Abstract
H2A is a nucleosome core subunit involved in organizing DNA into a chromatin structure that is often inaccessible to regulatory enzymes. Replacement of H2A by its variant H2A.Z renders chromatin accessible at enhancers and promoters. However, it remains unclear how H2A.Z functions so differently from canonical H2A. Here we report the genome-wide identification of proteins that directly interact with H2A and H2A.Z in vivo using a novel strategy, bPPI-seq. We show that bPPI-seq is a sensitive and robust technique to identify protein-protein interactions in vivo. Our data indicate that H2A.Z-interacting proteins and H2A-interacting proteins participate in distinct biological processes. In contrast to H2A-interacting proteins, the H2A.Z-interacting proteins are involved in transcriptional regulation. We found that the transcription factor Osr1 interacts with H2A.Z both in vitro and in vivo. It also mediates H2A.Z incorporation to a large number of target sites and regulates gene expression. Our data indicate that bPPI-seq can be widely applied to identify genome-wide interacting proteins under physiological conditions.
Similar content being viewed by others
Introduction
In eukaryotic nuclei, nucleosomes, the building block of chromatin, are formed by wrapping 146 bp of DNA around a histone octamer consisting two copies each of H2A, H2B, H3 and H4. While the vast majority of nucleosomes are composed of canonical core histones, a minor fraction of them are associated with histone variants. H2A.Z-containing nucleosomes are involved in multiple biological processes1,2,3. In contrast to H2A nucleosomes, H2A.Z-containing nucleosomes are specifically localized to transcriptional regulatory regions such as promoters and enhancers4,5,6,7,8,9. While H2A nucleosomes are generally involved in packaging DNA and condensation of chromatin, incorporation of H2A.Z facilitates targeting of transcription factors and chromatin regulators by maintaining chromatin accessibility8,9,10,11,12. Although crystallographic and biochemical studies revealed structural differences between H2A and H2A.Z nucleosomes13, the underlying basis of the drastic differences in their function and the mechanisms that target H2A.Z to specific chromatin regions remains unclear.
Several studies have reported identification of proteins that co-purify with H2A.Z nucleosomes using affinity purification followed by mass spectrometry analysis (AP-MS). A number of proteins are found to co-purify with H2A.Z14,15,16. In particular, ANP32E was identified as an H2A.Z-specific chaperone that mediates deposition of H2A.Z on chromatin16,17. INO80 was identified as an H2A.Z-specific remodeler to remove H2A.Z for promoting homologous recombination in human cells18. While these studies identified some interesting H2A.Z-interacting proteins that may be involved in the downstream function of H2A.Z, it remains unclear how H2A.Z finds its specific target sites on chromatin. Thus more comprehensive knowledge of interacting proteins is needed to understand the functional differences and targeting mechanisms of H2A and H2A.Z. However, current AP-MS strategies suffer from several limitations: (1) weak interactions may not be detected; (2) interactions that require a physiological chromatin environment may not be detected; (3) some proteins detected as interacting with H2A.Z may not interact directly but instead their interaction may be mediated by other components of the H2A.Z-containing nucleosomes; (4) some detected interactions may occur only in vitro but not in vivo; and (5) the proteins pulled down by an antibody vary depending on the binding and washing conditions, which may result in very low reproducibility of data from different experiments in the same lab or between labs even if the same antibody is being used.
Because a comprehensive catalog of H2A- and H2A.Z-interacting proteins is needed to understand their functional differences, we developed a sensitive in vivo strategy, termed bPPI-seq (bait Protein-Protein Interaction-sequencing), to detect genome-wide interacting proteins and used this strategy to compare the H2A- and H2A.Z-interacting proteins genome-wide. Our results indicated that H2A.Z-interacting proteins are enriched for proteins that regulate transcription while such proteins do not constitute the H2A-interacting proteins, thus providing novel insights into the mechanisms underlying their functional difference. We found that H2A.Z interacts with 20 DNA-binding transcription factors and further demonstrate that Osr1, a zinc-finger transcription factor, facilitates targeting of H2A.Z to regions containing Osr1 motifs, providing evidence that sequence-specific transcription factors mediate targeting of H2A.Z to specific chromatin regions. Our results indicate that bPPI-seq is an effective strategy to identify genome-wide protein interactions in mammalian cells.
Results
Reconstituting active GFP by fusion of its N-terminus with H2A.Z and its C-terminus with ANP32E
An active GFP can be reconstituted from two non-fluorescent fragments of GFP brought together by two interacting proteins19. Using this strategy, α-actinin 4 (ACTN4) was identified as a substrate of the serine/threonine protein kinase AKT1, which mediates its translocation to cell membrane20. To test if an active GFP can be reconstituted from two fragments fused to H2A.Z and an H2A.Z-interacting protein, we co-expressed the fusion protein (GFPN-H2A.Z) between GFP N-terminus and H2A.Z, and the fusion protein (ANP32E-GFPC) between GFP C-terminus and ANP32E, a known H2A.Z-interacting protein16,17 in NIH 3T3 cells. The cells turned green after 48 h of expression (Figure 1A, left panel), while very few green cells were detected when GFPN-H2A.Z was co-expressed with a non-interacting protein Tinf2-GFPC (Figure 1A, right panel). These results indicate that H2A- and H2A.Z-interacting proteins can be detected by the GFP reconstitution assay in vivo.
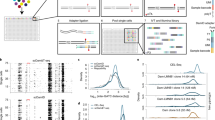
Genome-wide detection of H2A- and H2A.Z-interacting proteins using bPPI-seq. (A) Co-expression of GFPN-H2A.Z with ANP32E-GFPC reconstitutes an active green fluorescent protein. The constructs expressing full-length fusion proteins in pEF1a vector were co-transfected into NIH 3T3 cells and examined under microscope 48 h post transfection. Tinf2-GFPC is used as a negative control. (B) Isolation of GFP+ cells by flow cytometry sorting. The stable cell lines expressing GFPN-H2A.Z, GFPN-H2A or GFPN alone were transduced with three ReMTH-GFPC1, 2 and 3 viral particles to tag endogenous genes with GFPC domain, selected by puromycin for 5 days, followed by Doxycycline induction for 48 h before sorting. (C) Flow chart for preparation of the H2A.Z-interacting protein libraries (bPPI-seq) from RNAs isolated from the sorted cells. (D) Genome Browser image showing the bPPI-seq reads in the gene encoding H2A.Z-interacting protein ANP32E. The y-axis was normalized to library sequencing depth. The peak location indicates the fusion site of hybrid mRNA between C-terminal half (GFPC) and the mRNA of endogenous gene. The panels from top to the bottom are bPPI-seq GFPN (two replicates), bPPI-seq GFPN-H2A (two replicates), bPPI-seq GFPN-H2A.Z (two replicates) and RNA-seq.
Random gene tagging with ERM vectors
To unbiasedly identify H2A- and H2A.Z-interacting proteins, we randomly tagged endogenous genes by exon trapping using an enhanced retroviral mutagen vector21, which contains the cDNA of GFPC and a splicing donor (SD) driven by a Doxycycline-inducible promoter. After integration upstream or inside an endogenous gene, the splice donor mediates the formation of a hybrid transcript of GFPC fused in frame to the downstream endogenous exons, thus expressing a GFPC fusion protein.
Ectopically expressed GFPN-H2A.Z behaves similarly to endogenous H2A.Z
To identify all proteins that interact with H2A and H2A.Z, first, we established three stable cell lines of GFPN, GFPN-H2A and GFPN-H2A.Z in which the expression levels of GFPN-H2A or GFPN-H2A.Z were comparable to that of endogenous H2A or H2A.Z (Supplementary information, Figure S1). Furthermore, GFPN-H2A.Z in the GFPN-H2A.Z stable cell line was incorporated into chromatin and enriched in transcriptional regulatory regions in a similar way to endogenous H2A.Z, as shown by the genome browser snapshots, TSS profile analysis and Venn diagrams (Figure 2).
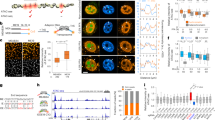
GFPN-H2A.Z is incorporated into chromatin with enrichment in transcriptional regulatory regions similar to endogenous H2A.Z. (A) A snapshot of genome browser image of ChIP-seq data for endogenous H2A.Z and GFPN-H2A.Z-HA. The top panel corresponds to H2A.Z ChIP-seq in shLuc control cells. The bottom panel corresponds to HA CHIP-seq in the GFPN-H2A.Z-HA stable cell line. (B) Venn diagram showing that H2A.Z- and GFPN-H2A.Z-HA-enriched regions highly overlap. (C) Average TSS profile plots for the tag density of the H2A.Z ChIP-seq in control cells and HA ChIP-seq in the GFPN-H2A.Z-HA stable cell line. (D) TSS profile plots for the logarithm of the density of the H2A.Z and GFPN-H2A.Z-HA ChIP-seq experiments.
To further test whether GFPN-H2A.Z shares similar functions with endogenous H2A.Z, we chose to perform rescue experiments by ectopically expressing GFPN-H2A.Z in H2A.Z knockdown cells. First, we compared the chromatin binding patterns of ectopic GFPN-H2A.Z with those of endogenous H2A.Z using ChIP-seq by titrating the concentration of transfected GFPN-H2A.Z construct. From a wide range of DNA amounts used for transfection (0, 0.75, 1.5, 3.0, 6.0, 12 μg), we found that although there is some increased background binding, the GFPN-H2A.Z peaks still correspond closely to endogenous H2A.Z peaks, as shown by the genome browser snap shots, TSS profile analysis and Venn diagrams (Supplementary information, Figures S2 and S3). Thus, we chose to use 3.0 μg of the GFPN-H2A.Z construct in the rescue experiment. While the ectopic expression of GFPN-H2A.Z somehow suppressed the expression of endogenous H2A.Z in the cells, the total H2A.Z level (GFPN-H2A.Z + H2A.Z) in transfected cells was about two-fold of the H2A.Z level in control cells by western blotting (Supplementary information, Figure S4A). Finally, GFPN-H2A.Z rescued the knockdown of endogenous H2A.Z assessed using reverse-transcription qPCR assays (Supplementary information, Figure S4B). These results indicate that GFPN-H2A.Z functions similarly with the endogenous H2A.Z.
Genome-wide identification of H2A- and H2A.Z-interacting proteins
To identify H2A- and H2A.Z-interacting proteins, the stable cell lines expressing GFPN alone, GFPN-H2A and GFPN-H2A.Z were infected with virus carrying three reading frames of GFPC (ReMTH-GFPC1, 2 and 3 viral particles) to tag endogenous genes with the GFPC domain. Following puromycin selection and subsequent induction by Doxycycline treatment for 48 h, green cells were collected by flow cytometry sorting. About 0.5% or 1.1% of cells expressing GFPN-H2A.Z or GFPN-H2A were GFP-positive, while very few cells expressing the GFP N-terminal domain showed green signals (Figure 1B). To identify the genes encoding H2A- or H2A.Z-interacting proteins, total RNAs from 50 to 150K sorted cells were reverse transcribed using oligo dT primers and fragmented to 200-500 bp by sonication (Figure 1C). Following end repair and ligation of a Y-shaped T7 adaptor containing a MmeI recognition site to both ends, the DNA fragments between 250 and 650 bp were isolated from gels. In the second PCR using a T7 primer and three Mme-SD-primers for the three reading frames of the GFPC fusion vector, fusion cDNAs containing HA and SD sequences were amplified. Digestion with MmeI cleaves DNA 20 bp away from its recognition site and thus was used to remove the HA and T7 primer sequences. The resulting DNA fragments were blunt-ended and used to construct RNA-seq libraries (Figure 1C). Six bPPI-seq libraries were pooled and sequenced on Hiseq2500. The short reads were mapped onto the mouse genome sequence to identify the genes encoding the interacting proteins. As expected, ANP32E was detected as a protein that specifically interacts with H2A.Z (Figure 1D). The location of short sequence reads reveals the fusion sites of the endogenous mRNA with GFPC, which are indicated by sharp signals at the exons where fusion occurs, as shown in the genome browser images (Figure 1D). Fusion events may occur at different exons of a gene as indicated by multiple peaks along the gene (Figure 1D). The fusion sites provide information on the location of a protein domain that is required for interacting with the bait protein in the cells. Because of the requirement of spatial proximity for reconstitution of an active GFP from the two GFP fragments, most proteins detected by this strategy show direct interactions with the bait.
To identify H2A- and H2A.Z-interacting proteins, we aligned all reads from the three bPPI-seq libraries, including GFPN, GFPN-H2A and GFPN-H2A.Z, to mouse genome mm9 using TopHat22. The read density in the coding regions of genes was computed for each library. Genes were required to satisfy several criteria to be classified as potential interacting proteins (see Materials and Methods). A significant feature allowing a protein to be considered as interacting with H2A or H2A.Z was if the difference of read density of the gene was significant in comparison to the GFPN control library, computed using the method of Audic and Claverie23 (false-positive discover rate < 0.000001; fold-change of read densities over eight-fold.)
Using the above criteria, we identified 217 and 295 H2A.Z- and H2A-interacting proteins, respectively (Figure 3A; Supplementary information, Tables S1, S2). Overall, the average molecular weights of H2A.Z- and H2A-interacting proteins were comparable to those of all proteins (Supplementary information, Figure S5; Supplementary information, Tables S3, S4). Among the H2A.Z-interacting proteins identified by our bPPI-seq assay, we found six proteins, including Anp32e, Actb, H2afz, Phf14, Mbtd1 and Smarca4, that had been discovered by mass spectrometry analysis14,16,24 and co-IP11 as H2A.Z partners, indicating that the assay is able to detect previously identified interacting proteins.
H2A- and H2A.Z-interacting proteins are enriched in distinctive biological processes. (A) Venn diagram showing that H2A and H2A.Z interact with different sets of proteins. (B) Detection of H2A- and H2A.Z-interacting proteins by bPPI-seq is highly reproducible between replicate experiments. The log2 rpkm values of mRNAs of interacting proteins between replicate experiments are compared in scatter plots. (C) A heat map showing the significance of the gene ontology (GO) terms for the H2A.Z-interacting proteins and H2A-interacting proteins. H2A-interacting proteins and H2A.Z-interacting proteins are involved in distinct biological processes. Highlighted terms are transcription-related terms based on the definitions of GO terms. The color level represents the negative logarithm of the adjusted P-value for the significance of the GO terms. The intensity of red represents greater significance of the GO terms whereas blue represents less significant GO terms. A cutoff of color level = 4.3 (corresponding to a P-value of 0.05) is applied to consider if the GO terms are statistically significant for the gene sets.
To test the reproducibility of bPPI-seq libraries, we plotted the logarithm of read density between two biological replicates (Figure 3B). The replicate samples are highly correlated (r = 0.78, 0.83, respectively).
H2A.Z-interacting proteins
Among the 217 H2A.Z-interacting proteins, 131 (60%) are nuclear proteins and the rest are cytoplasmic proteins. This is biologically relevant since H2A.Z functions in the nucleus. It is likely that the cytoplasmic proteins interact with H2A.Z during its translation in the cytoplasmic compartment and/or during its transportation from cytoplasm to nucleus. It is also possible that some cytoplasmic proteins may have unknown functions in the nucleus where they interact with H2A.Z.
The bPPI-seq data indicated an interaction between H2A.Z and Smarca4 in accord with previous co-IP data11 showing H2A.Z interacts with Smarca4 in mouse endoderm/hepatic progenitor cells, where H2A.Z and Foxa2 recruit nucleosome remodeling complexes including Smarca4, enabling nucleosome depletion and ES cell differentiation11. We also detected an interaction of H2A.Z with INO80, previously shown to mediate the deposition of H2A.Z into chromatin in both yeast and human cells18,25,26,27,28,29,30. The identification of the interaction between INO80 and H2A.Z in our mouse data would suggest the conservation of these interactions and that the promotion of homologous recombination in mouse by INO80 via the removal of H2A.Z occurs similarly to the analogous process in humans.
In addition to the above examples, we also found three data sets in the literature reporting identification of proteins co-purifying with the H2A.Z nucleosomes using affinity purification followed by mass spectrometry analysis14,16,24. These studies identified 45, 55 and 75, proteins, respectively with only seven identified proteins being shared among the three studies (Supplementary information, Table S5) that are mainly components of the SRCAP and TIP60 complexes14,16,24. ANP32E is a component of these complexes, which directly interacts with H2A.Z and functions as an H2A.Z chaperone16. The low reproducibility of these different data sets indicates the high variability of the strategy, which may reflect experimental conditions. Among the proteins identified by AP-MS include the H2A.Z-chaperone ANP32E, of which the direct interaction with H2A.Z is unarguably supported by crystallographic data16,17. Consistently, our bPPI-seq data also revealed these proteins as H2A.Z-interacting proteins (Supplementary information, Table S5). In addition, bPPI-seq also identified H2A.Z itself as an H2A.Z-interacting protein. This may be an indication of the direct contact of the two H2A.Z peptides within a nucleosome core particle as revealed by its crystal structure13. Together these results indicate that proteins known to directly interact with H2A.Z can be identified by both AP-MS and bPPI-seq.
H2A and H2A.Z interact with distinctive protein sets
H2A is a structural protein for nucleosome assembly while H2A.Z is involved in setting up chromatin structure at regulatory regions required for transcriptional activation or repression8. Identification of their interacting partners in the nucleus may shed light on the mechanisms underlying functional differences between them. Among the H2A- and H2A.Z-interacting proteins, there are 265 H2A-specific and 187 H2A.Z-specific proteins, respectively. To gain a perspective of the biological processes in which they might function, we performed a gene ontology analysis31 upon both sets of interacting proteins that we identified. This revealed the H2A.Z-interacting proteins and H2A-interacting proteins to be involved in distinct biological processes (Figure 3C). Gene ontology terms were manually selected reflecting transcriptional regulation based on their definitions and are highlighted in Figure 3C. The H2A.Z-interacting proteins are enriched in those with transcriptional-related terms in contrast to H2A-interacting proteins (Figure 3C). Among the H2A.Z-interacting proteins are 20 transcription factors including Osr1, a protein that may facilitate the targeting of H2A.Z or enable the downstream events of transcriptional activation by recognizing specific DNA sequences. These results are consistent with the previous observation that H2A.Z nucleosomes are involved in the regulation of transcription.
Validation of H2A.Z-interacting proteins
To verify that the identified proteins do interact with H2A.Z, we randomly selected 24 proteins from the H2A.Z-interacting protein list and generated the GFPC fusion protein expression constructs, including the zinc-finger transcription factor Osr1. We also selected an H2A.Z-interacting protein ANP32E, and a non-interacting protein Tinf2 as the respective positive and negative controls. As expected, transfection of the ANP32E construct into the GFPN-H2A.Z stable cell line gave rise to green fluorescence (Figure 4A) whereas the negative control Tinf2 did not. Among the 24 randomly selected proteins, 19 of them (79%) gave rise to green fluorescence (Figure 4A; Supplementary information, Table S6). To further validate the interactors using a different strategy, we performed co-immunoprecipitation assays. We found all the 14 GFPC fusion proteins that we tested could co-immunoprecipitate with GFPN-H2A.Z (Figures 4B and 5A). Notably, although four of the tested proteins have been defined as cytoplasmic proteins in the literature, they displayed nuclear GFP signals in our assay (Figure 4A; Supplementary information, Table S6) and also co-immunoprecipitated with GFPN-H2A.Z (Figure 4B). Therefore, these results validate the proteins identified by bPPI-seq as H2A.Z-interacting proteins and so indicate that bPPI-seq is a robust technique for detecting bait protein-protein interactions in vivo.
Validation of the H2A.Z-interacting proteins. (A) Reconstitution of active GFPs by co-expression of the fusion proteins GFPC-H2A.Z interactor and GFPN-HA-H2A.Z. The construct expressing the fusion protein between GFPC and an H2A.Z-interacting protein was transfected into the stable cell line expressing GFPN-HA-H2A.Z fusion protein and induced using Doxycyline for 48 h. Images of GFP and DAPI staining signals were taken (10×). (B) Co-IP of GFPN-HA-H2A.Z with the fusion protein between GFPC and H2A.Z-interacting proteins. Whole-cell extracts were made from the cells expressing the GFPN-HA-H2A.Z and its interacting fusion proteins as described in (A) and immunoprecipitated using HA antibody. The immunoprecipitates were resolved by SDS-PAGE and blotted by anti-FLAG antibody that detects the H2A.Z interactor fusion protein (left panel) and HA antibody that detects the GFPN-HA-H2A.Z fusion protein (right panel), respectively. The asterisks indicate cytoplasmic proteins based on literature.
Osr1 interacts directly with H2A.Z in vivo and in vitro. (A) Osr1 co-immunoprecipitates with H2A.Z but not H2A from cell extracts. Whole-cell extracts were prepared from NIH3T3 cells co-transfected with Osr1-FLAG-GFPC or Tinf2-FLAG-GFPC and GFPN-HA-H2A or GFPN-HA-H2A.Z for 48 h, followed by immunoprecipitation using anti-FLAG or anti-HA antibodies. The immunoprecipitates were washed with buffers containing 300 or 500 mM NaCl and detected by western blotting with anti-HA antibody. (B) Cartoons showing the full length and truncated H2A.Z proteins expressed in bacteria as MBP fusion proteins, which were purified using Amylose beads. The helices and M6 acid patch domains are indicated. The full length and different truncated Osr1 proteins were expressed in bacteria as GST fusion proteins and purified using glutathione-Sepharose 4B beads. (C) GST or GST-Osr1 fusion proteins bound to beads were incubated with different MBP-H2A.Z fusion proteins. The H2A.Z proteins captured by the beads were detected by western blotting using anti-HA antibody. (D) MBP or MBP-H2A.Z fusion proteins bound to beads were incubated with different GST-Osr1 fusion proteins. The Osr1 proteins captured by the beads were detected by western blotting using anti-FLAG antibody.
Osr1 directly interacts with H2A.Z in vivo and in vitro
The SWR1 and INO80 complexes are responsible for deposition of H2A.Z into chromatin in yeast25,26,27,28,29,30. In mammalian cells, the SRCAP, p400/TIP60 complexes, INO80 chromatin remodeler, and the H2A.Z-specific chaperone, ANP32E14,16,17,24 are implicated in mediating H2A.Z's deposition into chromatin. However, these proteins do not have sequence-specific DNA-binding activities and thus it remains unclear how H2A.Z finds its remarkably specific target sites on chromatin. H2A.Z was reported to act together with the Foxa2 transcription factor to regulate nucleosome structure and gene activation11, suggesting that sequence-specific DNA-binding factors may contribute to its targeting on chromatin. To test whether sequence-specific factors facilitate the binding of H2A.Z to its specific target sites, we further validated the interaction between H2A.Z and Osr1. NIH 3T3 cells were transfected with GFPC-FLAG-Osr1 and either GFPN-HA-H2A or GFPN-HA-H2A.Z for 2 days and cell extracts from these cells were subjected to immunoprecipitation with either anti-FLAG or anti-HA antibodies. As expected, both GFPN-HA-H2A and GFPN-HA-H2A.Z nucleosomes were efficiently pulled down by anti-HA antibody (Figure 5A, top panel, lanes 2, 4, 6, 8). However, only H2A.Z but not H2A was co-immunoprecipitated with Osr1 by anti-FLAG antibody (Figure 5A, top panel, compare lanes 1 and 3 with 5 and 7). This is consistent with the observation that Osr1 interacts specifically with H2A.Z as we had detected by the bPPI-seq assay. As a negative control, we found that Tinf2-FLAG-GFPC was not co-immunoprecipitated with either H2A or H2A.Z (Figure 5A, bottom panel). These results further confirmed that Osr1 specifically interacts with H2A.Z in the cells. To further test whether H2A.Z directly interacts with Osr1 and the domains involved in the interaction, we expressed and purified the full-length and different domains of H2A.Z and Osr1 in bacteria (Figure 5B) and examined their interactions in vitro (Figure 5C and 5D). We immobilized either GST or GST-Osr1 fusion protein on glutathioine beads and found that the latter was able to bind H2A.Z and that this required its middle domain (amino acids 41-80) (Figure 5C). However, in order to recapitulate full binding, both the middle and C-terminal domains of H2A.Z were needed, consistent with previous findings that the acid patch M6 region located in the C-terminal part is crucial for the function of H2A.Z and its interaction with other proteins3,16,29,32,33. By comparison, both the N- and C-terminal domains of Osr1 exhibited binding to MBP-H2A.Z beads (Figure 5D).
Osr1 and H2A.Z co-localize on chromatin and regulate shared targets
The finding that Osr1 and H2A.Z interact directly both in vitro and in vivo suggests that they may co-localize on chromatin. Indeed, ChIP-seq assays indicated that H2A.Z and Osr1 co-localize extensively on chromatin (Figure 6A and 6B). We found that 60% of the co-localizing regions for H2A.Z- and Osr1 corresponded to promoters, whereas 80% of regions occupied only by H2A.Z corresponded to enhancers (Supplementary information, Figure S6A). Interestingly, the promoters bound by both H2A.Z and Osr1 exhibited significantly higher expression levels than the promoters bound by H2A.Z alone (Supplementary information, Figure S6B).
Osr1 facilitates H2A.Z targeting and co-regulates gene expression with H2A.Z. (A) Genome browser image showing the co-localization of H2A.Z and Osr1 ChIP-Seq signals on chromatin. (B) Venn diagram showing the overlap of Osr1 and H2A.Z peaks in the genome. The number in red (blue) in the intersection region refers to the number of Osr1 (H2A.Z) peaks overlapping with H2A.Z (Osr1) peaks. The difference between the number of overlapping peaks is due to the overlap between multiple Osr1 peaks and a single H2A.Z peak. The P-value for the significance of overlap was calculated by using the hypergeometric distribution, in which the smaller number of overlapped peaks was used. (C) The changes in gene expression by knocking down H2A.Z and Osr1 are highly correlated. The gene expression in shLuc, shH2A.Z and shOsr1 cells was measured using RNA-seq. The fold change of gene expression (FC) was calculated for shH2A.Z and shOsr1 by comparing to shLuc. (D) Venn diagram showing the overlap of up-regulated genes after knocking down H2A.Z and Osr1. The P-values were calculated by using the hypergeometric distribution with 22 490 total genes. (E) Venn diagram showing the overlap of down-regulated genes after knocking down H2A.Z and Osr1. The P-value was calculated by using the hypergeometric distribution with 22 490 total genes.
These results suggest that H2A.Z and Osr1 may regulate the expression of shared target genes. To test this hypothesis, we knocked down either H2A.Z or Osr1 and analyzed the mRNA expression profiles in the knockdown cells using RNA-seq. Knocking down H2A.Z led to the upregulation of 1 799 genes and downregulation of 2 346 genes (Supplementary information, Table S7). Knocking down Osr1 led to 927 genes being up-regulated and 1 051 down-regulated genes (Supplementary information, Table S8). Interestingly there was a significant correlation between the up- and down-regulation of genes in response to knockdown of H2A.Z or Osr1 (Figure 6C) and large degrees of overlap (Figure 6D and 6E), suggesting that they not only co-localize to the same target regions on chromatin but also contribute to the expression of the same set of target genes.
Osr1 contributes to H2A.Z targeting on chromatin
To test whether Osr1 contributes to the deposition of H2A.Z on chromatin, we analyzed the ChIP-Seq binding profiles of H2A.Z in control and Osr1 knockdown cells. Knockdown of Osr1 resulted in 112 significantly decreased and 26 significantly increased H2A.Z binding peaks, respectively (Supplementary information, Table S9). Using the genomic SELEX method34 with bacterially expressed Osr1 protein, we identified a consensus binding motif for Ors1 (Figure 7A), which is similar to previously reported Osr1 motif35. We found that 40% of the increased peaks of H2A.Z association had the Osr1 motif whereas a significantly larger fraction (69%) of the decreased H2A.Z peaks contained the Osr1 motif (Figure 7B and 7C). This suggests that the presence of an Osr1 motif may direct Osr1 binding to the site and influence the targeting of H2A.Z. To confirm that Osr1 binding is indeed related to the observed decrease in H2A.Z targeting in the Osr1 knockdown cells, we separated the H2A.Z peaks into two groups: one with Osr1 binding and the other without Osr1 binding based on Osr1 ChIP-seq data (Figure 7D). Interestingly, the same trend was observed as using the Osr1 motif (Figure 7E), indicating that the sequence-specific recognition of DNA by Osr1 contributes to the targeting of H2A.Z to chromatin.
Osr1 facilitates incorporation of H2A.Z to regions containing target motifs. (A) The Osr1 motif logo identified by SELEX using bacterially expressed GST-Osr1 protein and mouse genomic DNA. (B) Volcano plot of changes in H2A.Z ChIP-Seq peaks after knocking down Osr1. The blue and green circles indicate the H2A.Z peaks without and with an Osr1 motif, respectively. The vertical red lines indicate 1.5-fold change and horizontal red line indicates a significant P-value cutoff. (C) A significantly larger fraction of decreased H2A.Z peaks contain an Osr1 motif than the increased H2A.Z peaks. The fractions of the H2A.Z peaks containing Osr1 motif were calculated according to Supplementary information, Table S9. (D) Volcano plot of changes in H2A.Z ChIP-Seq peaks following knockdown of Osr1. The blue and green circles indicate the H2A.Z peaks without and with Osr1 binding, respectively. The vertical red lines indicate a two-fold change and horizontal red line indicates a significant P-value cutoff. (E) The observed relative frequency of decreased and increased H2A.Z peaks that have Osr1 binding. The frequency of the H2A.Z peaks that have Osr1 binding was calculated according to Supplementary information, Table S9.
Discussion
The canonical core histone H2A is a structural subunit for nucleosome assembly and packaging of DNA in the nucleus. Displacement of H2A with the variant H2A.Z is associated with nucleosome destabilization and chromatin accessibility at regulatory regions of transcription such as enhancers and promoters. While these two closely related proteins differ by 30% in amino-acid sequence, what underlies their functional difference is not fully clear. In this study, we developed the novel bPPI-seq assay as a genome-wide screen to identify proteins that interact with H2A or H2A.Z using stable cell lines that expressed fusion proteins of the N-terminal part of GFP with H2A or H2A.Z. The fusion proteins were expressed at levels similar to or lower than the endogenous protein. Furthermore, the H2A.Z fusion protein was incorporated into chromatin in a similar manner to endogenous H2A.Z and was able to rescue the H2A.Z knockdown. Thus, it is very likely that expression of these fusion proteins does not globally disrupt chromatin structure or gene expression in the cells. Using bPPI-seq, we identified 217 H2A.Z-interacting proteins and 295 H2A-interacting proteins. Our data indicated that the majority of the proteins interacting with H2A or H2A.Z fall into different functional groups. While H2A.Z-interacting proteins are highly enriched for transcriptional regulation functions, such proteins are not enriched in the H2A-interacting set. This suggests that the functional differences between H2A and H2A.Z may be controlled by their interacting proteins.
Deposition of H2A.Z onto chromatin is DNA replication-independent and mediated by the ANP32E-containing SRCAP/TIP60 complexes14,16,17,24. However, it is not clear how H2A.Z finds its highly specific target sites in the genome. While our data and those of others suggested that deposition of H2A.Z facilitates the binding of transcription factors to their target sites8,11, we hypothesized that H2A.Z's direct interaction with sequence-specific transcription factors may guide H2A.Z to specific regions in the genome. Indeed, we found that many sequence-specific DNA-binding transcription factors, including Osr1, directly interact with H2A.Z. Our results indicate that Osr1 directly interacts with H2A.Z, co-localizes with H2A.Z on chromatin and contributes to the targeting of H2A.Z to a large number of sites containing the recognition motif of Osr1. The observation that H2A.Z and Osr1 appear to regulate a shared set of genes could be explained either by them functioning in the same pathway or by Osr1 expression requiring H2A.Z. We suggest that both mechanisms could contribute to the regulation of the shared target genes because H2A.Z targeting appears to be regulated by Osr1 in cells.
In this study, we randomly picked Osr1 to demonstrate the sensitivity and robustness of the relatively weak interactions detected by bPPI-seq. Since bPPI-seq revealed multiple transcription factors that interact with H2A.Z, it is highly likely that other transcription factors may also be involved in the recruitment of H2A.Z to target regions and it would be interesting to test this in future studies.
We have shown that bPPI-seq is an effective, rapid and inexpensive method for a genome-wide screen for proteins interacting with a bait protein under physiological conditions. Compared to the current affinity purification/mass spectrometry method, our bPPI-seq method has the following advantages: (1) the proteins detected by bPPI-seq are most likely direct interactors because of the requirement of spatial proximity to reconstitute active GFP; (2) bPPI-seq is capable of detecting weak and transient interactions that require a physiological environment and may not be detectible by other methods; (3) bPPI-seq detects protein interactions in vivo under physiological conditions and thus avoids in vitro artifacts; and (4) bPPI-seq reveals the location of potential interacting domains within a protein based on the fusion site. One caveat for using bPPI-seq is the necessity to control the expression of bait proteins; too high expression of bait proteins, such as by transient expression, may result in detection of artifactual interactions. Therefore, we cannot exclude the possibility that there may be some false positives using bPPI-seq due to its sensitivity. Thus other strategies such as co-localization on chromatin by ChIP-Seq, if applicable, or co-IP may be needed to confirm a true interaction. Although here, we have only demonstrated the applicability of bPPI-seq to the identification of proteins interacting with the chromatin proteins H2A and H2A.Z, we fully expect the approach can be widely applied to identify interacting partners of proteins from other cellular compartments. Thus the method will be instrumental for understanding protein components and interaction networks in various cellular processes.
Materials and Methods
Stable cell lines for bPPI-seq
NIH-3T3 Tet-on cells were transfected with pTight-GFPN, pTight-GFPN-H2A or pTight-GFPN-H2A.Z. After 48 h transfection, we treated the cells with Hygromycin B (200 μg/ml) to select stably transfected cells. After 2 weeks, clones were picked in which the expression level of GFPN, GFPN-H2A or GFPN-H2A.Z was comparable to that of endogenous H2A or H2A.Z with 2 μg/ml Doxycycline treatment for 48 h.
Cell lines, plasmids and antibodies
NIH/3T3 Tet-on 3G cells (Clontech), GP2 293 cells (Clontech) and Lenti-X 293T cells (Clontech) were routinely cultured in DMEM + GlutaMAX (Gibco) supplemented with 10% FBS (Sigma). ReMTH-GFPC1/2/3 and Bait-GFPN-Hygro plasmids were provided by Drs Zhiyong Ding and Gordon Mills (M.D. Anderson Cancer Center)36. The pEF1a-Bait-GFPN-Hygro plasmid was constructed by replacing the CMV promoter of Bait-VNN-Hygro with the EF1a promoter from pEF1a-IRES-AcGFP1 (Clontech). cDNAs of mouse H2A and H2A.Z were subcloned in-frame with an HA tag into the pEF1a-GFP-N-Hygro Vector. The mouse H2B (HistH2bc), Osr1, Tinf2 and ANP32E cDNAs were subcloned in-frame with a Flag tag and GFP-C into the pEF1a-Hygro Vector. The virus packaging vectors were pEco (Clontech) for retrovirus and pGreenPuro, ps-PAX2 and plp-VSVG for lentivirus (System Biosciences). pMal-c2 vector (New England Biolabs) and pGSTag vector (Addgene) were used for expression of MBP and GST fusion proteins.
The following antibodies were used in this study: anti-HA (Roche, 11867423001); anti-FLAG (Sigma, F1804); anti-HA-HRP (Santa Cruz, sc7392 HRP); anti-FLAG-HRP (Sigma, A8592); anti-Osr1 (Santa Cruz, sc68392); anti-H2A.Z (Abcam, ab4174).
Screening for H2A.Z-interacting proteins
ReMTH-VNC 1, 2 and 3 (three different reading frames) and pEco were co-transfected into GP2 293 cells using Lipofectamine 2000 (Invitrogen). Supernatants containing packaged viral particles were collected 48 h after transfection. About 30 ml viral particles were used to infect 5 x 107 3T3 Tet-on 3G cells expressing GFPN-H2A.Z, GFPN-H2A or GFPN alone to avoid multiple integrations. 48 h post-infection, cells were selected in DMEM containing 2 μg/ml puromycin (Sigma) for 2 days and maintained in 1 μg/ml puromycin for 3 days. Then, an equal number of cells infected with viruses for the three reading frames were pooled and induced by 2 μg/ml Doxycycline for 48 h before sorting for green cells. GFP-positive cells were sorted with FACSAria (BD Biosciences).
bPPI-seq library preparation and sequencing
Total RNAs of the sorted cells were extracted using QIAzol (Qiagen) and purified with miRNeasy micro kit (Qiagen) and DNase set (Qiagen). Purified RNAs were reverse transcribed using PrimeScript Reverse Transcriptase (Takara) with a PolyT primer. The cDNAs were amplified by a GFPC-specific (VNC primer) and T7 primers, and fragmented to 200 to 500 bp by sonication. The DNA ends were repaired using an End-It DNA End-repair Kit (Epicentre), followed by treatment with Klenow Fragment exo- (New England Biolabs) to generate a protruding 3′A base used for adaptor ligation. Following ligation of a pair of top-Phops-oligo-17bp/T7Mme I-18bp adaptors, fragments between 250 and 650 bp were isolated from agarose gel. Then the DNA was amplified by MmeI-SD-1, MmeI-SD-2, MmeI-SD-3 and T7 primers to add a MmeI recognition site at both ends, followed by digestion with MmeI (New England Biolabs). After treatment with Klenow Enzyme (New England Biolabs), indexed libraries were prepared with a Multiplexing Sample Preparation Oligonucleotide Kit (Illumina) according to the user's manual. The fragments between 250 and 500 bp were isolated for sequencing on an Illumina next-generation sequencing platform. The primers used for library preparation are listed below:
PolyT primer: 5′-GCAAATACGACTCACTATAGGGATCCTTTTTTTTTTTTTTTT-3′
VNC primer: 5′-AAAGACCCCAACGAGAAGC-3′
T7 primer: 5′-GCAAATACGACTCACTATAGGGATC-3′
top-Phosp-oligo-17bp: 5′-GGTGGATGGATGATGTGAGAATATA-3′
T7MmeI-18bp: 5′-GCAAATACGACTCACTATAGGGATCTCCGACACATCATCCATCCACCT-3
MmeI-SD-1: 5′-CGTCCCAGACTTCCGACGTCGATATATCTGGAGAAAG-3′
MmeI-SD-2: 5′-GACTACGCAGGTCCGACATCGATATATCTGGAGAAAG-3′
MmeI-SD-3: 5′-CAGGTCGACCATCCGACGGCGATATATCTGGAGAAAG-3′
Co-immunoprecipitation to detect interactions in mammalian cells
The HA-tagged H2A or H2A.Z constructs were co-transfected with one of the four constructs expressing FLAG-tagged H2B, ANP32E, Osr1 or Tinf2 protein into NIH3T3 Tet-on 3G cells. 48 h post-transfection, whole-cell lysates were prepared by washing the cells twice with PBS and lysed in ice-cold lysis buffer containing 20 mM Tris-HCl pH 7.5, 1% NP40, 1 mM EDTA, 150 mM NaCl, 10% glycerol and proteinase inhibitor. Chromatin lysates were prepared as described37. Anti-FLAG and anti-HA antibody were conjugated with protein G beads (Invitrogen). The lysates were incubated with antibody-protein G beads with rotation at 4 °C overnight. The beads were then washed four times with lysis buffer containing 150-500 mM NaCl and 0.1% NP-40 or 1× buffer D37. The bound proteins were eluted from the beads by boiling in SDS sample buffer with 100 mM DTT for 5 min. For western blotting, samples were resolved on SDS-PAGE gels according to standard procedures.
MBP-H2A.Z and GST-Osr1 pull-down assay
The various constructs expressing fusion proteins between H2A.Z (full length), H2A.Z-N (amino acids 1-40), H2A.Z-N+M (amino acids 1-80), H2A.Z-M (amino acids 41-80), H2A.Z-M+C (amino acids 41-127), H2A.Z-C (amino acids 81-127) or H2A (full length) and maltose-binding protein (MBP) were made by inserting the corresponding cDNAs into pMal-C2 between BamHI and HindIII sites. Constructs expressing fusion proteins between Osr1 (full length), Osr1-N (amino acids 1-133) or Osr1-C (amino acids 134-266) and GST with a FLAG tag at the C-terminus were made by inserting the corresponding cDNAs into pGSTag between XbaI and XhoI sites. MBP and GST fusion proteins were expressed in BL21-Gold-DE3 cells (Stratagene). Briefly, cells transformed with the expression constructs were cultured in 3 ml Luria Broth with 100 μg/ml carbenicilin (Invitrogen) at 37 °C overnight, which was diluted into 150 ml Luria Broth containing 100 μg/ml carbenicilin and further incubated for 3 h at 37 °C with shaking. The expression of fusion proteins was induced by 1 mM IPTG (Corning) for 2 h. Cells were collected by low-speed centrifugation, re-suspended in 3 ml binding buffer (20 mM Tris-HCl pH 7.5, 150 mM NaCl, 1 mM EDTA, 1 mM DTT, 0.1% NP40, 5% glycerol and proteinase inhibitor), and lysed by three cycles of freeze-and-thaw. Following sonication and high-speed centrifugation, the supernatant was mixed with Amylose Resin (New England Biolabs) for MBP fusion protein or glutathione-Sepharose 4B beads (GE Healthcare) for GST fusion protein for 1 h at 4 °C. The beads were washed four times with binding buffer. The bound proteins were eluted from one-half of the beads while keeping another half beads with proteins still bound in binding buffers. MBP fusion protein was eluted by 20 mM Tris-HCl pH 7.5, 200 mM NaCl, 1 mM DTT and 10 mM maltose. GST fusion protein was eluted by 50 mM Tris-HCl pH 7.5 and 10 mM reduced glutathione. Then MBP or GST fusion protein was mixed with GST fusion protein-bound beads or MBP fusion protein-bound beads, respectively, and incubated with rotation overnight at 4 °C. The beads were washed five times with binding buffer containing 150-500 mM NaCl and bound proteins were eluted with SDS sample buffer with 100 mM DTT, and resolved by SDS-PAGE. Western blotting was performed as described above.
GST-Osr1 genomic-SELEX
GST-Osr1 fusion protein on sepharose beads was generated as described above. Mouse genomic DNA (1 g) was sonicated in a Diagenode Bioruptor (level M, for a total of 10 min alternating 15 s on and 15 s off) to a size range of 100-400 bp. Genomic-SELEX was performed as described34 with some modification. Briefly, washed unloaded sepharose beads and GST-Osr1 beads were incubated with gentle agitation for 30 min at 37 °C in the presence of 500 ng of sonicated DNA. The resin was washed five times with SELEX binding buffer (20 mM Tris-HCl pH 8.0, 10% glycerol, 1 mM DTT, 300 mM NaCl). DNA and protein complex were eluted by elution buffer (50 mM Tris-HCl, pH 7.5 and 10 mM reduced glutathione). Eluted DNA was purified by MinElute Reaction Cleanup Kit (Qiagen). The DNA ends were repaired and a protruding 3′ A generated as described above. Following ligation of a pair of Solexa adaptors to the repaired ends, 150 to 400 bp fragments were recovered from agarose gel. Then DNA was amplified using the index primers for 18 cycles and the 150-400 bp fragments were isolated from agarose gel again. About 300 ng of the resultant library DNA was added again to the new empty sepharose beads and GST-Osr1 beads, and the process was repeated four times. The libraries enriched at rounds two, three and four were sequenced using Illumina high-throughput sequencing.
Knockdown of H2A.Z and Osr1 by shRNA
The RNA interference construct targeting mouse Osr1 was generated by inserting the target oligos into pGreenPuro Lentivector (System Biosciences) and the shH2A.Z construct was described previously8. The target gene sequences are listed below:
shLuc: 5′-GTGCGTTGTTAGTACTAATCCTATTT-3′
shOsr1: 5′-CAGATATATTCACTCCAAA-3′
shH2A.Z: 5′-CCTTATTATCTCAGGACTCTA-3′ (This target sequence is located in the 3′ UTR of H2A.Z so that the rescuing H2A.Z cDNA is resistant to this shRNA.)
Transfection for ChIP-Seq and RNA-Seq
A wide concentration range of pTight-H2A.Z-GFPN was used for transfection (0, 0.75, 1.5, 3.0, 6.0, 12 g together with 12, 11.25, 10.5, 9, 6, 0 μg of control pTight-GFPN, respectively) into tet on 3G cells. About 48 h later 2 μg/ml Doxycycline was added to induce expression for 48 h before fixing cells for anti HA Chip-seq. The shLuc control or knockdown cells were also cross-linked with formaldehyde, and chromatin fractions were prepared by sonication. The ChIP-seq experiments were performed as described4. The RNA-Seq was performed as described38.
Data analysis
bPPI-Seq data analysis We constructed three bPPI-Seq libraries, namely void GFPN, H2A-GFPN and H2A.Z-GFPN. The reads from each bPPI-Seq library were mapped to the mouse genome (mm9) using tophat22. The number of reads located on coding region of each gene was counted in each library. Interactions with the GFPN-fused-protein will lead to insertion of tet-responsive promoter and lead expression of the interacting genes. Selected potentially interacting proteins were required to satisfy three criteria. (1) Genes should have moderate mRNA expression (RPKM >6) quantified by the RNA-seq data. (2) They should have more than six counts per million mapped reads from the bPPI-seq data. (3) The standard deviation of two replicates of the H2A.Z or H2A bPPI-seq (in RPKM) should be ≤ 4. Among these potential interacting proteins, a protein was considered to interact with H2A or H2A.Z if the difference of read density of the gene was significant as compared to the GFPN control library, using the method developed by Audic and Claverie23 (fasle-positive discover rate < 0.000001; fold-change of read densities over eight-fold).
Gene ontology analysis H2A.Z- and H2A-specific interacting proteins are obtained by filtering out the overlap between the two sets of overlapping interacting proteins. We performed gene ontology analysis for the two sets of interacting proteins using the category of biological process in Gene Ontology Consortium31.
ChIP-seq data analysis Raw sequence reads were aligned to the reference mouse genome (UCSC, mm9) by Bowtie39. We allowed for one read per position matching and removed redundant reads for each ChIP-seq library for the alignment. The enriched regions of H2A.Z binding were identified using SICER (window sizes = 200 bp, gap = 200 bp, E-value < 0.001, score ≥ 100)40. The enriched regions of Osr1 binding were identified using model-based analysis for ChIP-Seq (MACS) with P-value cutoff equal to 1e-540.
RNA-seq data analysis We identified 22490 uniquely known genes from the mouse genome (mm9) using the UCSC genome browser database41. To quantify the mRNA level of genes, the RKPM (reads per kilobase of transcript per million mapped reads) values were computed over gene exons from the RNA-seq data sets. Differentially expressed genes between treated and control cells were identified by edgeR42 using two replicates in each condition (FDR < 0.01 and fold-change > 1.5 or < 2/3).
Correlation analysis between two experiments We quantified the similarity between the log2-fold changes (knockdown over control) for the two knockdown experiments (shH2A.Z and shOsr1) for both H2A.Z ChIP-seq data sets and RNA-seq data sets. For the H2A.Z ChIP-seq, all identified peaks from shLuc, shOsr1 and shH2A.Z experiments in one of the replicates were considered, and a set of N combined peaks was obtained by combining the identified peaks if they had any portions of overlap. We calculated the log2-fold changes of read density (normalized by library size) of H2A.Z binding in knockdown experiments versus control for all the N combined peak regions. The Pearson correlation coefficient was computed between the log2-fold changes of the two knockdown experiments to quantify their similarity. The same calculation was applied to the RNA-seq data sets, but the gene exon regions were considered instead of the combined peak regions and the RPKM measure was used instead of the read density measure. We also applied the same strategy to examine the reproducibility between two RNA-seq replicates, but we computed the correlation between the log2 RPKM measure of the two replicates instead of the log2-fold change. Note that combined peak regions (in ChIP-seq analysis) or genes (in RNA-seq analysis) were excluded if the log2 of the total read counts per million (CPM) from both replicates of the knockdown experiment and control experiment in the regions was less than 3.
Motif analysis
To find the Osr1 binding motifs, we applied a motif search algorithm called MEME-ChIP43 to the target genomic sequences identified by the GST-Osr1 SELEX assay.
The raw sequence reads from GST-Osr1 SELEX were aligned to the mouse genome (mm9) by Bowtie, and 144 Osr1-enriched regions were identified using MACS. Among these regions, 67 peaks were selected based on their P-values. In particular, a region size equalling 200 bp was considered for each of the peaks, and each region was located +100 and −100 bp around the peak summit defined by MACS. By applying MEME-ChIP to the sequences in these selected regions and requiring the maximum width of the searching motif to be eight, we found a motif GCTACTGC (E-value = 1.2e-72) as shown in Figure 7A. The motif we found was highly consistent with the Osr1 binding motif discovered by others35.
Analysis of differential H2A.Z binding between shOsr1 and shLuc
We quantitatively compared two H2A.Z ChIP-seq data sets from shOsr1 and shLuc cells in order to identify differential H2A.Z binding sites (peaks) between the two samples. In particular, we employed a normalization method to the two samples, in which the scale factor for normalizing the sequencing depth was computed based on the average of the read densities in the ChIP-enriched regions. Compared to the traditional sequencing depth normalization method44 (i.e., read counts are normalized by the number of all mapped reads), our normalization method was probably more suitable to the ChIP-seq data sets in which a real biological global decrease/increase of the total number of reads occurred in the knockdown samples.
Prior to the computation of differential peaks, the sequencing data were required to go through several pre-processed steps described as follows:
(1) MACS/SICER was used to find out the coordinates of the ChIP-enriched regions (Peaks) for both samples.
(2) A unified set of peaks was obtained from the two samples by taking the union of their peaks found in step 1 and combining any two peaks if they are overlapped. A peak in the unified set was considered to be a “common” peak if it overlapped with at least one peak in both samples.
(3) The number of reads (RK and RC) in each unified peak region was computed for both samples, where the subscript K and C correspond to the knockdown and control samples, respectively.
(4) The read density was computed for each peak (rK and rC), corresponding to the number of read counts divided by the length of the peak.
(5) The values M and A were calculated for each peak i using the following equations:

and

where rki and rCi are the read density of the peak i.
Next we compared the read densities in the knockdown and control samples. To normalize the read densities and remove any systematic bias between the two samples, we assumed that the average of the read densities of the most ChIP-enriched regions was conserved. This assumption was analogous to the assumption that the average of the activities of the house keeping genes is conserved.
In our analysis, we focused on the set of common peaks defined in the data pre-processing step 2. About 1 000 common peaks with the largest value of A were selected. Using the above assumption, a scale factor S was calculated to make the average of the read density in the 1 000 peaks to be the same, where S = <rk>/<rC>, and <rk> or <rC > is the average of the read densities in the 1 000 common peaks of knockdown and control samples, respectively. The new read densities of the knockdown sample were equal to rki divided by S.
For each peak in the unified set, we calculated a P-value to quantify the significance of the differential H2A.Z binding using the method developed by Audic and Claverie23. In particular, the values of normalized read counts were used as inputs in the method23, which was equal to (rki/S) multiply the width of the region. A peak was determined to be a significant differential binding peak if P-value < 0.001, |M| > log2(1.5), and A > 25. A volcano plot of −log10(P-value) versus M was shown in Figure 7B for peaks with A > 25.
Dependency of Osr1 motif and the directions of H2A.Z differential peaks
To elucidate the relationship between Osr1 and H2A.Z binding, we examined whether the directions of the differential H2A.Z binding regions were dependent on the existence of an Osr1 binding motif within the regions. In our analysis, we used FIMO45, a TFBS motif identification software, to find the occurrence of the Osr1 binding motif within the differential H2A.Z bindings regions. The occurrence is considered to be significant with a P-value < 0.0001.
The result was shown in a 2 × 2 contingency table (Supplementary information, Table S10), which has a very significant P-value that equals to 0.0 (calculated by χ2-test). Therefore, as shown by the contingency table, the direction of differential H2A.Z bindings is strongly dependent on the existence of an Osr1 binding motif within the binding region. The relative frequency for each element in the contingency table was computed in which each element was divided by the sum of its row. A result of the relative frequencies was plotted in Figure 7C, which showed that H2A.Z binding was more likely to be decreased in regions containing the Osr1 binding motif when Osr1 was knocked down.
Data accession
The sequencing data have been deposited in GEO (GSE80425).
Author Contributions
KZ conceived the project. YZ performed the experiments. WLK analyzed the data. SL, KC, WJ, QT and WL contributed to the experiments or data analysis. BN and KZ directed the project. YZ, WLK and KZ wrote the manuscript.
Competing Financial Interests
The authors declare no competing financial interests.
Accession codes
References
Larochelle M, Gaudreau L . H2A.Z has a function reminiscent of an activator required for preferential binding to intergenic DNA. EMBO J 2003; 22:4512–4522.
Faast R, Thonglairoam V, Schulz TC, et al. Histone variant H2A.Z is required for early mammalian development. Curr Biol 2001; 11:1183–1187.
Clarkson MJ, Wells JR, Gibson F, Saint R, Tremethick DJ . Regions of variant histone His2AvD required for Drosophila development. Nature 1999; 399:694–697.
Barski A, Cuddapah S, Cui K, et al. High-resolution profiling of histone methylations in the human genome. Cell 2007; 129:823–837.
Zhang H, Roberts DN, Cairns BR . Genome-wide dynamics of Htz1, a histone H2A variant that poises repressed/basal promoters for activation through histone loss. Cell 2005; 123:219–231.
Weber CM, Henikoff JG, Henikoff S . H2A.Z nucleosomes enriched over active genes are homotypic. Nat Struct Mol Biol 2010; 17:1500–1507.
Ku M, Jaffe JD, Koche RP, et al. H2A.Z landscapes and dual modifications in pluripotent and multipotent stem cells underlie complex genome regulatory functions. Genome Biol 2012; 13:R85.
Hu G, Cui K, Northrup D, et al. H2A.Z facilitates access of active and repressive complexes to chromatin in embryonic stem cell self-renewal and differentiation. Cell Stem Cell 2013; 12:180–192.
Subramanian V, Fields PA, Boyer LA . H2A.Z: a molecular rheostat for transcriptional control. F1000Prime Rep 2015; 7:01.
Marques M, Laflamme L, Gervais AL, Gaudreau L . Reconciling the positive and negative roles of histone H2A.Z in gene transcription. Epigenetics 2010; 5:267–272.
Li Z, Gadue P, Chen K, et al. Foxa2 and H2A.Z mediate nucleosome depletion during embryonic stem cell differentiation. Cell 2012; 151:1608–1616.
Skene PJ, Henikoff S . Histone variants in pluripotency and disease. Development 2013. 140:2513–2524.
Suto RK, Clarkson MJ, Tremethick DJ, Luger K . Crystal structure of a nucleosome core particle containing the variant histone H2A.Z. Nat Struct Biol 2000; 7:1121–1124.
Choi J, Heo K, An W . Cooperative action of TIP48 and TIP49 in H2A.Z exchange catalyzed by acetylation of nucleosomal H2A. Nucleic Acids Res 2009; 37:5993–6007.
Vardabasso C, Gaspar-Maia A, Hasson D, et al. Histone variant H2A.Z.2 mediates proliferation and drug sensitivity of malignant melanoma. Mol Cell 2015; 59:75–88.
Obri A, Ouararhni K, Papin C, et al. ANP32E is a histone chaperone that removes H2A.Z from chromatin. Nature 2014; 505:648–653.
Mao Z, Pan L, Wang W, et al. Anp32e, a higher eukaryotic histone chaperone directs preferential recognition for H2A.Z. Cell Res 2014; 24:389–399.
Alatwi HE, Downs JA . Removal of H2A.Z by INO80 promotes homologous recombination. EMBO Rep 2015; 16:986–994.
Hu CD, Chinenov Y, Kerppola TK . Visualization of interactions among bZIP and Rel family proteins in living cells using bimolecular fluorescence complementation. Mol Cell 2002; 9:789–798.
Ding Z, Liang J, Lu Y, et al. A retrovirus-based protein complementation assay screen reveals functional AKT1-binding partners. Proc Natl Acad Sci USA 2006; 103:15014–15019.
Liu D, Yang X, Yang D, Songyang Z . Genetic screens in mammalian cells by enhanced retroviral mutagens. Oncogene 2000; 19:5964–5972.
Trapnell C, Pachter L, Salzberg SL . TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 2009; 25:1105–1111.
Audic S, Claverie JM . The significance of digital gene expression profiles. Genome Res 1997; 7:986–995.
Ruhl DD, Jin J, Cai Y, et al. Purification of a human SRCAP complex that remodels chromatin by incorporating the histone variant H2A.Z into nucleosomes. Biochemistry 2006; 45:5671–5677.
Kobor MS, Venkatasubrahmanyam S, Meneghini MD, et al. A protein complex containing the conserved Swi2/Snf2-related ATPase Swr1p deposits histone variant H2A.Z into euchromatin. PLoS Biol 2004; 2:E131.
Krogan NJ, Keogh MC, Datta N, et al. A Snf2 family ATPase complex required for recruitment of the histone H2A variant Htz1. Mol Cell 2003; 12:1565–1576.
Mizuguchi G, Shen X, Landry J, Wu WH, Sen S, Wu C . ATP-driven exchange of histone H2AZ variant catalyzed by SWR1 chromatin remodeling complex. Science 2004. 303:343–348.
Ranjan A, Mizuguchi G, FitzGerald PC, et al. Nucleosome-free region dominates histone acetylation in targeting SWR1 to promoters for H2A.Z replacement. Cell 2013; 154:1232–1245.
Wu WH, Alami S, Luk E, et al. Swc2 is a widely conserved H2AZ-binding module essential for ATP-dependent histone exchange. Nat Struct Mol Biol 2005; 12:1064–1071.
Yen K, Vinayachandran V, Pugh BF . SWR-C and INO80 chromatin remodelers recognize nucleosome-free regions near +1 nucleosomes. Cell 2013; 154:1246–56.
Huang da W, Sherman BT, Lempicki RA . Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc 2009; 4:44–57.
Jensen K, Santisteban MS, Urekar C, Smith MM . Histone H2A.Z acid patch residues required for deposition and function. Mol Genet Genomics 2011; 285:287–296.
Subramanian V, Mazumder A, Surface LE, et al. H2A.Z acidic patch couples chromatin dynamics to regulation of gene expression programs during ESC differentiation. PLoS Genet 2013; 9:e1003725.
Reiss DJ, Mobley HL . Determination of target sequence bound by PapX, repressor of bacterial motility, in flhD promoter using systematic evolution of ligands by exponential enrichment (SELEX) and high throughput sequencing. J Biol Chem 2011; 286:44726–38.
Badis G, Berger MF, Philippakis AA, et al. Diversity and complexity in DNA recognition by transcription factors. Science 2009; 324:1720–1723.
Ding Z, Liang J, Lu Y, et al. A retrovirus-based protein complementation assay screen reveals functional AKT1-binding partners. Proc Natl Acad Sci USA 2006; 103:15014–15019.
Draker R, Ng MK, Sarcinella E, Ignatchenko V, Kislinger T, Cheung P . A combination of H2A.Z and H4 acetylation recruits Brd2 to chromatin during transcriptional activation. PLoS Genet 2012; 8:e1003047.
Wei G, Hu G, Cui K, Zhao K . Genome-wide mapping of nucleosome occupancy, histone modifications, and gene expression using next-generation sequencing technology. Methods Enzymol 2012; 513:297–313.
Langmead B, Trapnell C, Pop M, Salzberg SL . Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 2009; 10:R25.
Zhang Y, Liu T, Meyer CA, et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol 2008; 9:R137.
Karolchik D, Hinrichs AS, Furey TS, et al. The UCSC Table Browser data retrieval tool. Nucleic Acids Res 2004; 32(Database issue):D493–D496.
Robinson MD, McCarthy DJ, Smyth GK . edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010; 26:139–140.
Machanick P, Bailey TL . MEME-ChIP: motif analysis of large DNA datasets. Bioinformatics 2011; 27:1696–1697.
Kharchenko PV, Tolstorukov MY, Park PJ . Design and analysis of ChIP-seq experiments for DNA-binding proteins. Nat Biotechnol 2008; 26:1351–1359.
Grant CE, Bailey TL, Noble WS . FIMO: scanning for occurrences of a given motif. Bioinformatics 2011; 27:1017–1018.
Acknowledgements
We thank the NHLBI DNA Sequencing Core facility for sequencing the libraries and the NHLBI Flow Cytometry Core facility for sorting the cells. The work was supported by Division of Intramural Research, NHLBI, NIH. BN was supported by the National Key Research and Development Project (2016YFA0502203). YZ was supported by a PhD Student Fellowship from the China State Scholarship Fund (2013-2014).
Author information
Authors and Affiliations
Corresponding authors
Additional information
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
Supplementary information
Supplementary information, Figure S1
The expression of GFPN, GFPN-H2A and GFPN-H2A.Z fusion proteins were induced by DOX. (PDF 88 kb)
Supplementary information, Figure S2
Comparison between ChIP-seq data for endogenous H2A.Z and over-expressed GFPN-H2A.Z. (PDF 153 kb)
Supplementary information, Figure S3
Venn diagrams for comparing the overlap between the ChIP-seq enriched regions of the endogenous H2A.Z in shluc control cells and the ectopically expressed GFPN-H2A.Z-HA proteins in cells transfected with (A) 12μg, (B) 6μg, (C) 3μg, (D) 1.5μg, (E) 0.75 μg, and (F) 0μg of GFPN-H2A.Z-HA expression construct. (PDF 108 kb)
Supplementary information, Figure S4
GFPN-H2A.Z is incorporated into chromatin and rescues the function of endogenous H2A.Z. (PDF 78 kb)
Supplementary information, Figure S5
Boxplots of molecular weights of three groups of proteins: (1) All proteins, (2) H2A.Z-interacting proteins identified by bPPI-seq, (3) H2A-interacting proteins identified by bPPI-seq. (PDF 48 kb)
Supplementary information, Figure S6
Promoters bound by both H2A.Z and Osr1 exhibit higher expression levels than those bound by only H2A.Z. (PDF 114 kb)
Supplementary information, Table S1
List of H2A.Z-interacting proteins and their RPKM and FDR values. (XLS 56 kb)
Supplementary information, Table S2
List of H2A-interacting proteins and their RPKM and FDR values. (XLS 55 kb)
Supplementary information, Table S3
List of H2A.Z-interacting proteins and their molecular weights. (XLS 52 kb)
Supplementary information, Table S4
List of H2A-interacting proteins and their molecular weights. (XLS 59 kb)
Supplementary information, Table S5
Comparison of H2A.Z-interacting proteins identified by different groups and different methods (PDF 187 kb)
Supplementary information, Table S6
Validation tests using GFP reconstitution and co-IP assays for 24 randomly selected proteins. We also used ANP32E and Tinf2 as the positive and negative controls, respectively. (XLS 11 kb)
Supplementary information, Table S7
Up- and down-regulated genes by knocking down H2A.Z. (XLS 441 kb)
Supplementary information, Table S8
Up- and down-regulated genes by knocking down Osr1. (XLS 223 kb)
Supplementary information, Table S9
Contingency table showing the number of altered H2A.Z peaks with or without Osr1 motifs by knocking down Osr1. (PDF 318 kb)
Supplementary information, Table S10
Contingency table showing the number of altered H2A.Z peaks with or without Osr1 binding by knocking down Osr1 (PDF 296 kb)
Rights and permissions
About this article
Cite this article
Zhang, Y., Ku, W., Liu, S. et al. Genome-wide identification of histone H2A and histone variant H2A.Z-interacting proteins by bPPI-seq. Cell Res 27, 1258–1274 (2017). https://doi.org/10.1038/cr.2017.112
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2017.112
Keywords
This article is cited by
-
The histone variant H2A.Z in gene regulation
Epigenetics & Chromatin (2019)
-
Catching global interactions in vivo
Cell & Bioscience (2017)