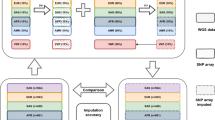
Abstract
Genetic matching potentially provides a means to alleviate the effects of incomplete Mendelian randomization in population-based gene–disease association studies. We therefore evaluated the genetic-matched pair study design on the basis of genome-wide SNP data (309 790 markers; Affymetrix GeneChip Human Mapping 500K Array) from 2457 individuals, sampled at 23 different recruitment sites across Europe. Using pair-wise identity-by-state (IBS) as a matching criterion, we tried to derive a subset of markers that would allow identification of the best overall matching (BOM) partner for a given individual, based on the IBS status for the subset alone. However, our results suggest that, by following this approach, the prediction accuracy is only notably improved by the first 20 markers selected, and increases proportionally to the marker number thereafter. Furthermore, in a considerable proportion of cases (76.0%), the BOM of a given individual, based on the complete marker set, came from a different recruitment site than the individual itself. A second marker set, specifically selected for ancestry sensitivity using singular value decomposition, performed even more poorly and was no more capable of predicting the BOM than randomly chosen subsets. This leads us to conclude that, at least in Europe, the utility of the genetic-matched pair study design depends critically on the availability of comprehensive genotype information for both cases and controls.
Similar content being viewed by others
Introduction
In both classical epidemiology and clinical research, potential confounders are usually controlled for by one of two different means, matching or randomization. In genetic studies, however, including the large number of genome-wide association (GWA) studies that have recently been published,1, 2, 3 only so-called ‘Mendelian’ randomization has been employed to control for genetic confounders, whereas matching by genotype has not played an important role.4 Nevertheless, there has always been some awareness among genetic epidemiologists that Mendelian randomization may fail, thereby leading to false positive reports of disease genes or to biased effect size estimates.5 One possible cause of such failure may be systematic differences in terms of the rate at which individuals with a particular phenotype or genotype are sampled from genetically distinct populations. Therefore, two statistical methods to retrospectively rectify genetic imbalances in case-control studies were developed in the late 1990s, both of which rely upon genotyping loci that are unrelated to the genetic variants under study (ie unlinked and not in linkage disequilibrium). The ‘genomic control’ approach6 uses marker genotypes to correct the employed test statistic, whereas ‘structured association’7 infers the number of populations represented in a sample, and then assigns each individual to one of these populations with a certain probability.
With the possibility to effectively genotype large numbers of single nucleotide polymorphisms (SNPs) in large numbers of individuals, using microarray technology,8 the effects of imperfect Mendelian randomization can, in principle, also be alleviated by genetic matching. If individuals from different samples such as cases and controls were as closely matched as possible in terms of their identity-by-state (IBS) status at a large number of SNPs, it may be surmised that most systematic population genetic differences would be eliminated between the ensuing sub samples. However, genetic matching would have to be based on markers from outside the genomic region under study to avoid over-matching. This implies that, in practise, repeated matching may be necessary if multiple or even GWA assessments are due. In any case, genetic matching could of course be accomplished efficiently with the use of genome-wide microarray data, but such a costly strategy may not be necessary if a set of ‘best genetic match’ (BGM) markers could be established in advance that are capable of capturing the major population genetic characteristics of relevant extant populations. Once a set of BGM markers has been found, it can be used in two ways: either to retrospectively confirm whether two samples of interest were genetically well-matched or to select members of matched samples prospectively, before any additional genotyping.
Recruitment of phenotypically well-characterized control samples is one of the major bottlenecks of genetic epidemiological and pharmacogenetic research. The use of common controls across different association studies has proven to be an efficient solution to this problem, pioneered at a local level by the Wellcome Trust Case Control Consortium (WTCCC),3 and since adopted, for example, by the US-American Genetic Association Information Network (GAIN)1 and the German National Genome Research Network (‘Nationales Genomforschungsnetz’, NGFN).9 However, the number and geographical distribution of control samples required for the common controls approach to be feasible at a broader geographical level are currently unknown.
In the present study, we investigated three issues related to the genetic-matched pair study design, using genome-wide SNP data from across Europe: (1) the prospects of identifying a small subset of SNPs that accurately predict the ‘best’ genome-wide matching partner of a given individual, (2) the distribution of ‘best’ genetic-matching partners between the European subpopulations and (3) the inter-individual variability in terms of the uniqueness of the ‘best’ genetic-matching partner. To this end, we analyzed the genotypes of 309 790 markers obtained from the GeneChip Human Mapping 500K Array Set in 2457 individuals, ascertained at one of 23 recruitment sites. The European population is important in this context, not only because of the historical interest in these people and their descendants in the Americas, Australia and elsewhere, but also because they are a major focus of both genetic epidemiological and pharmacogenetic research.1, 3
Material and methods
Samples, genotyping and quality control
The GeneChip Human Mapping 500K Array (Affymetrix) was used to genotype 500 568 SNPs in 2514 individuals from 23 different sampling sites (henceforth, termed ‘subpopulations’), distributed over 20 different European countries. Subpopulation sizes ranged from 12 to 500 individuals (Table 1). Sex ratios differed markedly between subpopulations, with some comprising only females or males, respectively. Genotyping was carried out at six different facilities. For further details, see Lao et al.10
Array-based SNP genotypes were subjected to stringent quality control as described earlier.10 Briefly, markers, which had a genotype call rate ≥93%, were monomorphic, located on the X chromosome or had a per marker call rate ≤90% in at least one genotyping facility were excluded, as were those showing a significant (P<0.05) deviation from Hardy–Weinberg equilibrium (HWE) in at least one subpopulation. Individuals deemed genetic outliers to their subpopulation of origin, based on low average IBS to the remaining individuals, were omitted from the respective subpopulation. In total, quality control left 2457 individuals (97.6%) and 309 790 markers (62.4%) for inclusion in subsequent analyses. The set of quality controlled markers will henceforth be referred to as marker set C. Ascertainment of a marker set for genetic matching was carried out with internal validation, using 2/3 of the members of each subpopulation (ie, 1638 randomly chosen individuals) as the training set, and using the remainder (819 individuals) as the validation set (Table 1).
All data were stored as either flat files or in a customized database with an interface to the R statistical software. All data analysis, except for the IBS estimation, was done in R version 2.4.111 using customized scripts. IBS calculations and selection of marker sets were carried out using custom C++ programs. All software is available from the authors on request.
Best genetic match marker set
For the ascertainment of a marker subset M of C that would allow us to identify ‘best’ genetic-matching partners, we will use a set-specific criterion, Δ(M) that is related to the IBS between given individuals and their matching partners, as selected on the basis of M (see below). In this context, we will use the term ‘best overall match’ (BOM) to denote that individual or group of individuals who maximize the average pair-wise IBS with the individual of interest for the complete marker set C. Ideally, we would want to ascertain a subset of markers that consistently lead to the selection of matching partners with an IBS with the reference individual that is close to the IBS between the reference individual and its BOM.
More formally, if the genotype (g), of a given SNP is encoded by the dose of one of its two alleles (ie, as 0, 1 or 2), then the IBS between any two individuals x and y equals 1−∣g(x)−g(y)∣/2 for that SNP. Here, g(x) and g(y) denote the genotypes of x and y, respectively. For a marker set M, let iM(x,y) be the average IBS, taken over all markers in M, and let iM(x) denote the maximum iM(x,y), taken over all individuals y other than x. Finally, if M⊆N are two nested marker sets, let iM,N(x) be the average iN(x,y) taken over all y for which iM(x,y)=iM(x). For a marker set M⊆C, Δ(M) is defined as the average difference ∣iC(x)−iM,C(x)∣, taken over all individuals x and weighted by the inverse of the size of the subpopulation to which x belongs.
We used forward selection from marker set C to ascertain marker sets that successively minimized the Δ criterion. The ensuing marker sets will be referred to as the best genetic match (BGM) marker sets. Upper and lower baselines for Δ were computed as follows. The upper baseline was obtained from randomly chosen marker sets of varying size (10–100 in steps of 10), with 1000 sets sampled for each set size value. The lower baseline was obtained from marker sets that theoretically should have captured most of the genetic variation present in the individuals under study, ie sets for which any additional marker would have been in strong linkage disequilibrium with the markers already included. Each chromosome was thus divided into bins of 20 kb, based on the mean swept radius of 500 kb estimated for the European population.12, 13 The swept radius is the distance at which the average association between two markers, measured by r2, is reduced to approximately one-third (more precisely, e−1) of its initial value. A bin size of 20 kb therefore ensures an average r2 of e−10/500=0.98 between markers in the bin. Markers were then randomly selected from bins, one at a time, and Δ calculated for the resulting marker set. The described selection process was repeated 1000 times and the mean Δ value taken as the lower baseline, ie the expectation of Δ at r2-based saturation.
Ancestry-sensitive marker set
To compare the BGM set, which focuses on inter-individual genetic variation with a marker set that was ascertained with the aim to highlight inter-population variation, we generated an ancestry-sensitive marker (ASM) set using the singular value decomposition (SVD) method with redundant marker reduction described by Paschou et al.14, 15 Global allele frequencies were used to interpolate missing data as suggested by the authors. Some 228 individuals were eliminated from the training set during PCA analysis with Eigensoft216 using the standard criterion of having an ancestry coefficient >6 standard deviations in at least one of the eigenvector axes. SVD was carried out with SVDLIBC (version 1.34, http://tedlab.mit.edu/~dr/SVDLIBC), a C library based on the SVDPACK library.17 Rank-revealing QR matrix decomposition was carried out in Octave version 2.0.1718 to reduce the redundancy of the first 5000 markers, ordered by the first SVD eigenvector. This resulted in a set of the same size (ie 100 markers) as the BGM set.
Distribution of best genetic match pairs
A count matrix was generated that contains, for each pair of subpopulations, the number of times an individual in the first subpopulation had their BOM in the second population. Cell counts were tested for a deviation from the null hypothesis that BOMs were drawn randomly from subpopulations using a two-tailed exact test as implemented in the R routine binom.test. A plot of directed graphs representing the relationships between individuals and their BOMs was generated using Graphviz.19
False positive rates
Thresholds for the false positive rates of population-based gene–disease associations in Europe were determined from contrived case-control experiments, using PLINK version 1.0320 on all markers in set C (Fisher's exact test on allele frequencies). These mock studies were carried out for all pair-wise combinations of subpopulations, each time labeling one subpopulation as ‘cases’ and the other as ‘controls’. The percentage of markers with P-values <0.05 was reported. As the variance of the P-value is inversely related to sample size, false positive rates were not estimated for subpopulations with sample sizes <20 (PT, HU and RO; see Table 1 for subpopulation abbreviations).
Results
Best genetic match and ancestry sensitive marker sets
Two subsets of markers (BGM and ASM) were ascertained from the complete marker set using either IBS-based forward selection or SVD with redundant marker reduction, respectively. As the decrease in Δ as a function of marker set size levelled off very rapidly (see Figure 1), BGM marker selection was terminated at 100 SNPs (Supplementary Table 1). For the sake of comparability, the ASM set was chosen so as to contain the same number of markers as the BGM set (Supplementary Table 2). Interestingly, the top 5000 markers of the provisional ASM set included various SNPs annotated to genes known to stratify the European gene pool as a result of recent positive selection acting differently in different geographic regions, including HERC221 (ranked 7), OCA222 (ranked 33), LCT23 (ranked 262) and TYRP124 (ranked 1138).
IBS-based forward selection of best genetic match (BGM) marker sets. The upper baseline for Δ is illustrated by box-whisker plots, each generated from 1000 random selections of a marker set of given size. The lower baseline for Δ (dotted line) is provided by a marker set for which any additional markers could be expected to be in strong linkage disequilibrium (r2>0.98) with at least one marker already included in that set (for details, see text). Selection of the BGM marker sets is depicted by a solid line; the performance of ASM sets of various sizes is illustrated by a dashed line. All Δ values were calculated from the validation set of individuals. The training set Δ values obtained for the BGM marker sets are included for reference (dash-dotted line).
A graphical representation of the forward selection process leading to the BGM set is provided in Figure 1. In the validation set, the Δ criterion decreased by ∼10% until it levelled off at ∼20 markers, and decreased only marginally thereafter. Although forward selection on the training set showed a promising reduction in Δ value, the validation Δ for the 100 top markers comprising the BMG set was still at 9.3 × 10−5, which is 14.3% lower than the upper (random) baseline but exceeds the lower baseline of 1.5 × 10−5 by a factor of six. This implies that the genome-wide similarity of two European individuals is hard to predict with sufficient accuracy on the basis of a small, specifically selected marker set, and that the little benefit that can be gained in this respect already arises from 100 markers or even fewer. By comparison, the capacity of the ASM set for BOM prediction was found to be indistinguishable from the upper (random) baseline, ie, it performed no better than randomly drawn marker sets.
Distribution of best overall matches (BOMs)
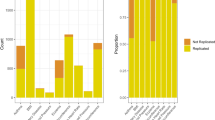
A significant amount of genetic similarity between the European subpopulations is revealed by an assessment of the subpopulation of origin of BOMs (Table 2). In a considerable proportion of cases (1868/2457 or 76.0%), the BOM of a given individual belonged to a different subpopulation than the individual itself. That this was particularly so when individuals or BOMs came from subpopulations with large sample sizes (DE1, DE2 and NL) was presumably due to the wider range of genetic diversity captured by these samples, but may also reflect their concurrent geographic location in central Europe. On the other hand, for some relatively isolated subpopulations (FI and IT2) the source of the BOM was mostly the subpopulation itself, reflecting their separation also seen in genetic barrier analysis and, in the case of the Finns, principle component analysis.10 Closer inspection at the individual level revealed that some individuals were disproportionately more often selected as BOMs than others (Figure 2). Thus, of the 2457 individuals examined, 1860 (75.7%) were never deemed a BOM at all. This is significantly higher than the expected number (1553.3, 63.2%) if BOMs were drawn at random (χ2=165.1, 1 df, P<0.001). At the same time, 120 individuals were chosen as BOMs at least five times, which is a significant excess over expectation (9.0, 0.36%, χ2=1401.9, 1 df, P<0.001). The subpopulation of origin of the 10 most frequently ascertained BOMs was generally among those central Europeans who also had the largest sample size (DE1 five, DE2 two and NL one), with the notable exception of DK (59 individuals, yet holding two of the top 10 positions; Figure 2). Interestingly, barring of the 10 most frequently chosen BOMs left the number of times the BOM was found outside the subpopulation of origin of the individual of interest virtually unchanged (1862/2457 or 75.8%, Supplementary Table 4). A graphical representation of the BOM relationships between individuals is provided in a directed graph illustrating the complexity of networks of matches (Figure 3).
Distribution of the number of times an individual was deemed a BOM. The observed distribution is marked by circles. Also included is a Poisson distribution with the same mean as the sample mean (marked by squares), which approximately corresponds to the theoretical expectation if best overall matching (BOM) were selected at random. The codes of the subpopulation of origin of the 10 most frequently selected BOMs are given at the upper right edge of the plot.
False positive rates
Although it is admittedly unlikely that a researcher would actually carry out a population-based gene–disease association study in which cases and controls were sampled from different countries, without adjusting for population origin in one way or another, measurement of the false positive rates expected from such undertaking is of general interest as a gauge of the magnitude of stratification pertaining in the European population. Mock false positive rates for pairs of subpopulations (Supplementary Table 3) ranged from 0.039 (CZ and PO) to 0.208 (DE1 and IT1), with a median of 0.070. Subpopulations sampled from the same political country often had false positive rates indicative of little or no population stratification, although this was not always the case (DE1–DE2: 0.089). Many neighboring countries also had false positive rates close to those expected under the null hypothesis, indicating the absence of major population differences as well (eg UK-IE: 0.042, NL-DK: 0.051, EL-YU: 0.047, CH-AT: 0.039, FR-DE2: 0.051).
Discussion
This is the first study to evaluate the genetic (ie, IBS-) matched pair study design with genome-wide SNP data of a large number of European individuals from across the continent. The high number of best genetic-matching partners found in different subpopulations corroborates earlier reports of a considerable amount of genetic similarity between the European subpopulations,4, 10, 14, 25, 26, 27 particularly those in close geographic proximity. The surprising inter-individual variability observed in terms of the number of times a person was chosen as the best genetic-matching partner of others does not necessarily imply that the relationship between genetic and geographic distance in a given sample hinges on a small number of people. Thus, when the most frequently chosen matching partners were barred in our analysis, the proportion of best matches found outside the subpopulation of origin of the respective index person remained virtually unchanged.
We observed that the best genetic-matching partner for a genome-wide marker set such as the Affymetrix GeneChip Human Mapping 500K Array cannot be predicted from a small, specifically selected subset of markers alone, but that the information required to make such predictions is distributed evenly across all markers. This leads us to conclude that, at least in Europe, the utility of the genetic-matched pair study design depends critically on the availability of comprehensive genotype information for both cases and controls. In practise, this would mean that shared controls should ideally be genotyped for all relevant genome-wide marker sets, thereby allowing the chromosome-specific choice of best matching partners for given case individuals on the basis of the remainder of the genome.
A distinction must obviously be made between ASM, collections of which have been described in recent papers,14, 25, 26, 27, 28 and the BGM marker set that we attempted to generate. As the genetic within-subpopulation variation in Europe is much greater than the between-subpopulation variation, it is not unlikely for any two individuals from different subpopulations to be genetically more similar to each other than any two individuals from the same subpopulation. In this sense, an ASM marker set consists of markers that differentiate subpopulations, whereas a BGM marker set should contain variants that highlight genetic similarity at the individual level. Although the two concepts are complimentary, the marker sets fit to each task need not be the same, and the existence of one set does not necessitate the existence of the other. Obviously, markers that arose on early branches of the corresponding, region-specific coalescence tree of the extant Europeans would provide good ASM, but they cannot at the same time identify nearest neighbors at the tips of the tree. Such identification requires a much higher resolution of the tree topology, and therefore many more markers. Consequently, no adequately sized BGM set could be constructed in our study and the ASM set selected with established methodology was no more capable of identifying the best genetic-matching partner of an individual than a randomly chosen marker set.
Recently, two independent applications of genetic matching have been reported in the context of GWA studies,4, 29 both of which relied on information derived from PCA of genotypes to match individuals. In the first study, using US-American type 1 diabetes patients and German controls, Luca et al4 carried out ‘full’ matching wherein matches consist of clusters of individuals that contain at least one case and one control. Matching was based upon a distance measure with the top eigenvectors as coordinates, weighted by the eigenvalues to exaggerate differences in dimensions of greater importance. In the second study, Heath et al29 undertook a PCA on a large pan-European group of individuals and proposed a method to predict the population affiliation of a sample of unknown origin from the eigenvector matrix of its genotypes. As both methods are likely to reduce spurious genetic differences between cases and controls in disease association studies, basing their matching criteria on eigenvectors from PCA is strongly reminiscent of selecting ASM. However, as we have shown above, matching with ASM is less efficient than best overall genetic matching particularly in Europe, where the within-subpopulation genetic variation is known to be much greater than the between-subpopulation variation. Indeed, the conclusion by Luca et al4 that some individuals remain ‘unmatchable’ by their approach is not surprising bearing in mind that ASM can only capture a miniscule proportion of the actual inter-individual genetic differences in a given population.
The false positive rates derived in our study from mock genetic case-control experiments represent an upper limit to the likely consequences of sharing samples in continent-wide scientific collaborations. In this respect, the rate estimates also rationalize collaborative genetic epidemiological and pharmacogenetic research in Europe; from the data we have compiled, it seems as if research projects combining cases from neighboring subpopulations and matching them against common control samples, such as those provided by the WTCCC,3 GAIN1 and NGFN,9 may indeed be valid.
In conclusion, we found that the pattern of pair-wise genetic matching in the European population was more complex than anticipated. Best genetic matches occurred frequently across the continent in our study, and disproportionately often involved a small group of individuals. Ascertainment of a subset of markers that accurately predicts best overall genetic matches turned out to be infeasible.
References
GAIN Collaborative Research Group Manolio TA, Rodriguez LL, Brooks L et al: New models of collaboration in genome-wide association studies: the Genetic Association Information Network. Nat Genet 32007; 9: 1045–1051.
Hirschhorn JN : Genetic approaches to studying common diseases and complex traits. Pediatr Res 2005; 57: 74R–77R.
The Wellcome Trust Case Control Consortium: Genome-wide association study of 14 000 cases of seven common diseases and 3000 shared controls. Nature 2007; 447: 661–678.
Luca D, Ringquist S, Klei L et al: On the use of general control samples for genome-wide association studies: genetic matching highlights causal variants. Am J Hum Genet 2008; 82: 453–463.
Davey Smith G, Ebrahim S : What can mendelian randomisation tell us about modifiable behavioural and environmental exposures? BMJ 2005; 330: 1076–1079.
Devlin B, Roeder K : Genomic control for association studies. Biometrics 1999; 55: 997–1004.
Pritchard JK, Stephens M, Donnelly P : Inference of population structure using multilocus genotype data. Genetics 2000; 155: 945–959.
Wang WY, Barratt BJ, Clayton DG, Todd JA : Genome-wide association studies: theoretical and practical concerns. Nat Rev Genet 2005; 6: 109–118.
Wichmann HE, Gieger C, Illig T, MONICA/KORA_Study_Group: KORA-gen – resource for population genetics, controls and a broad spectrum of disease phenotypes. Gesundheitswesen 2005; 67: 26–30.
Lao O, Lu TT, Nothnagel M et al: Correlation between genetic and geographic structure in Europe. Curr Biol 2008; 18: 1241–1248.
R Development Core Team: R: A language and environment for statistical computing. R Foundation for Statistical Computing: Vienna, 2008.
Morton NE, Zhang W, Taillon-Miller P, Ennis S, Kwok PY, Collins A : The optimal measure of allelic association. Proc Natl Acad Sci USA 2001; 98: 5217–5221.
Wollstein A, Herrmann A, Wittig M et al: Efficacy assessment of SNP sets for genome-wide disease association studies. Nucleic Acids Res 2007; 35: e113.
Paschou P, Drineas P, Lewis J et al: Tracing sub-structure in the European American population with PCA-informative markers. PLoS Genet 2008; 4: e1000114+.
Paschou P, Ziv E, Burchard EG et al: PCA-correlated SNPs for structure identification in worldwide human populations. PLoS Genet 2007; 3: 1672–1686.
Patterson N, Price AL, Reich D : Population structure and eigenanalysis. PLoS Genet 2006; 2: e190.
Berry MW : Large scale singular value computations. Int J Supercomput Appl 1992; 6: 13–49.
Eaton JW : GNU Octave Manual. Network Theory Unlimited: Bristol, 2002.
Gansner ER, North SC : An open graph visualization system and its applications to software engineering. Softw Pract Exp 2000; 30: 1203–1233.
Purcell S, Neale B, Todd-Brown K et al: PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 2007; 81: 559–575.
Kayser M, Liu F, Janssens AC et al: Three genome-wide association studies and a linkage analysis identify HERC2 as a human iris color gene. Am J Hum Genet 2008; 82: 411–423.
Duffy DL, Montgomery GW, Chen W et al: A three-single-nucleotide polymorphism haplotype in intron 1 of OCA2 explains most human eye-color variation. Am J Hum Genet 2007; 80: 241–252.
Bersaglieri T, Sabeti PC, Patterson N et al: Genetic signatures of strong recent positive selection at the lactase gene. Am J Hum Genet 2004; 74: 1111–1120.
Voight BF, Kudaravalli S, Wen X, Pritchard JK : A map of recent positive selection in the human genome. PLoS Biol 2006; 4: e72.
Bauchet M, McEvoy B, Pearson LN et al: Measuring European population stratification with microarray genotype data. Am J Hum Genet 2007; 80: 948–956.
Price AL, Butler J, Patterson N et al: Discerning the ancestry of European Americans in genetic association studies. PLoS Genet 2008; 4: e236.
Seldin MF, Shigeta R, Villoslada P et al: European population substructure: clustering of northern and southern populations. PLoS Genet 2006; 2: e143.
Tian C, Hinds DA, Shigeta R et al: A genomewide single-nucleotide-polymorphism panel for Mexican American admixture mapping. Am J Hum Genet 2007; 80: 1014–1023.
Heath SC, Gut IG, Brennan P et al: Investigation of the fine structure of European populations with applications to disease association studies. Eur J Hum Genet 2008; 16: 1413–1429.
Acknowledgements
All sample donors are gratefully acknowledged for their participation. We thank the following colleagues for their help and support: J Kooner and J Chambers of the LOLIPOP study and D Waterworth, V Mooser, G Waeber and P Vollenweider of the CoLaus study for providing access to their collections through the GlaxoSmithKline-sponsored Population Reference Sample (POPRES) project; K King for preparing the POPRES data; M Simoons, E Sijbrands, A van Belkum, J Laven, J Lindemans, E Knipers and B Stricker for their financial contribution to the Rotterdam study; P Arp, M Jhamai, W van IJken and R van Schaik for generating the Rotterdam study dataset; T Meitinger, P Lichtner, G Eckstein and all genotyping staff at the Helmholtz Zentrum München for generating the KORA study dataset; H von Eller-Eberstein for providing access to the PopGen data; R Borup, C Schjerling, H Ullum, E Haastrup and numerous colleagues at the Copenhagen University Hospital Blood Bank for making the Danish data available; and S Brauer for DNA sample management. We also wish to thank Affymetrix for making the GeneChip Human Mapping 500K Array genotypes of the CEPH-CEU trios publicly available, and the Centre d’Etude du Polymorphisme Humain (CEPH) for the original sample collection. This work was supported by the Netherlands Forensic Institute (M Ka), Affymetrix (M Ka and M Kr), the German National Genome Research Network and the German Federal Ministry of Education and Research (H-EW, SS, M Kr and PN); the Helmholtz Zentrum München – German Research Center for Environmental Health, Neuherberg and the Munich Center of Health Sciences as part of LMUinnovativ (H-EW), the Netherlands Organization for Scientific Research (AGU: NWO 175.010.2005.011), the European Commission (AGU: GEFOS; 201865, AS: LD Europe; QLG2-CT-2001-00916); the Czech Ministry of Health (MM: VZFNM 00064203 and IGA NS/9488-3), Helse-Vest, Regional Health Authority Norway (LAB), the Swedish National Board of Forensic Medicine (GH: RMVFoU 99:22, 02:20) and the Academy of Finland (AS: 80578, OMLL, JP: 109265 and 111713). None of the funding organization had any influence on the design, conduct or conclusions of the study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Rights and permissions
About this article
Cite this article
Lu, T., Lao, O., Nothnagel, M. et al. An evaluation of the genetic-matched pair study design using genome-wide SNP data from the European population. Eur J Hum Genet 17, 967–975 (2009). https://doi.org/10.1038/ejhg.2008.266
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2008.266
Keywords
This article is cited by
-
The more the merrier? How a few SNPs predict pigmentation phenotypes in the Northern German population
European Journal of Human Genetics (2016)