observed between groups, thus validating the potential of the pooling-based GWAS to identify SNPs differing in allele frequencies.
Detailed analysis of the selected locus on chromosome 7 suggested an association between PEX and two SNPs, rs2107856 and rs2141388, located in intron 11 of the CNTNAP2 gene. This association was observed both in the discovery and replication cohorts of German patients. Though modest given the OR of about 1.4, the risk conferred to the disease of these two SNPs is typical for many susceptibility variants associated with complex diseases.
In contrast, no association with these SNPs was evident in the Italian patients’ cohort. Insufficient statistical power due to the limited number of Italian subjects available or other factors such as population specific differences in disease-susceptibility loci could explain this difference. Further replication in other study groups is nevertheless required. Moreover, the detected association is more frequently observed in PEX patients carrying LOXL1 nonrisk haplotypes. This might suggest an additionally LOXL1-independent mechanism.
CNTNAP2 is a large gene spanning 2.3 mb of DNA on chromosome 7 and is coding for contactin-associated protein-like 2 (CNTNAP2, also termed Caspr2) a neuronal membrane protein and member of the neurexin superfamily,36, 37 possibly involved in potassium channel trafficking.36, 38 So far it has been suggested as a candidate gene for various neuropsychiatric disorders,39, 40, 41, 42, 43, 44, 45 however little is known about its specific function and regulation. This protein contains multiple interaction domains such as laminin G, fibrinogen-like domains and epidermal growth factor (EGF)-repeat domains that are also found in members of the fibrillins and fibulins protein families. It is interesting to note that the two associated SNPs are located in an intronic region with no annotated functional element, but next to exons coding for the EGF-repeat domains, known to be involved in specific protein–protein interactions.46 Interaction of CNTNAP2 with cytoskeletal components may thus be important for membrane stabilization.
Because, to the best of our knowledge, studies on ocular expression of CNTNAP2 are limited to the rat optic nerve,47 we documented in this study the detailed expression pattern of CNTNAP2 in human ocular tissues. In agreement with published data on neuronal expression, CNTNAP2 mRNA and protein expression levels were found to be highest in the retina in clear association with retinal ganglion cells and axons. In addition, CNTNAP2 protein was found to be ubiquitously expressed in virtually all ocular tissues, where it predominantly localized to cell membranes of epithelial, endothelial, vascular, smooth muscle, glial and neuronal cells. Although no significant differences in expression levels could be observed between PEX and control tissues, there was a tendency toward reduced expression of CNTNAP2 mRNA in retinal tissues of glaucomatous eyes, independent of the presence of PEX. This preliminary observation has to be substantiated by analyzing larger numbers of glaucoma and non-glaucoma specimens.
Immunoelectron microscopy revealed, however, a reduced membrane-associated staining for CNTNAP2 at sites of de novo formation of PEX fibrils. PEX fibrils have been reported to be multifocally produced by various intraocular cell types including the nonpigmented ciliary epithelium, the iris pigment epithelium, the trabecular and Schlemm's canal endothelium, the corneal endothelium as well as vascular endothelial and smooth muscle cells of the iris.2 Based on the observation that CNTNAP2 is expressed along the membranes of cells that are involved in the de novo formation of PEX fibrils in anterior segment tissues, and in view of the fact that PEX fibrillogenesis and assembly occur in close association with cell membranes, a functional implication of CNTNAP2 in PEX pathophysiology is well conceivable. It may be speculated that alterations in membrane stabilization or ion channel function could contribute to these abnormal matrix processes, which are in fact known to be associated with cell-surface irregularities, basement membrane destruction and degenerative membrane alterations.
Although the precise role of CNTNAP2 in the pathogenesis of PEX syndrome and PEXG remains to be elucidated, the present pooling-based genetic approach together with expression and localization patterns in ocular tissues provides evidence for CNTNAP2 as a putative candidate gene for PEX disease. Nevertheless, investigations of the associated SNPs in other populations are needed to further strengthen the association of CNTNAP2 with PEX syndrome.
References
Schlotzer-Schrehardt U, Naumann GO : Ocular and systemic pseudoexfoliation syndrome. Am J Ophthalmol 2006; 141: 921–937.
Ritch R, Schlotzer-Schrehardt U : Exfoliation syndrome. Surv Ophthalmol 2001; 45: 265–315.
Damji KF, Bains HS, Stefansson E et al: Is pseudoexfoliation syndrome inherited? A review of genetic and nongenetic factors and a new observation. Ophthalmic Genet 1998; 19: 175–185.
Orr AC, Robitaille JM, Price PA et al: Exfoliation syndrome: clinical and genetic features. Ophthalmic Genet 2001; 22: 171–185.
Ovodenko B, Rostagno A, Neubert TA et al: Proteomic analysis of exfoliation deposits. Invest Ophthalmol Vis Sci 2007; 48: 1447–1457.
Sharma S, Chataway T, Burdon KP et al: Identification of LOXL1 protein and Apolipoprotein E as components of surgically isolated pseudoexfoliation material by direct mass spectrometry. Exp Eye Res 2009; 89: 479–485.
Burdon KP, Sharma S, Hewitt AW et al: Genetic analysis of the clusterin gene in pseudoexfoliation syndrome. Mol Vis 2008; 14: 1727–1736.
Krumbiegel M, Pasutto F, Mardin CY et al: Exploring functional candidate genes for genetic association in german patients with pseudoexfoliation syndrome and pseudoexfoliation glaucoma. Invest Ophthalmol Vis Sci 2009; 50: 2796–2801.
Ritland JS, Utheim TP, Utheim OA et al: Effects of APOE and CHRNA4 genotypes on retinal nerve fibre layer thickness at the optic disc and on risk for developing exfoliation syndrome. Acta Ophthalmol Scand 2007; 85: 257–261.
Yilmaz A, Tamer L, Ates NA, Camdeviren H, Degirmenci U : Effects of apolipoprotein E genotypes on the development of exfoliation syndrome. Exp Eye Res 2005; 80: 871–875.
Krumbiegel M, Pasutto F, Mardin CY et al: Apolipoprotein E genotypes in pseudoexfoliation syndrome and pseudoexfoliation glaucoma. J Glaucoma, In press.
Thorleifsson G, Magnusson KP, Sulem P et al: Common sequence variants in the LOXL1 gene confer susceptibility to exfoliation glaucoma. Science 2007; 317: 1397–1400.
Fan BJ, Pasquale L, Grosskreutz CL et al: DNA sequence variants in the LOXL1 gene are associated with pseudoexfoliation glaucoma in a US clinic-based population with broad ethnic diversity. BMC Med Genet 2008; 9: 5.
Challa P, Schmidt S, Liu Y et al: Analysis of LOXL1 polymorphisms in a United States population with pseudoexfoliation glaucoma. Mol Vis 2008; 14: 146–149.
Ramprasad VL, George R, Soumittra N, Sharmila F, Vijaya L, Kumaramanickavel G : Association of non-synonymous single nucleotide polymorphisms in the LOXL1 gene with pseudoexfoliation syndrome in India. Mol Vis 2008; 14: 318–322.
Aragon-Martin JA, Ritch R, Liebmann J et al: Evaluation of LOXL1 gene polymorphisms in exfoliation syndrome and exfoliation glaucoma. Mol Vis 2008; 14: 533–541.
Mossbock G, Renner W, Faschinger C, Schmut O, Wedrich A, Weger M : Lysyl oxidase-like protein 1 (LOXL1) gene polymorphisms and exfoliation glaucoma in a Central European population. Mol Vis 2008; 14: 857–861.
Mori K, Imai K, Matsuda A et al: LOXL1 genetic polymorphisms are associated with exfoliation glaucoma in the Japanese population. Mol Vis 2008; 14: 1037–1040.
Fuse N, Miyazawa A, Nakazawa T, Mengkegale M, Otomo T, Nishida K : Evaluation of LOXL1 polymorphisms in eyes with exfoliation glaucoma in Japanese. Mol Vis 2008; 14: 1338–1343.
Gong WF, Chiang SW, Chen LJ et al: Evaluation of LOXL1 polymorphisms in primary open-angle glaucoma in southern and northern Chinese. Mol Vis 2008; 14: 2381–2389.
Hayashi H, Gotoh N, Ueda Y, Nakanishi H, Yoshimura N : Lysyl oxidase-like 1 polymorphisms and exfoliation syndrome in the Japanese population. Am J Ophthalmol 2008; 145: 582–585.
Lemmela S, Forsman E, Onkamo P et al: Association of LOXL1 gene with Finnish exfoliation syndrome patients. J Hum Genet 2009; 54: 289–297.
Pasutto F, Krumbiegel M, Mardin CY et al: Association of LOXL1 common sequence variants in German and Italian patients with pseudoexfoliation syndrome and pseudoexfoliation glaucoma. Invest Ophthalmol Vis Sci 2008; 49: 1459–1463.
Macgregor S, Zhao ZZ, Henders A, Nicholas MG, Montgomery GW, Visscher PM : Highly cost-efficient genome-wide association studies using DNA pools and dense SNP arrays. Nucleic Acids Res 2008; 36: e35.
Craig JE, Hewitt AW, McMellon AE et al: Rapid inexpensive genome-wide association using pooled whole blood. Genome Res 2009; 19: 2075–2080.
Johnson C, Drgon T, Liu QR et al: Pooled association genome scanning for alcohol dependence using 104 268 SNPs: validation and use to identify alcoholism vulnerability loci in unrelated individuals from the collaborative study on the genetics of alcoholism. Am J Med Genet B Neuropsychiatr Genet 2006; 141B: 844–853.
Butcher LM, Meaburn E, Dale PS et al: Association analysis of mild mental impairment using DNA pooling to screen 432 brain-expressed single-nucleotide polymorphisms. Mol Psychiatry 2005; 10: 384–392.
Butcher LM, Davis OS, Craig IW, Plomin R : Genome-wide quantitative trait locus association scan of general cognitive ability using pooled DNA and 500K single nucleotide polymorphism microarrays. Genes Brain Behav 2008; 7: 435–446.
Comabella M, Craig DW, Camina-Tato M et al: Identification of a novel risk locus for multiple sclerosis at 13q31.3 by a pooled genome-wide scan of 500,000 single nucleotide polymorphisms. PLoS ONE 2008; 3: e3490.
Hinds DA, Seymour AB, Durham LK et al: Application of pooled genotyping to scan candidate regions for association with HDL cholesterol levels. Hum Genomics 2004; 1: 421–434.
Pearson JV, Huentelman MJ, Halperin RF et al: Identification of the genetic basis for complex disorders by use of pooling-based genomewide single-nucleotide-polymorphism association studies. Am J Hum Genet 2007; 80: 126–139.
Barrett JC, Fry B, Maller J, Daly MJ : Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 2005; 21: 263–265.
Schlotzer-Schrehardt U, Pasutto F, Sommer P et al: Genotype-correlated expression of lysyl oxidase-like 1 in ocular tissues of patients with pseudoexfoliation syndrome/glaucoma and normal patients. Am J Pathol 2008; 173: 1724–1735.
Zenkel M, Kruse FE, Junemann AG, Naumann GO, Schlotzer-Schrehardt U : Clusterin deficiency in eyes with pseudoexfoliation syndrome may be implicated in the aggregation and deposition of pseudoexfoliative material. Invest Ophthalmol Vis Sci 2006; 47: 1982–1990.
Schlotzer-Schrehardt U, Zenkel M, Nusing RM : Expression and localization of FP and EP prostanoid receptor subtypes in human ocular tissues. Invest Ophthalmol Vis Sci 2002; 43: 1475–1487.
Poliak S, Gollan L, Martinez R et al: Caspr2, a new member of the neurexin superfamily, is localized at the juxtaparanodes of myelinated axons and associates with K+ channels. Neuron 1999; 24: 1037–1047.
Einheber S, Zanazzi G, Ching W et al: The axonal membrane protein Caspr, a homologue of neurexin IV, is a component of the septate-like paranodal junctions that assemble during myelination. J Cell Biol 1997; 139: 1495–1506.
Horresh I, Poliak S, Grant S, Bredt D, Rasband MN, Peles E : Multiple molecular interactions determine the clustering of Caspr2 and Kv1 channels in myelinated axons. J Neurosci 2008; 28: 14213–14222.
Verkerk AJ, Mathews CA, Joosse M, Eussen BH, Heutink P, Oostra BA : CNTNAP2 is disrupted in a family with Gilles de la Tourette syndrome and obsessive compulsive disorder. Genomics 2003; 82: 1–9.
Friedman JI, Vrijenhoek T, Markx S et al: CNTNAP2 gene dosage variation is associated with schizophrenia and epilepsy. Mol Psychiatry 2008; 13: 261–266.
Alarcon M, Abrahams BS, Stone JL et al: Linkage, association, and gene-expression analyses identify CNTNAP2 as an autism-susceptibility gene. Am J Hum Genet 2008; 82: 150–159.
Arking DE, Cutler DJ, Brune CW et al: A common genetic variant in the neurexin superfamily member CNTNAP2 increases familial risk of autism. Am J Hum Genet 2008; 82: 160–164.
Bakkaloglu B, O’Roak BJ, Louvi A et al: Molecular cytogenetic analysis and resequencing of contactin associated protein-like 2 in autism spectrum disorders. Am J Hum Genet 2008; 82: 165–173.
Strauss KA, Puffenberger EG, Huentelman MJ et al: Recessive symptomatic focal epilepsy and mutant contactin-associated protein-like 2. N Engl J Med 2006; 354: 1370–1377.
Zweier C, de Jong EK, Zweier M et al: CNTNAP2 and NRXN1 are mutated in autosomal-recessive Pitt-Hopkins-like mental retardation and determine the level of a common synaptic protein in Drosophila. Am J Hum Genet 2009; 85: 655–666.
Nakayama M, Nakajima D, Nagase T, Nomura N, Seki N, Ohara O : Identification of high-molecular-weight proteins with multiple EGF-like motifs by motif-trap screening. Genomics 1998; 51: 27–34.
Brown AA, Xu T, Arroyo EJ et al: Molecular organization of the nodal region is not altered in spontaneously diabetic BB-Wistar rats. J Neurosci Res 2001; 65: 139–149.
Acknowledgements
We thank all patients and control individuals for participation in this study; Juliane Niedziella for invaluable help with patient recruitment; Olga Zwenger and Petra Rothe for expert technical assistance. This study was supported by Grant no. SFB 539 from the German Research Foundation
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Rights and permissions
About this article
Cite this article
Krumbiegel, M., Pasutto, F., Schlötzer-Schrehardt, U. et al. Genome-wide association study with DNA pooling identifies variants at CNTNAP2 associated with pseudoexfoliation syndrome. Eur J Hum Genet 19, 186–193 (2011). https://doi.org/10.1038/ejhg.2010.144
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2010.144
Keywords
This article is cited by
-
Analysis of genetically determined gene expression suggests role of inflammatory processes in exfoliation syndrome
BMC Genomics (2023)
-
Is pseudoexfoliation glaucoma a neurodegenerative disorder?
Journal of Biosciences (2021)
-
De novo variants in an extracellular matrix protein coding gene, fibulin-5 (FBLN5) are associated with pseudoexfoliation
European Journal of Human Genetics (2019)
-
Evaluation of CNTNAP2 gene rs2107856 polymorphism in Turkish population with pseudoexfoliation syndrome
International Ophthalmology (2019)
-
A common variant mapping to CACNA1A is associated with susceptibility to exfoliation syndrome
Nature Genetics (2015)