Abstract
Hirschsprung's disease (HSCR) is a congenital disorder in which ganglion cells are absent in variable portions of the lower digestive tract according to which patients are classified. The RET gene is the major HSCR gene, although reduced penetrance of RET mutations and variable expression of HSCR phenotype indicates that more than one gene is required. An unidentified RET-dependent modifier on 3p21 appears to be necessary for transmission of the short HSCR (S-HSCR) phenotype. We investigated 6 Mb of the 3p21 region on a quest for the HSCR-susceptibility locus. Fifty-eight S-HSCR case–parent trios were genotyped using Sequenom technology for 214 tag single nucleotide polymorphisms (SNPs) distributed along 6 Mb of the 3p21 region. A five-marker haplotype, spanning a 118 kb gene-rich region, was found to be overtransmitted to affected offspring. The associated haplotype encompasses three genes involved in neurological phenotypes. Importantly, this association was replicated in an independent sample of 172 S-HSCR cases and 153 unrelated controls. Ranking markers by proximity to candidate genes or by expected functional consequences could be used in follow-up studies to finally pinpoint this HSCR locus.
Similar content being viewed by others
Introduction
Hirschsprung's disease (HSCR) is a congenital disorder in which ganglion cells are absent in variable portions of the lower digestive tract. There is significant racial variation in the incidence of the disease and it is most often found among Asians (2.8 per 10 000 life births).1, 2 HSCR patients can be classified according to the severity of the phenotype into long (L-SHCR) and short (S-HSCR) segment aganglionosis. HSCR presents mostly sporadically, although it can be familial with a complex pattern of inheritance, including low, sex-dependent penetrance and phenotypic variability. The male/female ratio (M/F) is ≈4:1 among S-HSCR patients and ≈1:1 among L-HSCR patients. Aganglionosis is attributed to a failure of neural crest cells (enteric ganglion precursors) to migrate, proliferate or survive in the gastrointestinal tract. The RET gene, encoding a tyrosine-kinase receptor, is the major HSCR gene and its expression is crucial for the development of the enteric ganglia.3 Other HSCR genes identified so far mainly code for protein members of interrelated signalling pathways involved in the development of enteric ganglia: RET, endothelin receptor B and the transcriptional regulator SOX10 signalling pathways,4, 5, 6, 7 although mutations in genes other than RET account only for 7% of the cases. Reduced penetrance of RET-coding sequence (CDS) mutations and variable expression of HSCR phenotype indicates that the disease could result from the combined effect of RET and genes of these signalling pathways, whereby the outcome would be altered RET expression.8, 9, 10, 11 Mutations in RET regulatory regions have been shown to contribute to HSCR either alone, or in combination with variants in RET CDS or in other susceptibility genes. Indeed, common RET SNPs encompassing the whole gene, including the promoter, were found strongly associated with HSCR either singly or combined (haplotype).12, 13, 14, 15, 16, 17, 18, 19, 20 Importantly, a functionally relevant SNP was identified within a HSCR-associated RET haplotype.21 This ‘common’ mutation, which lies in a putative enhancer element within RET intron 1, has low penetrance, small sex-dependent effect and explains only a small fraction of the HSCR cases. RET non-coding region mutations allow to speculate that proteins, encoded by yet unknown loci, could modify RET expression and act as disease-promoting or -suppressing genes (modifiers).16, 21 HSCR can, therefore, be defined as a complex disorder with multifactorial inheritance that requires RET and other interacting disease susceptibility alleles. As in many other complex diseases, the manifestation of the phenotype may result from the combination of common variants (SNPs) in several genes. Additional genes are necessary to explain not only the disease incidence but also its complex pattern of inheritance. Current data indicate that S-HSCR manifestation requires the effect of RET (major gene with large effect) and that of two unidentified RET-dependent modifiers mapped to 3p21 and 19q12.22 The 3p21 region was contained in the 70 Mb of the p arm of chromosome 3, which was genotyped by the Hong Kong HapMap Group for the HapMap project.23 We identified a high LD gene-rich region (6 Mb) comprised in 3p21 and subsequently, we investigated the region on a quest for the HSCR-susceptibility locus. Elucidating HSCR is especially relevant in China where Hirschsprung's disease has one of the highest incidences in the world.
Materials and methods
Samples
The institutional review board of The University of Hong Kong together with the Hospital Authority granted ethical approval for this project (IRB: UW 03-227 T/227). Blood samples were drawn from patients, their parents and controls after obtaining informed consent (parental consent in newborns and children below age 7 years).
Fifty-eight S-HSCR case–parent trios, 172 S-HSCR cases and 153 unrelated controls were included in this study. Parents were clinically unaffected. The male/female ratio was 5:1.
Marker selection
A total of 1693 SNP markers (density of 1 SNP per 5 kb approximately) encompassing the 3p21 region (from 47 to 53 Mb) were downloaded from HapMap data release 2005-03_16a_phase I (Build 34) for the Chinese Han from Beijing (CHB). Eight-hundred and fourteen common SNPs with a minor allele frequency of ≥5% were chosen and input in an ‘in-house’-built clustering programme, CLUSTAG24 for selection of tag SNPs. The cluster-merging threshold was set at r2=0.8.
Assay design and genotyping
We employed a Sequenom platform (Sequenom MassARRAY system, Sequenom, San Diego CA, USA)25 for assay design and genotyping. SNP sites were amplified by PCR in multiplex format in 384-microtiter plates by a pair of specifically designed forward and reverse PCR primers. The length of the amplicons for SNP capture ranged from 60 to 120 base pairs (bp). Following genomic amplification of the target regions, PCR products were treated with shrimp alkaline phosphatase for 20 minutes at 37 °C to dephosphorylate any residual nucleotides and to prevent their future incorporation and interference with the primer extension assay. Extension primers, DNA polymerase, and a cocktail mixture of deoxynucleotides (dNTPs) and dideoxynucleotide triphosphates (ddNTPs) were added to each mix. These were then followed by cycles of homogeneous MassEXTEND™ reaction probed by the extension primers for each SNP. The MassARRAY™ typer software version 3.1 was then used to read out the extended mass and assign the genotype call.
Quality control
For each 384-well plate, 20 samples were duplicated and four wells were filled with H2O (blank) to crosscheck contamination and reliability of the system. A whole plate was considered failed if: (i) no SNPs had passed the call rate of >80% and/or (ii) if the success rate of duplicate check had been <99.5% and that of the blank <90% and/or (iii) the success rate of the blank check alone had been <75%.
SNPs were removed from the analysis when: (i) they were not called at least in 90% of the individuals; (ii) they were monomorphic; (iii) their genotype frequencies deviated from the Hardy–Weinberg equilibrium expectation (P<0.01) or (iv) a Mendelian error rate of >0.01 was detected.
Individuals with less than 90% successful genotype calls were removed from the study.
Data analysis
The publicly available software Whap,26 Haploview27 and FUGUE (http://www.sph.umich.edu/csg/abecasis/fugue/)28 were used for statistical analyses of single markers and haplotypes for both family-based and case–control association tests. These programmes implement a standard EM approach to estimate haplotype frequencies.
The disease prevalence was set at 0.028. Empirical and global P-values for each test were obtained by running 1000 permutation tests.
Results and discussion
SNP selection, genotyping and quality control
After having implemented CLUSTAG, 218 markers were selected as tags. Four tag SNPs did not pass the quality control or SpectroDESIGNER failed to provide a successful genotyping assay. Therefore, a total of 214 SNPs were initially genotyped in 58 trios. One-hundred and seventy-three assays passed the data evaluation and were considered as successful. Genotype calls were >90% for all individuals included in this study.
Family-based association study
Single marker and haplotype analysis
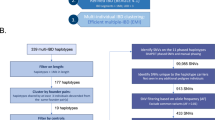
One-, three- and five-marker ‘sliding’ window TDT analyses showed association with HSCR within a narrow region (Figure 1). The optimum window-size subject to the minimum type I error was that of five markers. Five consecutive markers encompassing a 129 059 bp region were therefore identified. Although none of the markers were individually associated with HSCR (Table 1), haplotypes 6 and 7 (CCGGT and CCGGC, respectively) were associated with decreased risk to HSCR. Haplotype 1 (CCTAT) was more frequently transmitted to the affected offspring, although it did not reach statistical significance (Table 2).
Results of one (a), three (b) and five (c) marker ‘sliding’ window TDT analyses. At each position, the results of any covering window are averaged to produce a final statistic. This statistic is tested by permutations only (global significance values will also be given for the maximum and summed statistics).
Case–control association study
To verify the results obtained from the family-based analyses, 172 independent S-HSCR cases and 153 unrelated controls were also genotyped for the five markers comprised in the HSCR-associated haplotype observed previously. Single- and five-marker haplotype analyses were performed using Whap, Haploview and FUGUE. Similar results were obtained with all programmes and these are represented in Tables 3, 4. Marker rs747654 (marker 4 in Table 1) was found significantly associated with HSCR. This marker had the smallest P-value in the family-based association study conducted in 58 trios, even though, it did not reach statistical significance. Five-marker haplotype analyses showed that the CCTAT haplotype was significantly over-represented in patients (haplotype 1 in Table 2), hence conferring risk to HSCR. On the other hand, the CCGGT haplotype (haplotype 6 in Table 2) was significantly under-represented in the affected individuals, presumably conferring protection. Thus, the results from the family-based study could be replicated in an independent sample. Of note is the fact that ‘allele A’ of marker rs747654 was encompassed by the ‘risk’ haplotype while ‘allele G’ of the same marker by the ‘protective’ ‘haplotype’. Similar analyses were performed after having classified the individuals according to their gender. No significant differences were found when genders were compared, probably due to the small sample size and to the fact that the majority of patients were male. This was expected since no parental gender bias of transmission of susceptibility to HSCR at the 3p21 locus had previously been noted.22
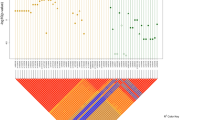
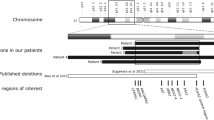
Since differences in LD between patient and control groups may help pinpoint the location of the causative locus in any given haplotype, we investigated the detailed LD in the region encompassed by the 3p21 five-marker haplotype in both patients and controls. Figure 2 represents the LD plot among these five markers in both HSCR patients and controls together with a representation of the genes encompassed in the region. Pair-wise LD values are higher in HSCR patients than in controls and consequently, LD blocks differ when using the ‘confidence interval’ method for block definition.29 According to HapMap data for CHB (Figure 3a), a spot for recombination exists between rs1841178 and rs3774808 (markers 1 and 2), which is in line with the overall lower LD values observed in our control population.
Biological implications
To investigate the putative biological implications of the HSCR-associated haplotype to the disease, we studied the genes encompassed by the five markers and at the intrinsic characteristics of the chromosomal region. Six genes and five transcripts were found in the 3p21 region, spanning from 48357638 to 48475671 (NCBI build 35). Detailed location of markers and genes in the region are depicted in Table 5 and Figure 2. Importantly, some of the genes encompassed are responsible for neurological phenotypes in humans (TREX1, PLXNB1 and SCOTIN). The most HSCR-associated SNP, rs747654, lies intronic to several transcripts of the prime repair exonuclease 1 gene (TREX1). Notably, TREX1 is expressed in developing gut of mouse.30 The plexin B1 gene (PLXNB1), encoding a receptor for the transmembrane semaphorin SEMA4D,31 promotes cell adhesion and neurite outgrowth and plays an important role in the development of the nervous system.32 Interestingly, SEMA4D maps to 9q31 region, which is known to harbor another RET-dependent modifier.33 SCOTIN plays a crucial role amid the p53–p73 pathway governing apoptosis and whose misregulation is implicated in the pathogenesis of neuroblastoma (neural crest cell-derived tumor).34, 35, 36 Apoptosis of neural crest cells have been suggested as one of the causes of enteric aganglionosis.37, 38, 39
It is, therefore, tempting to speculate that dysfunction of any of these genes could contribute to HSCR. The five SNPs are, in fact, located in conserved regulatory regions (as indicated by comparative analysis of genomic sequences by using VISTA, data not shown) and could, therefore, overlap binding sites of regulatory proteins governing the transcription of the genes within the region. To investigate this possibility, we used rVISTA.40 None of the SNP alleles introduced or abolished a predicted binding site, although a nucleotide change at these positions could well interfere with the neighboring sites by reducing its accessibility. Further analyses are needed to evaluate the effect of the SNPs on these genes. These SNPs could also act as surrogate markers for other functional site(s) and/or act synergistically with other disease variants.
Outside the five-marker haplotype region lies the ARHGEF3 gene (Rho guanine nucleotide exchange factor), which encodes a Rho GTPase that plays a fundamental role in numerous cellular processes. ARHGEF3 has been identified as an HSCR candidate locus in mice.30 Selected additional SNPs lying in the ARHGEF3 region were also genotyped in all HSCR and control samples. Unfortunately, no HSCR-associated SNP was identified.
The initial study reporting 3p21 as a RET-dependent modifier was conducted in samples from patients of European origin.22 Although that 3p21 region is a ‘gene rich’ region in relatively high LD, the later is higher in the individuals of northern and western European ancestry (CEU) genotyped for HapMap than in CHB. This provides grounds for differences in the associated haplotypes between populations (Figure 3), which could be advantageous for the identification of the causative locus. Of note, rs747654 is monomorphic in the CEU population (G allele; Table 5), which implies that if allele A of rs747654 was a causative variant, it would be exclusive to Chinese population. A more parsimonious explanation would be that the yet unknown susceptibility variant lies within the LD region.
In this study, we investigated the 3p21 region for genes that may be implicated in the pathogenesis of HSCR disease. Our data indicates that a HSCR locus could be lying in this region, even though the statistical significance did not survive rigorous multiple-testing adjustments. Indeed, the inclusion of a large number of SNPs generates an enormous multiple testing problem, especially when studying all but the strongest effects. However, our preliminary findings on trios could be replicated in an independent sample. Ranking markers by proximity to candidate genes or by expected functional consequences could be used in follow-up studies to finally pinpoint the HSCR locus.
References
Amiel J, Lyonnet S : Hirschsprung disease, associated syndromes, and genetics: a review. J Med Genet 2001; 38: 729–739.
Torfs C : An epidemiological study of Hirschsprung' disease in a multiracial California population. Third International Meeting: Hirschsprung's disease and related neurocristophaties Evian, France 1998 2004.
Edery P, Lyonnet S, Mulligan LM et al: Mutations of the RET proto-oncogene in Hirschsprung's disease. Nature 1994; 367: 378–380.
Puffenberger EG, Kauffman ER, Bolk S et al: Identity-by-descent and association mapping of a recessive gene for Hirschsprung disease on human chromosome 13q22. Hum Mol Genet 1994; 3: 1217–1225.
Angrist M, Bolk S, Halushka M et al: Germline mutations in glial cell line-derived neurotrophic factor (GDNF) and RET in a Hirschsprung disease patient. Nat Genet 1996; 14: 341–344.
Hofstra RM, Valdenaire O, Arch E et al: A loss-of-function mutation in the endothelin-converting enzyme 1 (ECE-1) associated with Hirschsprung disease, cardiac defects, and autonomic dysfunction. Am J Hum Genet 1999; 64: 304–308.
Pingault V, Bondurand N, Kuhlbrodt K et al: SOX10 mutations in patients with Waardenburg-Hirschsprung disease. Nat Genet 1998; 18: 171–173.
Carrasquillo MM, McCallion AS, Puffenberger EG et al: Genome-wide association study and mouse model identify interaction between RET and EDNRB pathways in Hirschsprung disease. Nat Genet 2002; 32: 237–244.
McCallion AS, Stames E, Conlon RA et al: Phenotype variation in two-locus mouse models of Hirschsprung disease: tissue-specific interaction between Ret and Ednrb. Proc Natl Acad Sci USA 2003; 100: 1826–1831.
Lang D, Epstein JA : Sox10 and Pax3 physically interact to mediate activation of a conserved c-RET enhancer. Hum Mol Genet 2003; 12: 937–945.
Lang D, Chen F, Milewski R et al: Pax3 is required for enteric ganglia formation and functions with Sox10 to modulate expression of c-ret. J Clin Invest 2000; 106: 963–971.
Borrego S, Ruiz A, Saez ME et al: RET genotypes comprising specific haplotypes of polymorphic variants predispose to isolated Hirschsprung disease. J Med Genet 2000; 37: 572–578.
Sancandi M, Griseri P, Pesce B et al: Single nucleotide polymorphic alleles in the 5′ region of the RET proto-oncogene define a risk haplotype in Hirschsprung's disease. J Med Genet 2003; 40: 714–718.
Burzynski GM, Nolte IM, Osinga J et al: Localizing a putative mutation as the major contributor to the development of sporadic Hirschsprung disease to the RET genomic sequence between the promoter region and exon 2. Eur J Hum Genet 2004; 12: 604–612.
Garcia-Barcelo MM, Sham MH, Lui VC et al: Chinese patients with sporadic Hirschsprung's disease are predominantly represented by a single RET haplotype. J Med Genet 2003; 40: e122.
Garcia-Barcelo M, Ganster RW, Lui VC et al: TTF-1 and RET promoter SNPs: regulation of RET transcription in Hirschsprung's disease. Hum Mol Genet 2005; 14: 191–204.
Burzynski GM, Nolte IM, Bronda A et al: Identifying candidate Hirschsprung disease-associated RET variants. Am J Hum Genet 2005; 76: 850–858.
Fitze G, Cramer J, Serra A et al: Within-gene interaction between c.135 G/A genotypes and RET proto-oncogene germline mutations in HSCR families. Eur J Pediatr Surg 2003; 13: 152–157.
Fitze G, Cramer J, Ziegler A et al: Association between c135G/A genotype and RET proto-oncogene germline mutations and phenotype of Hirschsprung's disease. Lancet 2002; 359: 1200–1205.
Fitze G, Appelt H, Konig IR et al: Functional haplotypes of the RET proto-oncogene promoter are associated with Hirschsprung disease (HSCR). Hum Mol Genet 2003; 12: 3207–3214.
Emison ES, McCallion AS, Kashuk CS et al: A common sex-dependent mutation in a RET enhancer underlies Hirschsprung disease risk. Nature 2005; 434: 857–863.
Gabriel SB, Salomon R, Pelet A et al: Segregation at three loci explains familial and population risk in Hirschsprung disease. Nat Genet 2002; 31: 89–93.
The International HapMap consortium: A haplotype map of the human genome. Nature 2005; 437: 1299–1320.
Ao SI, Yip K, Ng M et al: CLUSTAG: hierarchical clustering and graph methods for selecting tag SNPs. Bioinformatics 2005; 21: 1735–1736.
Jurinke C, van den BD, Cantor CR et al: Automated genotyping using the DNA MassArray technology. Methods Mol Biol 2001; 170: 103–116.
Purcell S, Daly MJ, Sham PC : WHAP: haplotype-based association analysis. Bioinformatics 2007; 23: 255–256.
Barrett JC, Fry B, Maller J et al: Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 2005; 21: 263–265.
Abecasis GR, Cherny SS, Cookson WO et al: Merlin—rapid analysis of dense genetic maps using sparse gene flow trees. Nat Genet 2002; 30: 97–101.
Gabriel SB, Schaffner SF, Nguyen H et al: The structure of haplotype blocks in the human genome. Science 2002; 296: 2225–2229.
Heanue TA, Pachnis V : From the Cover: expression profiling the developing mammalian enteric nervous system identifies marker and candidate Hirschsprung disease genes. Proc Natl Acad Sci USA 2006; 103: 6919–6924.
Artigiani S, Comoglio PM, Tamagnone L : Plexins, semaphorins, and scatter factor receptors: a common root for cell guidance signals? IUBMB Life 1999; 48: 477–482.
Worzfeld T, Puschel AW, Offermanns S et al: Plexin-B family members demonstrate non-redundant expression patterns in the developing mouse nervous system: an anatomical basis for morphogenetic effects of Sema4D during development. Eur J Neurosci 2004; 19: 2622–2632.
Bolk S, Pelet A, Hofstra RM et al: A human model for multigenic inheritance: phenotypic expression in Hirschsprung disease requires both the RET gene and a new 9q31 locus. Proc Natl Acad Sci USA 2000; 97: 268–273.
Bourdon JC, Renzing J, Robertson PL et al: Scotin, a novel p53-inducible proapoptotic protein located in the ER and the nuclear membrane. J Cell Biol 2002; 158: 235–246.
Terrinoni A, Ranalli M, Cadot B et al: p73-alpha is capable of inducing scotin and ER stress. Oncogene 2004; 23: 3721–3725.
Rossi M, Sayan AE, Terrinoni A et al: Mechanism of induction of apoptosis by p73 and its relevance to neuroblastoma biology. Ann N Y Acad Sci 2004; 1028: 143–149.
Lantieri F, Griseri P, Ceccherini I : Molecular mechanisms of RET-induced Hirschsprung pathogenesis. Ann Med 2006; 38: 11–19.
Bordeaux MC, Forcet C, Granger L et al: The RET proto-oncogene induces apoptosis: a novel mechanism for Hirschsprung disease. EMBO J 2000; 19: 4056–4063.
Kapur RP : Early death of neural crest cells is responsible for total enteric aganglionosis in Sox10(Dom)/Sox10(Dom) mouse embryos. Pediatr Dev Pathol 1999; 2: 559–569.
Dubchak I, Ryaboy DV : VISTA family of computational tools for comparative analysis of DNA sequences and whole genomes. Methods Mol Biol 2006; 338: 69–89.
Acknowledgements
We thank all subjects who participated in the study. This work was supported by research Grants HKU 765407M and HKU 775907M from the Hong Kong Research Grants Council to MGB and PT respectively. SSC and PCS are supported by NIH Grant EY-12562.
Author information
Authors and Affiliations
Corresponding authors
Additional information
SNP genotype data were downloaded from HapMap (http://www.hapmap.org/downloads/); SNP sequences and genomic characteristics were obtained from Ensembl public database (http://www.ensembl.org/Multi/martview); Gene and phenotype information were obtained from Map Viewer of NCBI (http://www.ncbi.nlm.nih.gov/mapview/); SNP sequence alignment with domain were searched from Pfam (http://www.sanger.ac.uk/Software/Pfam/).
Rights and permissions
About this article
Cite this article
Garcia-Barceló, MM., Fong, Py., Tang, C. et al. Mapping of a Hirschsprung's disease locus in 3p21. Eur J Hum Genet 16, 833–840 (2008). https://doi.org/10.1038/ejhg.2008.18
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2008.18
Keywords
This article is cited by
-
Exome-Wide Association Study Identified New Risk Loci for Hirschsprung’s Disease
Molecular Neurobiology (2017)
-
Four new loci associations discovered by pathway-based and network analyses of the genome-wide variability profile of Hirschsprung’s disease
Orphanet Journal of Rare Diseases (2012)
-
L1CAM malfunction in the nervous system and human carcinomas
Cellular and Molecular Life Sciences (2010)
-
Genetic basis of Hirschsprung’s disease
Pediatric Surgery International (2009)