Abstract
Tuberous sclerosis complex (TSC) is an autosomal dominant disorder characterised by the development of hamartomas in a variety of organs and tissues. The disease is caused by mutations in either the TSC1 gene on chromosome 9q34 or the TSC2 gene on chromosome 16p13.3. The TSC1 and TSC2 gene products, TSC1 and TSC2, interact to form a protein complex that inhibits signal transduction to the downstream effectors of the mammalian target of rapamycin (mTOR). Here we investigate the effects of putative TSC1 missense mutations identified in individuals with signs and/or symptoms of TSC on TSC1–TSC2 complex formation and mTOR signalling. We show that specific amino-acid substitutions close to the N-terminal of TSC1 reduce steady-state levels of TSC1, resulting in the activation of mTOR signalling and leading to the symptoms of TSC.
Similar content being viewed by others
Introduction
Tuberous sclerosis complex (TSC) is an autosomal dominant disorder characterised by the development of hamartomas in a variety of organs and tissues, including the brain, skin and kidneys.1, 2
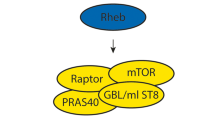
Mutations in either the TSC1 gene on chromosome 9q343 or the TSC2 gene on chromosome 16p13.34 cause TSC. The TSC1 and TSC2 gene products, TSC1 and TSC2, interact to form a protein complex. TSC2 contains a GTPase-activating protein domain and the TSC1–TSC2 complex acts on the rheb GTPase to prevent the rheb-GTP-dependent stimulation of cell growth through the mammalian target of rapamycin (mTOR).5 In cells lacking either TSC1 or TSC2, the downstream targets of mTOR, including p70 S6 kinase (S6K) and ribosomal protein S6, are constitutively phosphorylated.6, 7 The identification of the role of the TSC1–TSC2 complex in regulating mTOR activity has made it possible to compare TSC1 and TSC2 variants found in the normal population with variants identified in individuals with symptoms of TSC. The effects of amino-acid changes on TSC1–TSC2 complex formation, on the activation of rheb GTPase activity by the complex and on the phosphorylation status of S6K and S6 can be determined.8
Comprehensive screens for mutations at both the TSC1 and TSC2 loci have been performed in several large cohorts of TSC patients, and a wide variety of different pathogenic mutations have been described.9, 10, 11, 12, 13, 14, 15 Although ∼20% of the mutations identified in the TSC2 gene are missense changes, missense mutations in the TSC1 gene appear much less frequently. One simple explanation for this observation is that TSC1 missense mutations are rare because the majority of TSC patients have a mutation in the TSC2 gene. According to the TSC1 mutation database,16 22 putative missense mutations have been identified in TSC patients. However, only one of these is a confirmed de novo mutation. Here, we investigate the effects of 10 TSC1 missense changes (c.350T>C (p.L117P), c.539T>C (p.L180P), c.572T>A (p.L191H), c.671T>G (p.M224R), c.737G>A (p.R246K), c.913G>A (p.G305R), c.913G>T (p.G305W), c.1526G>A (p.R509Q), c.3103G>A (p.G1035S) and c.3290G>A (p.R1097H)) on TSC1–TSC2 function. We compared these TSC1 variants with wild-type TSC1 and three truncation variants: c.379_381delTGT (p.128delV), c.593_595delACT (p.N198F199delinsI) and c.2075C>T (p.R692X). Our analysis demonstrates that TSC1 missense mutations reduce steady-state levels of TSC1, resulting in increased mTOR activity and leading to the symptoms of TSC.
Materials and methods
Patient characteristics
Samples from patients with either a putative or definite diagnosis of TSC were received for mutation analysis. Details on clinical symptoms were obtained from the treating physicians who were sent a standardised clinical evaluation form (see Supplementary Table 1).
Mutation analysis
DNA was extracted from peripheral blood using standard techniques. Mutation analysis was performed as described earlier,13 or by direct sequence analysis of all TSC1 and TSC2 coding exons and exon/intron boundaries. In addition, both genes were analysed using the multiplex ligation-dependent probe amplification assay (MRC Holland, Amsterdam, The Netherlands).
To investigate whether the identified changes had an effect on splicing, three different splice-site prediction programs were used,17, 18, 19 as described earlier.13
Generation of constructs and antisera
Expression constructs encoding C-terminal YFP- and myc-tagged TSC1 variants were derived using the QuikChange site-directed mutagenesis kit (Stratagene, La Jolla, CA, USA). In each case, the complete open reading frame of the mutated construct was verified by sequence analysis. The other constructs used in this study have been described earlier.8, 20, 21 Polyclonal rabbit antisera specific for human TSC1 and TSC2 have been described earlier.21 Other antibodies were purchased from Cell Signaling Technology (Danvers, MA, USA).
Functional analysis of TSC1 variants
Expression of TSC1 variants in transfected cells
Human embryonal kidney (HEK) 293T cells seeded into 6-cm diameter dishes were transfected with a 1:1 mixture of the TSC1 and TSC2 expression constructs using Lipofectamine Plus (Invitrogen, Carlsbad, CA, USA), following the manufacturer's instructions. Two days after transfection, the cells were lysed in 50 mM Tris-HCl (pH 8.0), 100 mM NaCl, 50 mM NaF, 0.5 mM EDTA and 1% Triton X-100 plus protease inhibitors (Roche, Basel, Switzerland) and separated into supernatant and pellet fractions by centrifugation at 10 000 g for 10 min at 4°C as described earlier.22 Wild-type TSC1 and the TSC1 variants were immunoprecipitated from the supernatant fractions by incubation with a monoclonal antibody against the C-terminal myc epitope tag for 90 min at 4°C before incubation with Protein G beads (GE Healthcare, Uppsala, Sweden). After gentle agitation for 90 min at 4°C, the beads were washed three times with a >50-fold excess of lysis buffer. The immunoprecipitated proteins were detected by immunoblotting. Blots were developed using enhanced chemiluminescent detection (GE Healthcare).8
Immunoblot analysis of S6K T389 phosphorylation in cells overexpressing TSC1 variants
HEK 293T cells were transfected with a 4:2:1 mixture of the TSC1, TSC2 and S6K expression constructs. A total of 1.75 μg DNA was diluted in 200 μl Dulbecco's modified Eagle's medium (DMEM) containing 7 μg polyethyleneimine (Polysciences, Warrington, PA, USA). Where necessary, an empty expression vector (pcDNA3; Invitrogen) was added to make up the total amount of DNA. After 15 min at room temperature, the DNA/polyethyleneimine complexes were added to 80% confluent cells in 3.5-cm diameter dishes. After 4 h at 37°C, the transfection mixture was replaced with DMEM containing 10% foetal calf serum. Twenty-four hours after transfection, the cells were harvested and analysed by immunoblotting as before or by near infrared fluorescent detection on an Odyssey™ Infrared Imager (169 μm resolution, medium quality with 0 mm focus offset) (Li-Cor Biosciences, Lincoln, NE, USA). The integrated intensities of the protein bands were determined using the Odyssey software (default settings with background correction; 3-pixel width border average method). The mean ratios of the T389-phosphorylated S6K signal to the total S6K signal (T389/S6K) and the total TSC2 signal to the total TSC1 signal (TSC2/TSC1) were determined relative to wild-type TSC1 from at least three independent experiments (wild-type T389/S6K and TSC2/TSC1 ratios=1).
Immunofluorescent detection of S235/236 phosphorylation of ribosomal protein S6 in TSC1-deficient cells
Tsc1−/− mouse embryo fibroblasts (MEFs)7 were transfected with expression constructs encoding wild-type TSC1 or the TSC1 variants, using Lipofectamine Plus (Invitrogen), following the manufacturer's instructions. Twenty-four hours after transfection, S6 (S235/236) phosphorylation in the transfected cells was detected by immunofluorescent microscopy using a rabbit polyclonal antibody specific for S235/236-phosphorylated S6.23 TSC1 variants were identified either directly (for C-terminal-tagged TSC1-YFP variants) or with a mouse monoclonal antibody against the myc epitope tag (for C-terminal-tagged TSC1-myc variants). If possible, at least 50 cells expressing each TSC1 variant were counted per experiment and the number of cells showing a clear reduction in S6 (S235/236) phosphorylation was noted. The mean proportions of expressing cells with reduced S6 phosphorylation were calculated from at least three independent experiments.
Results
Patient characteristics and mutation analysis
The TSC1 c.350T>C (p.L117P) change was detected in two generations of a family with TSC (Figure 1a). The index patient (I:1) had epilepsy since the age of 22 years and fulfilled the diagnostic criteria for definite TSC, with facial angiofibroma, ungual fibroma, hypomelanotic macules, a shagreen patch and cerebral white matter migration lines. No mental disability was reported. The youngest child (II:2) had the c.350T>C change. Despite being somewhat hyperactive, this child developed normally until 4 years of age when epileptic seizures occurred and development stagnated. The child is now severely mentally retarded. No information was available about other signs of TSC. The eldest child (II:1), who could not yet be tested for the c.350T>C change, suffered severe anoxia at birth and has severe infantile encephalopathy with spastic tetraplegia and epilepsy.
Pedigrees of the investigated families with TSC. Arrows indicate the index cases. Open symbols indicate no signs or symptoms of TSC; black symbols indicate individuals with definite TSC; hatched symbols indicate individuals with possible TSC. A question mark indicates individuals where no clinical data were available. Genotypes are indicated for the individuals where DNA was available for testing. (a) Family with TSC and the TSC1 c.350T>C (p.L117P) variants. (b) Family with TSC and cosegregation of the TSC1 c.671T>G (p.M224R) and c.3103G>A (p.G1035S) variants. (c) Family with cosegregation of TSC and the TSC1 c.539T>C (p.L180P) variants. (d) Family with a de novo TSC1 c.572T>A (p.L191H) mutation.
The TSC1 c.671T>G (p.M224R) and TSC1 c.3103G>A (p.G1035S) changes were identified in two generations of a family with TSC24 (Figure 1b). The index case (II:3) had definite TSC with multiple shagreen patches, hypomelanotic macules, ungual fibromas, dental pits, epilepsy and severe mental disability. One parent (I:2) and both siblings (II:1 and II:2) of the index case also fulfilled the diagnostic criteria for definite TSC. Individual I:2, who was seizure-free and of below-average intelligence (IQ 73), had skin lesions pathognomonic for TSC. Individual II:1, who was of normal intelligence (IQ 94), had seizures, cortical tubers and multiple TSC skin lesions. Individual II:2 had epilepsy, below-average intelligence (IQ 67) and multiple TSC skin lesions. All the affected individuals in this family were heterozygous for the c.671T>G and c.3103G>A changes. Three affected individuals (I:2, II:1 and II:2) were heterozygous for a polymorphism in the TSC2 gene (TSC2 c.1276-32C>G). The index case was homozygous for the wild-type TSC2 1276-32C allele, consistent with TSC segregating with a mutation at the TSC1 locus in this family.
The TSC1 c.539T>C (p.L180P) change was detected in three generations of a family with TSC (Figure 1c). Individual I:2 had an ungual fibroma as the only reported sign of TSC. Individual II:2 had epilepsy, no mental disability, skin signs typical for TSC and a subependymal giant cell astrocytoma as well as other brain lesions consistent with a definite diagnosis of TSC. Individual II:3, the half-sibling of II:2, had typical TSC-associated skin lesions and possible mild mental retardation. Individual II:4, the other half-sibling of II:2, had a history of seizures during puberty and some skin tags, not typical of TSC. Individual III:1, the child of II:2, had epilepsy, and typical TSC-associated skin and brain lesions. Individual III:2, the sibling of III:1, had multiple cardiac rhabdomyoma and Wolff–Parkinson–White syndrome. No mental disability was reported for either individual III:1 or III:2. TSC did not cosegregate with markers mapping close to the TSC2 locus on chromosome 16p13.3, and no candidate TSC2 mutation was identified in the index case.
The TSC1 c.572T>A (p.L191H) change was identified in an individual who met the clinical criteria for definite TSC, including typical skin, heart and brain lesions. The individual had a history of seizures but no mental disability was reported. The c.572T>A change was absent in the individual's parents and child (Figure 1d). None of these individuals showed any signs of TSC.
Two TSC patients, one Finnish and one Dutch, were identified with the TSC1 c.737G>A (p.R246K) change. No information was available on the parents of either of these two individuals. The Finnish individual had typical TSC-associated skin, brain and kidney lesions. No mental disability or history of seizures was reported. The Dutch individual was also reported to have definite TSC. The sibling of this individual did not have any signs of TSC and did not carry the c.737G>A change.
The TSC1 c.1526G>A (p.R509Q) change was identified in a child of African origin, suspected of having TSC due to an echodensity that was detected prenatally in the septum of the heart. After birth, physical examination of the child did not reveal any signs of TSC but multiple congenital malformations that fitted with VACTERL (vertebral anomalies, atresia, cardiac malformations, tracheoesophageal fistula, renal anomalies and limb anomalies) association were identified. The same c.1526G>A change was identified in one of the parents. Neither parent reported any signs of TSC.
The TSC1 c.3290G>A (p.R1097H) change was identified in a child with cardiac rhabdomyoma who subsequently developed epilepsy at the age of 3 months. An MRI scan of the brain showed multiple subependymal nodules, cortical tubers and white matter abnormalities. This individual fulfilled the diagnostic criteria for definite TSC, with angiomyolipoma and multiple skin lesions. The c.3290G>A change was also identified in one of the parents; neither parent showed any signs of TSC.
The TSC1 c.379_381delTGT (p.128delV) change was detected in a child with epilepsy and a definite diagnosis of TSC (multiple skin signs, subependymal nodules and angiomyolipoma). No mental disability was reported and no information was available on the phenotypic or genetic status of the parents. The same change has been reported in another unrelated TSC patient.25 The TSC1 c.593_595delACT (p.N198F199delinsI), c.913G>A (p.G305R) and c.913G>T (p.G305W) missense changes have been reported earlier to cosegregate with TSC in three independent families.15, 26
In all the above cases, no other putative pathogenic mutations were identified and comparison of the allele ratios of the index cases and parents (where possible) did not reveal any evidence for somatic mosaicism in the leukocyte DNA.
Comparative analysis of TSC1 amino-acid substitutions
During our initial mutation screening, the L117P change was the only putative pathogenic TSC1 amino-acid substitution identified. We compared the L117P variant with wild-type TSC1, an earlier characterised TSC1 in-frame deletion (N198F199delinsI)27 and to a common TSC1 truncation mutation, R692X3, 10 (Figure 2). Wild-type TSC1 was detected predominantly in the post 10 000 g pellet fraction, whereas the three variants were detected in the supernatant fraction (Figure 2a). Upon coexpression of TSC2, the variants as well as wild-type TSC1 were detected in both subcellular fractions (Figure 2b). TSC2 was coimmunoprecipitated from the supernatant fraction with all three variants, although clearly less effectively than with wild-type TSC1.
Analysis of the TSC1 L117P variant. Wild-type TSC1 (wt) or the L117P, N198F199delinsI (del) or R692X variants (all containing a C-terminal myc epitope tag) were overexpressed in HEK 293T cells alone (a) or in the presence of coexpressed TSC2 (b). Cell lysates were separated into post 10 000 g centrifugation pellet and supernatant fractions. The TSC1 variants were detected with a monoclonal mouse antibody against the myc epitope tag (9B11; Cell Signaling Technology). TSC1–TSC2 complexes were immunoprecipitated from the supernatant fractions using the same antibody. TSC2 was detected with a polyclonal rabbit antiserum.8 (a) Expression of the TSC1 variants in the absence of TSC2. Wild-type TSC1 (wt) was detected predominantly in the pellet fraction. In contrast, the variants were detected predominantly in the supernatant fraction. The signals for the L117P and N198F199delinsI (del) variants are clearly less than wild-type TSC1 and the R692X variants (as detected with the 9B11 antibody). (b) Coexpression of TSC2 and the TSC1 variants. In the presence of TSC2, wild-type TSC1 and the variants were detected in both subcellular fractions. Coexpression of TSC2 resulted in a shift of wild-type TSC1 to the supernatant fraction and a shift of the variants to the pellet fraction. TSC1 and the TSC1 variants were immunoprecipitated with an antibody against the myc epitope tag. TSC2 was coimmunoprecipitated with all three variants. However, in each case, the TSC2 signal in the immunoprecipitate was clearly less than with the wild-type TSC1.
Next, we investigated whether the variants could inhibit mTOR signalling. Expression of wild-type TSC1 alone is insufficient to inhibit mTOR activity.28 Therefore, the variants were coexpressed with TSC2 and S6K in HEK 293T cells, and S6K T389 phosphorylation was analysed by immunoblotting (Figure 3a). To try and achieve comparable expression levels of the different variants, we used different quantities of the corresponding expression constructs. Coexpression of wild-type TSC1 and TSC2 resulted in a reduction in S6K T389 phosphorylation, even with low levels of the TSC1 expression construct. In contrast, coexpression of TSC2 with the variants did not reduce S6K T389 phosphorylation, indicating that they were unable to inhibit mTOR effectively. Compared with wild-type TSC1, the L117P and N198F199delinsI variants were detected at low levels, even when the quantity of transfected expression construct DNA was increased.
TSC1–TSC2-dependent inhibition of mTOR signalling by the TSC1 L117P variant. (a) T389 phosphorylation of S6K in HEK 293T cells coexpressing TSC2, myc-tagged S6K and myc-tagged wild-type TSC1 (wt) or the TSC1 L117P, N198F199delinsI (del) or R692X variants was determined by immunoblotting. Cells transfected with differing amounts of wild-type or variant expression constructs, as indicated, were analysed. The first lane on the left (control) corresponds to cells transfected with S6K and wild-type TSC1 expression constructs only (no TSC2 expression construct). The coexpression of only TSC2 and wild-type TSC1 inhibited S6K phosphorylation. (b) TSC1-dependent inhibition of S6 phosphorylation. YFP-tagged wild-type TSC1 (YFPTSC1) and the YFP-tagged N198F199delinsI (YFPdel) and L117P (YFPL117P) variants were expressed in Tsc1−/− MEFs. S6 S235/236 phosphorylation in the YFP-positive cells was detected by immunofluorescent microscopy using an antibody specific for S235/236-phosphorylated S6 (Cell Signaling Technology). As a control, S6 phosphorylation in cells expressing YFP only was also determined. At least 50 cells were counted per variant per experiment. Mean percentage scores of three separate experiments are shown. Values significantly different from the wild type are indicated.
To confirm that the L117P and N198F199delinsI variants were unable to inhibit mTOR, they were expressed in Tsc1−/− MEFs. These cells exhibit constitutive S6 (S235/236) phosphorylation.7 As shown in Figure 3b, wild-type YFP-tagged TSC1 reduced S6 phosphorylation in >60% of the YFP-positive Tsc1−/− MEFs, consistent with earlier results.8 In contrast, S6 phosphorylation was reduced in <30% of cells expressing the L117P or N198F199delinsI variants (paired t-test P<0.001). Nevertheless, a reduction in S6 phosphorylation was observed in a higher proportion of variant-expressing cells than in cells expressing YFP only (paired t-test P<0.02). Therefore, although both variants were clearly less effective than wild-type TSC, we could not exclude the possibility that the variants can antagonise mTOR activity.
As a result of on-going TSC1 and TSC2 mutation screening, we identified eight additional TSC1 single amino-acid changes: p.128delV, p.L180P, p.L191H, p.M224R, p.R246K, p.R509Q, p.G1035S and p.R1097H. The p.M224R and p.G1035S changes cosegregated with TSC in a single family (Figure 1b). To investigate the effects of these two amino-acid substitutions on TSC1 function, the corresponding single and double variants were compared with wild-type TSC1 and the R692X truncation. As shown in Figure 4a, the M224R variant and the M224R/G1035S double variant were detected at lower levels than wild-type TSC1 and the G1035S variant. Furthermore, the expression of wild-type TSC1 or the G1035S variant reduced S6 phosphorylation in >50% of transfected Tsc1−/− MEFs whereas <20% of the MEFs expressing the M224R, M224R/G1035S or R692X variants showed a reduction in S6 phosphorylation (Figure 4b), indicating that the M224R substitution is the pathogenic mutation in this family and that the G1035S substitution is a cosegregating neutral variant.24 We did not observe any differences between the M224R single mutant and the M224R/G1035S double mutant.
Results of the functional assays on the M224R, G1035S and M224R/G1035S variants. (a) TSC1–TSC2-dependent inhibition of S6K-T389 phosphorylation. TSC2, S6K and wild-type TSC1 (wt TSC1), or the TSC1 variants, were expressed in HEK 293T cells. T389 phosphorylation of S6K was determined by immunoblotting and was reduced in the presence of wild-type TSC1 and the G1035S variants only. S6K, wild-type TSC1 and the TSC1 variants were detected with an antibody against the myc epitope tag. The signal for the M224R and M224R/G1035S variants is reduced compared with wild-type TSC1 and the G1035S and R692X variants. A representative example of at least three separate experiments is shown. (b) Inhibition of S6 phosphorylation in Tsc1−/− MEFs. Wild-type myc-tagged TSC1 (TSC1) and the myc-tagged variants were expressed in Tsc1−/− MEFs. S6 S235/236 phosphorylation in the myc-positive cells was determined by double-label immunofluorescent microscopy using a rabbit polyclonal antibody specific for S235/236-phosphorylated S6 and a mouse monoclonal antibody specific for the myc tag. The percentage of Tsc1−/− MEFs expressing the TSC1 variants and showing a clear reduction in S6 phosphorylation is indicated. Because the signals for the M224R and M224R/G1035S variants were very low, it was difficult to unequivocally differentiate >50 expressing cells. Therefore, the total number of counted cells after two experiments is indicated above each bar.
To determine whether the 128delV, L180P, L191H, R246K, R509Q and R1097H amino-acid changes were pathogenic, we compared these variants with the previously characterised L117P and R692X variants, and with two additional putative missense variants, G305R and G305W15 (Figure 5). The R246K, G305R, G305W, R509Q and R1097H variants were detected at comparable levels with wild-type TSC1 (Figure 5a and b) and were just as effective at inhibiting S6K T389 phosphorylation (Figure 5a and c). In contrast, the 128delV, L180P and L191H variants were detected at low levels, similar to the L117P and M224R variants (Figure 5a and b). Furthermore, S6K T389 phosphorylation was not inhibited by the expression of these variants (Figure 5a and c). Consistent with the immunoblot data, the expression of the L117P, 128delV, L180P, L191H, M224R, M224R/G1035S and R692X variants in Tsc1−/− MEFs reduced S6 phosphorylation in <40% of cells, whereas wild-type TSC1 and the R246K, G305R, G305W, R509Q, G1035S and R1097H variants reduced S6 phosphorylation in >50% of the expressing cells (Figure 5d).
Analysis of the 128delV, L180P, L191H, R246K, G305R, G305W, R509Q and R1097H variants. (a) TSC1-dependent inhibition of S6K T389 phosphorylation. S6K T389 phosphorylation was determined by immunoblotting as before except that the blots were developed using Odyssey near infrared detection (Li-Cor Biosciences). Cells expressing S6K, TSC2 and wild-type TSC1 (wt) or the TSC1 variants were analysed. As controls, cells transfected with expression constructs for wild-type TSC1 and S6K only (TSC1/S6K), TSC2 and S6K only (TSC2/S6K), S6K only (S6K) or empty vector only (pcDNA3) were also analysed. S6K and the TSC1 variants were detected with an antibody specific for the myc epitope tag. The R692X truncation variant is indicated with a large arrowhead; molecular weight markers are indicated with small arrows. Expression of wild-type TSC1 or the R246K, G305R, G305W, R509Q, G1035S and R1097H variants clearly inhibited S6K phosphorylation. The signals for the L117P, 128delV, L180P, L191H and M224R variants were clearly reduced compared with wild-type TSC1 (see also panel b), and S6K T389 phosphorylation was increased (see also panel c). A representative example of three separate experiments is shown. (b) Relative expression of the TSC1 variants. The integrated intensities of the TSC1 and TSC2 signals for each variant were determined in three independent experiments using the Odyssey scanning software. The mean TSC2/TSC1 ratios, relative to the wild-type TSC1 (wt) (wild-type TSC1 TSC2/TSC1 ratio=1), were determined. Standard deviations are indicated. (c) TSC1-dependent inhibition of S6K T389 phosphorylation. The ratio of the T389 S6K phosphorylation signal intensity to the total S6K signal intensity (T389/S6K) was measured in three independent experiments. The integrated intensity of each band was determined using the Odyssey scanning software and the mean T389/S6K ratios, relative to the wild-type TSC1 (wt) (wild-type TSC1 T389/S6K ratio=1), were determined. Standard deviations are indicated. (d) Inhibition of S6 phosphorylation in Tsc1−/− MEFs. Wild-type myc-tagged TSC1 (wt) or the myc-tagged variants were expressed in Tsc1−/− MEFs. S6 S235/236 phosphorylation in the myc-positive cells was determined by immunofluorescent microscopy as before. At least 50 cells were counted per variant per experiment. Each variant was tested in at least two separate experiments. Mean percentages of Tsc1−/− MEFs expressing the TSC1 variants and showing reduced phosphorylation of S6 are shown. Standard deviations are indicated.
We concluded that the 128delV, L180P and L191H changes destabilise TSC1, resulting in increased mTOR activity. The R246K, G305R, G305W, R509Q, G1035S and R1097H amino-acid substitutions did not affect TSC1 function in our assays.
Splice-site prediction analysis
Splice-site prediction analysis of the identified variants was performed according to a standard protocol13 using three independent splice-site prediction programs.17, 18, 19 Effects were predicted for four of the variants: TSC1 c.379_381delTGT (p.128delV), c.737G>A (p.R246K) c.913G>A (p.G305R) and c.913G>T (p.G305W).
According to two of the prediction programs, the c.379_381delTGT mutation created a potential new acceptor site within exon 6. However, the values of the new site (0.48 and 0.33 in Fruitfly and NetGene2, respectively) were lower than the values of the wild-type acceptor site (0.98 and 0.97, respectively). It is therefore unlikely that this sequence change affects splicing.
The R246K, G305R and G305W amino-acid substitutions did not affect TSC1 function. However, the c.737G>A (p.R246K) nucleotide substitution altered the last nucleotide of exon 8 and all three programs indicated that this change disrupted the intron 8 splice donor site, leaving a new potential donor site 90 nucleotides downstream. Similarly, the c.913G>A (p.G305R) and c.913G>T (p.G305W) substitutions affect the last nucleotide of exon 9 and were predicted by all three programs to disrupt splicing to the intron 9 donor site. We concluded that these three variants are most likely pathogenic splice-site mutations, not missense mutations. Unfortunately, the effect of these sequence changes on TSC1 RNA splicing could not be confirmed experimentally because no new samples could be obtained.
An overview of the functional assays and splice-site analysis is presented in Supplementary Table 2.
Discussion
Mutation analysis of the TSC1 and TSC2 genes in individuals with TSC, and in those suspected of having the disease is important for diagnosis and genetic counselling. During our screening of a cohort of approximately 900 index cases with TSC, or possible TSC, we identified eight putative pathogenic TSC1 missense changes, c.350T>C (p.L117P), c.539T>C (p.L180P), c.572T>A (p.L191H), c.671T>G (p.M224R), c.737G>A (p.R246K), c.1526G>A (p.R509Q), c.3103G>A (p.G1035S) and c.3290G>A (p.R1097H), as well as a putative pathogenic in-frame deletion mutant, TSC1 c.379_381delTGT (p.128delV). We compared these variants with the earlier reported c.913G>A (p.G305R), c.913G>T (p.G305W), c.593_595delACT (p.N198F199delinsI) and c.2074C>T (p.R692X) mutants.2, 15, 26
The TSC1 L117P, 128delV, L180P, L191H, N198F199delinsI and M224R changes resulted in reduced levels of TSC1 and a reduction in TSC1-dependent inhibition of mTOR activity, as detected by immunoblotting. In each case, the functional characterisation was consistent with the genetic and phenotypic findings and we concluded that the changes were pathogenic.
The R246K, G305R, G305W, R509Q, G1035S and R1097H amino-acid substitutions did not affect TSC1 function in our assays. However, we concluded that the TSC1 c.737G>A (p.R246K), c.913G>A (p.G305R) and c.913G>T (p.G305W) changes were most likely pathogenic splice-site mutations. The c.737G>A transition was predicted to destroy the splice donor site at the 3′ end of exon 8 and the c.913G>A and c.913G>T substitutions affect the last nucleotide of exon 9, also disrupting the normal splice donor site. The c.1526G>A (p.R509Q), c.3103G>A (p.G1035S) and c.3290G>A (p.R1097H) variants did not appear to affect either TSC1 RNA splicing or TSC1 function. The c.1526G>A change was identified in a foetus suspected of having TSC. However, after birth, the child showed multiple congenital malformations that fitted with VACTERL association, and a normal physical examination of the child did not reveal any signs of TSC. We concluded that this variant is unlikely to cause TSC. The c.3103G>A variant cosegregated with the pathogenic c.671T>G (p.M224R) variant and TSC in a single family.24 We concluded that the p.G1035S substitution was a neutral variant, consistent with previous reports.29, 30 The c.3290G>A change was identified in a child with definite TSC and in one unaffected parent, and we concluded that it was also most likely a rare neutral variant.
Here we demonstrate that specific amino-acid substitutions close to the N-terminal of TSC1 (amino acids 117–224) reduce the steady-state levels of TSC1. The location and the effects of these changes on TSC1 function are very similar to a small number of missense mutations described recently in some cases of bladder cancer,31 suggesting that the pathogenetic mechanisms underlying TSC-associated lesions and tumours of the bladder may be related.
Several studies indicate that TSC1 mutations are associated with a less severe clinical presentation in TSC patients.9, 12, 13, 15 The small number of patients that we identified with a TSC1 missense mutation made it difficult to identify a specific phenotypic spectrum in this group. Nevertheless, the use of functional assays to differentiate between polymorphisms and pathogenic mutations, in TSC and other diseases, will not only facilitate the identification of pathogenic mutations but also help investigate possible genotype–phenotype correlations and provide insight into how specific amino-acid residues contribute to protein function.
References
Gomez M, Sampson J, Whittemore V eds: The Tuberous Sclerosis Complex. Oxford, UK: Oxford University Press, 1999.
Roach ES, DiMario FJ, Kandt RS et al: Tuberous sclerosis consensus conference: recommendations for diagnostic evaluation. National Tuberous Sclerosis Association. J Child Neurol 1999; 14: 401–407.
van Slegtenhorst M, de Hoogt R, Hermans C et al: Identification of the tuberous sclerosis gene TSC1 on chromosome 9q34. Science 1997; 277: 805–808.
The European Chromosome 16 Tuberous Sclerosis Consortium: Identification and characterization of the tuberous sclerosis gene on chromosome 16. Cell 1993; 75: 1305–1315.
Li Y, Corradetti MN, Inoki K et al: TSC2: filling the GAP in the mTOR signaling pathway. Trends Biochem Sci 2003; 28: 573–576.
Zhang H, Cicchetti G, Onda H et al: Loss of Tsc1/Tsc2 activates mTOR and disrupts PI3K-Akt signaling through downregulation of PDGFR. J Clin Invest 2003; 112: 1223–1233.
Kwiatkowski DJ, Zhang H, Bandura JL et al: A mouse model of TSC1 reveals sex-dependent lethality from liver hemangiomas, and up-regulation of p70S6 kinase activity in Tsc1 null cells. Hum Mol Genet 2001; 11: 525–534.
Nellist M, Sancak O, Goedbloed MA et al: Distinct effects of single amino acid changes to tuberin on the function of the tuberin–hamartin complex. Eur J Hum Genet 2005; 13: 59–68.
Jones AC, Shyamsundar MM, Thomas MW et al: Comprehensive mutation analysis of TSC1 and TSC2, and phenotypic correlations in 150 families with tuberous sclerosis. Am J Hum Genet 1999; 64: 1305–1315.
van Slegtenhorst M, Verhoef S, Tempelaars A et al: Mutational spectrum of the TSC1 gene in a cohort of 225 tuberous sclerosis complex patients: no evidence for genotype–phenotype correlation. J Med Genet 1999; 36: 285–289.
Au KS, Rodriguez JA, Finch JL et al: Germ-line mutational analysis of the TSC2 gene in 90 tuberous sclerosis patients. Am J Hum Genet 1998; 62: 286–294.
Dabora SL, Jozwiak S, Franz DN et al: Mutational analysis in a cohort of 224 tuberous sclerosis patients indicates increased severity of TSC2, compared with TSC1, disease in multiple organs. Am J Hum Genet 2001; 68: 64–80.
Sancak O, Nellist M, Goedbloed M et al: Mutational analysis of the TSC1 and TSC2 genes in a diagnostic setting: genotype–phenotype correlations and comparison of diagnostic DNA techniques in tuberous sclerosis complex. Eur J Hum Genet 2005; 13: 731–741.
Niida Y, Lawrence-Smith N, Banwell A et al: Analysis of both TSC1 and TSC2 for germline mutations in 126 unrelated patients with tuberous sclerosis. Hum Mutat 1999; 14: 412–422.
Au K-S, Williams AT, Roach ES et al: Genotype/phenotype correlation in 325 individuals referred for a diagnosis of tuberous sclerosis complex in the United States. Genet Med 2007; 9: 88–100.
Tuberous sclerosis database – Leiden Open Variation Database. [www.chromium.liacs.nl/lovd/index.php?select_db=TSC2].
NetGene2 Server. [www.cbs.dtu.dk/services/NetGene2].
SpliceSiteFinder. [www.genet.sickkids.on.ca/∼ali/splicesitefinder.html].
BDGP: Splice Site Prediction by Neural Network. [www.fruitfly.org/seq_tools/splice.html].
Nellist M, van Slegtenhorst MA, Goedbloed M et al: Characterization of the cytosolic tuberin–hamartin complex: tuberin is a cytosolic chaperone for hamartin. J Biol Chem 1999; 274: 35647–35652.
van Slegtenhorst M, Nellist M, Nagelkerken B et al: Interaction between hamartin and tuberin, the TSC1 and TSC2 gene products. Hum Mol Genet 1998; 7: 1053–1057.
Nellist M, Verhaaf B, Goedbloed MA et al: TSC2 missense mutations inhibit tuberin phosphorylation and prevent formation of the tuberin-hamartin complex. Hum Mol Genet 2001; 10: 2889–2898.
Jaeschke A, Hartkamp J, Saitoh M et al: Tuberous sclerosis complex tumor suppressor-mediated S6 kinase inhibition by phosphatidylinositide-3-OH kinase is mTOR independent. J Cell Biol 2002; 159: 217–224.
Jansen FE, Braams O, Vincken KL et al: Overlapping neurologic and cognitive phenotypes in patients with TSC1 or TSC2 mutations. Neurology 2008; 70: 908–915.
Lee-Jones L, Aligianis I, Davies PA et al: Sacrococcygeal chordomas in patients with tuberous sclerosis complex show somatic loss of TSC1 or TSC2. Genes Chromosomes Cancer 2004; 41: 80–85.
Goedbloed MA, Nellist M, Verhaaf B et al: Analysis of TSC2 stop codon variants found in tuberous sclerosis patients. Eur J Hum Genet 2001; 9: 823–828.
Hodges AK, Li S, Maynard J et al: Pathological mutations in TSC1 and TSC2 disrupt the interaction between hamartin and tuberin. Hum Mol Genet 2001; 10: 2899–2905.
Inoki K, Li Y, Zu T et al: TSC2 is phosphorylated and inhibited by Akt and suppresses mTOR signalling. Nat Cell Biol 2002; 4: 648–657.
Young JM, Burley MW, Jeremiah SJ et al: A mutation screen of the TSC1 gene reveals 26 protein truncating and 1 splice site mutation in a panel of 79 tuberous sclerosis patients. Ann Hum Genet 1998; 62: 203–213.
Dabora SL, Sigalas I, Hall F et al: Comprehensive mutation analysis of TSC1 using two-dimensional electrophoresis with DGGE. Ann Hum Genet 1998; 62: 491–504.
Pymar LS, Platt FM, Askham JM et al: Bladder tumour-derived somatic TSC1 missense mutations cause loss of function via distinct mechanisms. Hum Mol Genet 2008; 17: 2006–2017.
Acknowledgements
Financial support was provided by the US Department of Defense Congressionally-Directed Medical Research Program (grant no. TS060052) and the Michelle Foundation. We thank the family members who contributed to this study. Dr N Migone is thanked for helpful comments on the paper. The authors report no conflicts of interest.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website (http://www.nature.com/ejhg)
Supplementary information
Rights and permissions
About this article
Cite this article
Nellist, M., van den Heuvel, D., Schluep, D. et al. Missense mutations to the TSC1 gene cause tuberous sclerosis complex. Eur J Hum Genet 17, 319–328 (2009). https://doi.org/10.1038/ejhg.2008.170
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2008.170
Keywords
This article is cited by
-
Mutational analysis of TSC1 and TSC2 genes in Tuberous Sclerosis Complex patients from Greece
Scientific Reports (2017)
-
The Role of mTOR Inhibitors in the Treatment of Patients with Tuberous Sclerosis Complex: Evidence-based and Expert Opinions
Drugs (2016)
-
Genotype and cognitive phenotype of patients with tuberous sclerosis complex
European Journal of Human Genetics (2012)
-
Determining the pathogenicity of patient-derived TSC2 mutations by functional characterization and clinical evidence
European Journal of Human Genetics (2011)
-
Characterisation of TSC1 promoter deletions in tuberous sclerosis complex patients
European Journal of Human Genetics (2011)