Abstract
Leptospirosis is a widespread zoonotic disease caused by pathogenic spirochetes of the genus Leptospira that infects humans and a wide range of animals. By combining computational prediction and high-accuracy tandem mass spectra, we revised the genome annotation of Leptospira interrogans serovar Lai, a free-living pathogenic spirochete responsible for leptospirosis, providing substantial peptide evidence for novel genes and new gene boundaries. Subsequently, we presented a high-coverage proteome analysis of protein expression and multiple posttranslational modifications (PTMs). Approximately 64.3% of the predicted L. interrogans proteins were cataloged by detecting 2 540 proteins. Meanwhile, a profile of multiple PTMs was concurrently established, containing in total 32 phosphorylated, 46 acetylated and 155 methylated proteins. The PTM systems in the serovar Lai show unique features. Unique eukaryotic-like features of L. interrogans protein modifications were demonstrated in both phosphorylation and arginine methylation. This systematic analysis provides not only comprehensive information of high-coverage protein expression and multiple modifications in prokaryotes but also a view suggesting that the evolutionarily primitive L. interrogans shares significant similarities in protein modification systems with eukaryotes.
Similar content being viewed by others
Introduction
In the past decade, leptospirosis caused by pathogenic Leptospira species has been recognized as an important emerging infectious zoonose world widely 1. It is estimated that more than 500 000 human cases of severe leptospirosis occur annually in the world with a mortality rate of up to 23% 2. In addition, leptospirosis results in significant economic loss in a wide range of livestock. Leptospira interrogans is the most frequently reported pathogen responsible for leptospirosis, with its serogroup Icterohaemorrhagiae representing more than half of the leptospires found in human infections. Despite advances in prevention and therapy, the molecular mechanisms of pathogenesis in leptospirosis remain almost completely unknown. The genome sequences of the pathogenic L. interrogans serovar Lai and serovar Copenhageni, the pathogenic L. borgpetersenii serovar Hardjo and the saprophytic L. biflexa serovar Patoc have been reported 3, 4, 5, 6. The accomplishment of these sequencing projects greatly facilitated the studies of Leptospira physiology and pathology at the genomic 4, 5, 6, transcriptomic 7, 8 and proteomic levels 9, 10, 11, 12, 13, 14. However, there are discrepancies in the annotations for the same genome sequences by different institutions or by using diverse prediction methodologies, which hinders further studies 8. Recently, mass spectrometry-based approaches have been employed to support the annotations of prokaryotic and eukaryotic genomes, and to more accurately determine the existence and boundaries of genes 15, 16, 17.
Posttranslational modifications (PTMs) play crucial roles in regulating protein functions in bacterial physiology and virulence 18, 19, 20. In particular, phosphorylation, acetylation and methylation, which are the most extensively studied PTMs, are all acknowledged to be important for regulating protein activities 21, 22, 23. Recently, the analysis of phosphorylation at Ser/Thr/Tyr residues in the model bacteria Bacillus subtilis and Escherichia coli, and the analysis of acetylation at Lys residue in E. coli were accomplished by taking advantage of the modified peptide enrichment techniques 24, 25, 26. A variety of phosphoproteins and acetylproteins were found to be involved in metabolic processes. Nevertheless, such studies each mainly focus on a certain individual PTM, and thus do not represent the real image of protein status regulated by multiple PTMs. In Leptospira species, there has not yet been systematic analysis of PTMs. It is therefore highly desirable to develop a strategy for simultaneously detecting multiple types of PTMs together with the global protein expression in Leptospira species.
In this study, we described the investigations of genome annotation, global protein expression and multiple types of PTMs, including phosphorylation, acetylation and methylation in the pathogen L. interrogans serovar Lai. A combined map with extensive protein expression profile and multiple PTMs was established. Our proteomic data provided significant information for revising the genome annotation of L. interrogans, and offered additional insights into the physiology and pathogenesis of this organism.
Results
Rectifying the genome annotation of L. interrogans by utilizing MS/MS data
L. interrogans serovar Lai is a free-living Leptospira pathogen that can cause severe leptospirosis. Its genome has been sequenced in 2003, consisting of a 4.33 Mb large circular chromosome and a 0.36 Mb small chromosome 3. The original genome annotation released in GenBank revealed 4 727 protein-coding sequences (CDSs), including 16.0% CDSs shorter than 50 codons. These short CDSs are usually beyond the detection power of available gene prediction methods and many methods usually ignore such short CDSs. In a later revised annotation for the serovar Lai by Adler and his colleagues, almost all of these short CDSs were excluded and 3 614 CDSs were eventually annotated 8.
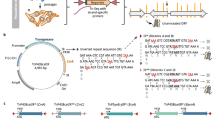
In this study, we rectified the annotation of the serovar Lai genome by combining computational prediction tools and the proteomic data (Figure 1). First, we performed a six-frame translation of the serovar Lai genome and retained all potential CDSs longer than 20 codons to construct a six-frame translation database that contained 70 903 candidate CDSs, 14.0 times and 18.6 times more than those in the originally published database and Adler's database, respectively. After filtering by stringent computational criteria, 3 943 CDSs remained as the computationally predicted CDSs.
In parallel, in order to improve the annotation of the serovar Lai genome, ∼1.37 million high-accuracy tandem mass spectrometry (MS/MS) spectra obtained by the Yin-yang multidimensional liquid chromatography (MDLC) system 27 coupled with LTQ-Orbitrap mass spectrometry (Yin-yang MDLC-MS/MS) from the serovar Lai were searched against the six-frame translation database. After filtering the identified peptides with a 1.0% false-positive ratio (FPR) 28, we selected those CDSs matched by at least two unique peptides to validate and rectify the genome annotation. Consequently, we found peptide evidence for 2 158 CDSs in the six-frame translation database. The majority of them (2 148 CDSs) were present in the computationally predicted CDSs database, while another ten CDSs were identified (Supplementary information, Table S1). Among them four novel CDSs were annotated for the first time in the serovar Lai genome, never showing up in either of the previously published annotations. Since the short CDSs were susceptible to being lost after the stringent filtration of computational prediction, the reason for the absence of these CDSs in the annotations may be due to their relatively short lengths. For instance, the ribosomal proteins, L33 (54 residues) and L36 (37 residues), were absent in the computation-based annotation, while we identified two unique peptides assigned to these two CDSs, and thus putting them back into our revised annotation.
In addition, we amended the gene boundaries of some CDSs by utilizing the MS/MS spectra data. Compared to the annotation released in GenBank, our annotation rectified the start codons of 558 CDSs. Proteomic data provided peptide evidence for new N-terminus of 31 CDSs. As an example, the DNA sequence from 2 197 816 to 2 199 286 in the large chromosome was originally annotated to contain two CDSs, LA2216 (2 197 816 to 2 197 965) and LA2217 (2 198 111 to 2 199 286), using TTG as their start codons from the +1 reading frame and the +2 reading frame, respectively, while in our computation-based annotation LA2216 was excluded and the start codon of LA2217 made an upstream shift in the same reading frame, leading to an increase of 72 codons (2 197 895 to 2 198 110) in the N-terminus of this gene (Figure 2). In this case, we identified five tryptic peptides containing 61 residues in the increased region, so we rectified the previous computation-based annotations for these two proteins according to our MS/MS data. On the other hand, our computation-based annotation might wrongly determine gene start codons. For example, the N-terminus of the gene LA1032 encoding enoyl-CoA hydratase predicted by our computational tools showed a decrease of 31 codons compared to that in the original annotation. We found four unique peptides containing 28 residues in this region by searching the six-frame translation database and therefore retrieved this correct start codon. Similarly, the start codons of another six protein-coding genes were found to be wrongly annotated and were eventually rectified according to the proteomic data.
Among protein-coding genes in the serovar Lai genome, 73 genes were previously predicted to be pseudogenes by Adler and coworkers 6. To our surprise, we detected peptides uniquely assigned to five proteins, which are encoded by pseudogenes (LA0703, LA1005, LA2083, LA4202 and LB007).
In conclusion, based on the proteomic data, we present a relatively reliable annotation of L. interrogans serovar Lai genome, including a total of 3 953 CDSs. In comparison to the previously released annotations, our annotation added 66 novel CDSs, among which 4 were purely derived from the MS/MS data.
Establishment of the proteomic profile of L. interrogans
Subsequently, by combining MS/MS spectra with the individual PTM database-searching strategy 29, we systematically analyzed the protein expression and multiple types of PTMs, including phosphorylation, acetylation and methylation in L. interrogans. In order to maximally obtain the phosphopeptide identification, we also utilized the titania beads as a supplement to enrich phosphopeptides. In total, 2 540 proteins accounting for 64.3% of the whole predicted L. interrogans proteins were assigned by 18 564 identified unique peptides (Supplementary information, Table S2). This represents the second highest proteome coverage so far for prokaryotes, only lower than that (88.6%) of the simple species Mycoplasma mobile (∼0.78 Mb), which has a genome of only a one-sixth size of that of L. interrogans 30. Recently, 2 221 (60.7%) out of 3 658 predicted proteins in L. interrogans serovar Copenhageni were identified 14, among which 2 178 had orthologs in the serovar Lai and 1 982 of these orthologs were identified in our study.
The high quality of our protein identification was based on the following multiple criteria: the FPR at the peptide level was lower than 1.0%; the average precursor ion mass tolerance of identified peptides was 3.7 p.p.m.; 88.4% of 2 540 identified proteins were mapped by at least two unique peptides; all modified peptides and the peptides singly assigned to proteins were manually checked (the MS/MS spectra of modified peptides are presented in Supplementary information, Figure S1).
We identified 22 and 27 proteins that were not predicted in the original annotation and Adler's annotation, respectively. Among them, 18 and 13 proteins were assigned by at least two unique peptides, respectively. Totally, among 66 proteins that were firstly annotated by us in the serovar Lai genome, 11 were identified by MS/MS and 7 were assigned by at least two unique peptides.
The serovar Lai genome contained 1 832 (46.3%) hypothetical proteins, including 886 conserved hypothetical proteins. There were 927 hypothetical proteins detected in our study and 410 belonged to conserved hypothetical proteins, among which 775 and 329 were assigned by at least two unique peptides.
Identification of PTMs in L. interrogans
We totally identified 32, 46, 104 and 58 proteins corresponding to phosphorylation, Lys acetylation, Glx (Glx denotes Glu/Gln) methylation and Lys/Arg methylation, respectively (Supplementary information, Table S3). Among these 223 modified proteins, we found 14 proteins that were modified by at least two different PTMs. There were 27 phosphorylated sites, 54 Lys-acetylated sites, 135 Glx-methylated and 64 Lys/Arg-methylated sites unambiguously detected in these modified proteins.
Phosphorylation of Ser/Thr/Tyr/Asp
Among the unambiguous 27 phosphosites, 13 (41.9%) were at Ser, 6 (21.4%) were at Thr, 5 (17.9%) were at Tyr and 3 (10.7%) were at Asp. More than 50% (13) of the sequences containing the Ser/Thr/Tyr-phosphorylated sites matched the known target motifs of nine eukaryotic protein kinases. Particularly, the target motif of the cAMP-dependent protein kinase PKA (R-X-pS/pT) was overrepresented among the phosphorylated Ser/Thr sites (P = 0.0064).
Acetylation of Lys
We identified 54 unambiguous Lys-acetylated sites. The sequences surrounding the Lys-acetylated sites revealed some consensus residues, such as Gly at the +1 position (P = 0.0020), Lys at the −5 position (P = 0.013) and the −2 position (P = 0.019).
Methylation of Glx
Among the 135 identified Glx-methylated sites, 114 (84.4%) were at Glu and 21 (15.6%) were at deamidated Gln. Glx methylation primarily occurred in the Glx-Glx pair, preferentially at the first Glx residue. Among the 49 Glx-Glx pairs containing methylation sites, 20 were methylated at both of the Glx residues, 12 were only at the first Glx residue, three were only at the second and the rest 14 were ambiguous. Furthermore, we analyzed the amino acids surrounding the methylated Glx-Glx pairs and found that small amino acids (A/G/S/T) at the −2 (P = 0.0027) and +2 (P = 0.040) positions of the Glx-Glx pair were overrepresented.
Methylation of Lys and Arg
Among 64 Lys/Arg-methylated sites, there were 13 (20.3%) monomethyl, 14 (21.9%) dimethyl and 13 (20.3%) trimethyl sites of Lys, along with 10 (15.6%) monomethyl and 14 (21.9%) dimethyl sites of Arg. Interestingly for LipL32, mono-, di- and tri- methylation at Lys171 were identified, respectively. Arg methylation has been found to play important roles in cellular processes in eukaryotes 31. We found this modification in L. interrogans by utilizing MS/MS data, however, further experiment validation is needed.
Functional classes of L. interrogans proteins
BLAST was performed against NCBI COG (Clusters of Orthologous Genes) database 32 to obtain function descriptions of L. interrogans proteins, and 55.8% of the entire predicted proteins and 67.9% of the MS-detected proteins had assigned functions in the COG database (Table 1). The distribution of protein proportions in different functional classes is illustrated in Figure 3. The MS-detected proteins, including the modified proteins, were classified into a wide variety of functional classes. In particular, three sets of modified proteins were statistically overrepresented in certain functional classes.
The identified phosphoproteins were overrepresented in signal transduction (P = 1.4E-8). Eight anti-anti-σ factors (LA0091, LA0653, LA0839, LA0861, LA1327, LA2434, LA3070 and LA3096) were detected to contain phosphosites in their STAS (Sulfate Transporter and Anti-Sigma factor antagonist) domains. In addition, LipL32 (LA2637), which is an important outer membrane lipoprotein and is specific for pathogenic Leptospira species, was found to be phosphorylated at Tyr170 residue, which was suggested by Pro-Q Diamond dye staining (Supplementary information, Figure S2) and validated by mass spectrometry analysis before and after alkaline phosphatase treatment (Figure 4).
(A) MS/MS spectrum of the phosphopeptide, LDDDDDGDDTpYKEER, of LipL32 (LA2637). (B) Pro-Q staining and Coomassie brilliant blue (CBB) staining of the gel bands containing LipL32 and intensity comparison of the peptides LDDDDDGDDTYKEER and QAIAAEESLK of LipL32 before (AP−) and after (AP+) alkaline phosphatase (AP) treatment. After dephosphorylation by AP, the nonphosphorylated isoform (LDDDDDGDDTYKEER) of the phosphopeptide increased its amount, while the amount of another peptide (QAIAAEESLK) was invariable. (C) The localization of the modified sites in the crystal structure of LipL32.
The acetylproteins were overrepresented in transcription regulation (P = 1.5E-4) and signal transduction (P = 0.0015). Acetylation sites were found in RNA polymerase β subunit, σ factors (LA2101 and LA2232) and RsbU homologs (LA2122, LA2435 and LB112). Some kinases were acetylproteins, including histidine kinases (LA1745 and LB290) and Ser/Thr kinase (LA1164), as were some proteins involved in the acetyl group transfer, such as the acetyl-CoA acetyltransferase (LA0457), the ribosomal-protein-serine-acetyltransferase (LA3315) and the histone deacetylase family protein (LA0915).
The Glx-methylated proteins were overrepresented in cell motility (P = 5.0E-5). Among the proteins involved in chemotaxis, four methyl-accepting chemotaxis proteins (MCPs) (LA0049, LA0676, LA2426 and LA4243) and five flagella proteins (FlgD (LA2849), FliF (LA2591), FlgB (LA0347), FlgH (LA2665) and FliK (LA2850)) contained Glx-methylated sites. The chemotaxis histidine kinase CheA1 (LA1251) was also detected with a Glx-methylated site.
The Lys/Arg-methylated proteins did not show preference for any functional class. For proteins involved in translation, the ribosomal proteins L1 (LA3423) and L27 (LA0851), the translational initiation factor IF-2 (LA0943) and the elongation factor Ts (LA3297) were found to be methylated at Lys or Arg. Thereamong, the K137 residue of the ribosomal protein L1 was found to have two methylation states, dimethylation and trimethylation. Likewise, we found all three different states of methylation occurring at K171 residue of LipL32.
Evolutionary conservation of L. interrogans proteins
To investigate the evolutionary conservation of L. interrogans proteins, we searched for orthologs of L. interrogans proteins against 49 bacterial species across the phylogenetic tree, as well as against 10 archaea and 9 eukaryotes by performing two-dimensional BLASP (Supplementary information, Table S4). The measurement of protein conservation followed the technique used in the recent study by Macek et al. 25. In brief, the ortholog number of a category of L. interrogans proteins in each analyzed species was counted and was divided by the number of this category of proteins. The percentage was reported as the conservation of this category of proteins in this species. For example, 969 out of 3 953 predicted L. interrogans proteins had their orthologs in E. coli K12, and therefore the conservation of the predicted L. interrogans proteins in E. coli K12 was considered as 24.5% (969/3 953). The MS-detected proteins were on average more conserved than the predicted proteins in L. interrogans database throughout the three superkingdoms, and especially than the unidentified proteins (Figure 5). For example, the conservation of the predicted total proteins in bacteria was averagely 24.3% and the conservation of the unidentified proteins was only 8.6%, while the conservation of the identified proteins was 33.0%. Our analysis indicated that the conserved proteins were mainly involved in essential housekeeping functions such as protein translation, and the metabolism of amino acids and nucleotides. In addition, the conservation of phosphoproteins (20.7% in bacteria) was lower than that of the MS-detected total proteins and that of the predicted proteins. Acetylproteins (35.1% in bacteria) and Glx-methylated proteins (32.1% in bacteria) were more conserved than the predicated proteins, but they did not show a pronounced enhancement in conservation compared to the MS-detected total proteins. Except in archaea, Lys- and Arg-methylated proteins showed a little more conservation than the MS-detected total proteins in bacteria (40.3% and 39.6%, respectively, vs 33.0%) and eukaryotes (26.6% and 23.6%, respectively, vs 17.8%).
Discussion
Improving genome annotation of L. interrogans by proteomic data
Although the prevalent gene prediction tools such as Glimmer 33 and ORPHUES 34 are very sophisticated tools for searching genes against the prokaryotic genomes, some challenges still remain. Those genes shorter than 50 codons seldom get across the stringent computational filtration and therefore are missed. The gene boundaries are often difficult to determine. In addition, for a number of hypothetical genes without functional assignments, their existence in organisms is uncertain. Importantly, reliable PTMs cannot be directly obtained from computational prediction. The genome of the pathogen L. interrogans serovar Lai has been completely sequenced and two divergent annotations by independent institutions are present. In this study, high-resolution MS/MS spectra data acquired by using Yin-yang MDLC-LTQ-Orbitrap-MS/MS provided large-scale peptide evidence for validating computationally predicted CDSs in the serovar Lai. Moreover, based on the MS/MS data, we retrieved 10 additional CDSs and found 4 novel CDSs, which complemented the annotation based on pure computational prediction. In addition, the identified peptides helped us to amend the gene boundaries of 38 CDSs incorrectly released in the GenBank or wrongly predicted by our computational prediction tools.
Protein expression profiling of L. interrogans
We detected 64.3% of predicted proteins of the serovar Lai in the normal growth phase. This represents the second highest proteome coverage in prokaryotes. More than 88.4% of MS-detected proteins were assigned by multiple unique peptides, and therefore our proteome data provide high-confidence for validating and rectifying the genome annotation of the serovar Lai. As an example, we detected peptides that mapped to five genes previously annotated as pseudogenes. Meanwhile, a wide range of hypothetical proteins were identified to be expressed in the serovar Lai. In addition, we compared protein identification in the serovar Lai with that in the serovar Copenhageni 14 and found that these two serovars had similar protein expression profiles in vitro: 89.0% expressed proteins in the serovar Copenhageni were also identified in the serovar Lai.
Biological relevance of modified proteins in L. interrogans
The modified proteins, including phosphoproteins, acetylproteins and methylproteins, are distributed in a variety of functional classes, suggesting that PTMs are extensively involved in cellular processes of L. interrogans. In particular, the acetylproteins of L. interrogans were overrepresented in transcription regulation, especially in the activity regulation of RNA polymerase complex. Meanwhile, a high percentage of phosphoproteins are involved in the regulation of the σ factors. According to the previous knowledge and the localizations of these modified sites in the components of RNA polymerase complex, it may be estimated that most of these modification events are employed to negatively regulate the activities of the targets. For example, the RNA polymerase β subunit (LA3420) and primary σ70-factor RpoD (LA2232) are acetylated in their critical DNA-binding domains, “RNA polymerase Rpb2, domain 2” for β subunit and “Region 4 domain” for RpoD, respectively. These acetylation events may disrupt the interaction between the corresponding proteins and DNA segments, since they reduce positive charges of the DNA-binding domains. The mechanism by which bacteria utilize the interaction between anti-σ factors and anti-anti-σ factors to regulate the activities of the alternative σ factors has been elucidated in B. subtilis and it is present in a wide range of bacteria 35, 36. Under normal conditions, anti-anti-σ factors are phosphorylated by anti-σ factors at serine residues in their STAS domains and release anti-σ factors to bind alternative factors, resulting in the silencing of stress-induced gene transcription. Eight anti-anti-σ factors were found to be phosphorylated at serine residues in their STAS domains in this study, indicating that a similar regulatory mechanism is also present in L. interrogans. Moreover, this may not be the sole way to negatively regulate the alternative σ factors under normal conditions. LA2101 is an alternative σ factor in L. interrogans, belonging to the extracytoplasmic function subfamily, in which most of the members respond to signals from the extracytoplasmic environment and trigger the transcription of stress-induced genes. We found one acetylation site in “Region 4 domain” of LA2101. This acetylation event may block the binding between this σ-factor and DNA, and therefore have the same effect on suppressing the σ-factor, like phosphorylation of anti-anti-σ factors. This finding suggests that bacteria may employ direct mechanisms to suppress the alternative σ-factors in the normal growth phase and the corresponding mechanisms should be further studied.
Previous studies about Glx methylation focused mainly on chemotaxis receptors. In this study, we screened Glx methylation on a global scale and found that Glx-methylated proteins were distributed in a variety of functional classes, suggesting that this type of modification participates in regulating more cellular processes than previously known. Meanwhile, the MS-detected Glx-methylated sites showed some sequence preferences that were found in chemotaxis receptors and the Glx-methylated proteins also showed overrepresentation in cell motility. Besides four MCP members, we found that the chemotaxis kinase CheA1 was Glx-methylated. In addition, several flagella proteins, including FlgD, FliF, FlgB, FlgH and FliK, were found to contain Glx-methylated sites (Figure 6). Glx methylation in these flagella proteins may participate in the regulation of flagella motility and influence the chemotaxis system of L. interrogans, beyond the level of chemoreceptors. We also found acetylation in LA2426. Although there have not yet been other reports about acetylation of MCPs, acetylation of CheY, the response factor in chemotaxis, has been validated to enhance signaling by CheY 37. Based on these findings, it is likely that novel regulatory mechanisms may be present in chemotaxis.
MS-detected known and novel modified proteins in the chemotaxis system of L. interrogans. Terms of “MCP”, “W”, “A”, “B”, “Y”, “R” and “A C” denote Methyl-accepting chemotaxis protein, CheW, CheA, CheB, CheY, CheR and adenylate cyclase, respectively. “pS”, “pT”, “pY”, “pH”, “pD”, “Me” and “Ac” denote phosphoserine, phosphothreonine, phosphotyrosine, phosphohistidine, phosphoasparagine, methyl-Glx and acetyl-Lys, respectively. The symbol “+” denotes the known and MS-detected modifications in our study and the symbol “*” denotes those first revealed in our study.
LipL32 is a major lipoprotein in the outer membrane and is specific for pathogenic Leptospira species 38. In vivo studies showed that it is the major target of the antibody response in humans and other animals. The peptide 160LDDDDDGDDTYKEER174 in LipL32 was detected with the unmodified and multiple modified states by mass spectrometry, suggesting the importance of this region to LipL32. Phosphorylated Tyr170, mono-, di- and tri-methylated Lys171 were identified during the individual database searching. A multi-modifications searching was carried out and the doubly modified peptide 160LDDDDDGDDTpYme3KEER174 (me3 denotes trimethyl-) was found. Additionally, we found methylation at the Glu250 residue in the C-terminus of LipL32. These PTM sites are localized in the regions containing two electronegative patches (dominated by Glu250, Glu251, Glu260 and Glu261, and by Glu138, Asp167 and Asp168) revealed by the crystal structure of LipL32 determined in the recent study 39. These two acid-rich patches were identified as potential binding sites for extracellular matrix (ECM) proteins such as laminin. These PTM events may influence the interaction between LipL32 and ECM proteins due to increasing (phosphorylation) electronegative charges or due to steric repulsions. Moreover, Tyr170 and Lys171 are localized in the human antibody epitope of LipL32 (residues 151-177). Therefore, studying the biological significances of these PTM events will provide insights into further understanding the immunological role of LipL32.
Evolutionary conservation analysis of the modified proteins
Previous conservation analysis indicated that phosphoproteins were more conserved than the nonphosphorylated proteins 24, 25. It should be noted that in these studies, the predicted nonphosphorylated proteins in the databases were used to compare with the MS-detected phosphoproteins. We reason that such comparisons should ideally be made among really expressed proteins. Although the full protein expression profile is hard to obtain, we reason that in L. interrogans, we have established a relatively comprehensive global protein expression profile that was obtained simultaneously along with the profile of modified proteins. Therefore, it may be more reasonable to draw conservation comparisons between MS-detected modified proteins and MS-detected unmodified proteins. By this methodology, we found that neither the acetylproteins nor the Glx-methylated proteins showed a pronounced enhancement in conservation compared to the MS-detected unmodified proteins and only Lys/Arg-methylated proteins appear to be a little more conserved throughout bacteria and eukaryotes. Phosphoproteins showed even poorer conservation than the MS-detected unmodified proteins. Recent studies also indicate that many phosphorylation and acetylation events may have been acquired during evolution and are considered as species-specific 40, 41. The phosphoproteins and acetylproteins in L. interrogans are significantly relevant to signal transduction and transcription regulation, which are involved in the adaptive response and gene expression. Therefore, we propose that the relatively unique lifestyles of this pathogen may partially explain the lack of more conservation of phosphoproteins and acetylproteins on average. Different from phosphoproteins and acetylproteins, quite a few of the Lys/Arg-methylated proteins are involved in the conserved function class and translation machinery, and therefore show more conservation from bacteria to eukaryotes.
Comparison of PTM patterns of L. interrogans with other species
Phosphorylation and acetylation have been systematically studied in E. coli. Compared to the identified modification sites and proteins in E. coli, those in L. interrogans represent different modification features. Although the titania enrichment technique was used to capture phosphopeptides and the obtained phosphopeptides were combined with those obtained by the Yin-yang MDLC technique, the identified phosphoproteins were much less than those identified in E. coli (Table 2). Meanwhile, there are differences in phosphorylation features between L. interrogans and E. coli. For example, the phosphoproteins in E. coli are overrepresented in carbohydrate metabolism, while those in L. interrogans are overrepresented in signal transduction. These differences may be partly explained by their carbon sources in culture. E. coli may use sugars as their carbon sources, and glycolysis is the primary central metabolism for this bacterium. It is well known that phosphorylation positively regulates glycolysis and the enzymes involved in glycolysis are of high abundance, therefore, a high percentage of the phosphosites in the glycolysis enzymes may be detected in E. coli. In contrast, L. interrogans utilizes long-chain fatty acids as the sole carbon source due to the lack of hexokinase, the key enzyme for the glycolysis 3. Since glycolysis does not exist in L. interrogans, it is not unusual that we detected few phosphosites in the proteins involved in carbohydrate metabolism. Another potential reason why fewer phosphosites were identified in L. interrogans may be due to the presence of more phosphatases in L. interrogans (25) compared to E. coli (5), which may lead to more frequent dephosphorylation in L. interrogans. In addition, the significantly overrepresented eukaryotic-like consensus sequences were not found in E. coli and B. subtilis. By contrast, more than a half of the unambiguous phosphosites in L. interrogans matched the target motifs of eukaryotic kinases, and genomic analysis indicates that L. interrogans contains more eukaryotic-like kinases and eukaryotic-like phosphatases than E. coli and B. subtilis. These findings at the proteomic and genomic levels indicate that L. interrogans contains eukaryotic-like Ser/Thr phosphorylation machinery. Research on the human pathogen Mycobacterium tuberculosis has revealed that Ser/Thr phosphorylation regulated by eukaryotic-like Ser/Thr kinases and phosphatases plays crucial roles in physiology and virulence 19, 42. In L. interrogans, some catalytic enzymes involved in Ser/Thr phosphorylation only exist in pathogenic Leptospira species (e.g., Ser/Thr kinases, LA1164 and LA3113). Therefore, this raises the possibility that the eukaryotic-like phosphorylation system is partly linked to virulence of L. interrogans. Further studies of the Ser/Thr phosphorylation system will deepen our understanding of the physiology and pathogenesis of L. interrogans. Like phosphorylation, there are also differences in the acetylation systems between L. interrogans and E. coli, including the consensus sequences flanking the acetylation sites and the overrepresented functional classes 26.
Previous studies in E. coli and Salmonella enterica have proposed a Glx methylation consensus sequence, Glx-Glx-X-X-A-S/T, and methylation occurred strictly at the second Glx residue 43. Another consensus sequence, A/S-sm-X-Glx-Glx-X-sm-A/S was found in an evolutionarily primitive bacterium Thermotoga maritime and the Glx-Glx methylation was prone to occur at the first Glx residue 44. In L. interrogans, the Glx methylation showed similar features of target sequences and functions with that in E. coli and that in T. maritime, suggesting that this modification is conserved in bacteria from evolutionarily primitive to evolutionarily advanced bacteria. It is noted that Glx methylation in L. interrogans showed more features of the target sequences in T. maritime rather than those in E. coli. For example, the Glx-Glx methylation in L. interrogans occurred preferentially at the first Glx residue and small amino acids at the −2 and +2 positions of the Glx-Glx pair were overrepresented. These similarities and differences may be associated with their evolution states. L. interrogans and T. maritime are both evolutionarily primitive and thus probably have more similarities in Glx methylation.
In summary, we present here a relatively reliable genome annotation for the human pathogen L. interrogans serovar Lai by utilizing proteomic data in combination with computational prediction. Meanwhile, the combined map of global protein expression and multiple PTMs will help us to better understand the special status of L. interrogans during evolution as a mammalian pathogen and provide additional views for further studies of the physiology and pathogenesis of L. interrogans. By mass spectrometry, quite a few Arg methylation sites were found. In combination with Ser/Thr phosphorylation features, this supports the suggestion that there are eukaryotic-like PTM machineries in L. interrogans, which may serve as potential therapeutic targets for leptospirosis.
Materials and Methods
Genome annotation
The genome sequence of L. interrogans serovar Lai strain Lai was downloaded from GenBank (http://www.ncbi.nlm.nih.gov/). Gene prediction tools, Glimmer 2.13 33 and ORPHUES 34 were used to predict genes. The protein coding sequences (CDSs) predicted by Glimmer and ORPHUES were searched against nr (NCBI non-redundant) database for functional annotation using BLAST and were searched against the Pfam, PRINTS, ProDom, Block and SMART databases for domain information using InterProScan 45. A six-frame translation of the entire genome of L. interrogans was carried out, which comprised 70 903 candidate CDSs (longer than 20 codons). The target-decoy database 28 consisting of forward and reversed sequences of these CDSs was constructed and subjected to MS/MS spectra searching.
Cell culture and protein preparation
The culture of L. interrogans serovar Lai type strain Lai (56601) was prepared as described previously 46. Briefly, cells were cultured to mid-log phase (at a density of ∼6.6 × 108 bacteria per ml) in liquid Ellinghausen-McCullough-Johnson-Harris (EMJH) medium at 28 °C with shaking under aerobic conditions; the cells were then harvested by centrifugation at 10 000× g for 10 min at 4 °C, followed by washing thrice in phosphate-buffered saline. The cell pellets were resuspended in the lysis buffer consisting of 2% SDS, 50 mM Tris-HCl (pH 8.0), 2 mM PMSF, 2 mM sodium fluoride and 2 mM sodium orthovanadate, sonicated, and then centrifuged at 25 000× g for 1 h. The concentration of protein extracts was determined by the bicinchoninic acid assay. On average, ∼1 mg of proteins could be obtained from 1.0 × 1010 bacteria. The proteins were reduced with 10 mM Dithiothreitol for 2 h at 37 °C, and carbamidomethylated with 50 mM iodoacetamide for 45 min at room temperature in darkness. Subsequently, the solution was incubated with four volumes of pre-cold acetone overnight at −20 °C and centrifuged at 25 000× g for 1 h to remove the supernatant.
In-solution digestion
The precipitated proteins were resuspended in 50 mM ammonium bicarbonate (pH 8.3) buffer and incubated with sequencing-grade modified trypsin (Promega) (1:50) with shaking for 4 h at 37 °C. Then, trypsin was added again to make the final protease/protein ratio up to 1:25. After 16 h, the digestion solution was ultrafiltered using 10 kDa Microcon Centrifugal Filter Devices (Millipore) to remove trypsin, and then the sample was lyophilized.
Yin-yang MDLC-MS/MS analysis
The Yin-yang MDLC system was performed as described previously with some modifications including using pH continuous gradient elution instead of pH step gradient elution and using SAX as the first loading 27, 47. Briefly, ∼1 mg of the sample was dissolved in 100 μl of pH 2.0 buffer (2 mM citric acid adjusted by formic acid), and then loaded onto the SCX column (10 μm, 320 μm × 100 mm, Column Technology Inc, CA, USA) by a syringe pump at a flow rate of 3 μl/min. The flow-through fraction of the SCX column was lyophilized and dissolved in 100 μl of pH 8.5 buffer (NH4OH and formic acid), then loaded onto the SAX column (10 μm, 320 μm × 100 mm, Column Technology Inc). Meanwhile, in order to better stabilize phosphohistidine and phosphoaspartate, we reversed the order of SCX and SAX by using SAX as the first loading. The sample was dissolved in 100 μl of pH 8.5 buffer, and then loaded first onto the SAX column by a syringe pump at a flow rate of 3 μl/min. The rest of the steps were performed as described above. The SCX/SAX column was coupled with a Surveyor liquid chromatography system (Thermo Fisher Scientific), consisting of a degasser, two MS pumps, an autosampler, two C18 trap columns (5 μm, 300 μm × 5 mm, Agilent Technologies) and an analytical C18 column (5 μm, 75 μm × 150 mm, Column Technology Inc) on-line. The HPLC solvents used were 0.1% formic acid (v/v) aqueous (A) and 0.1% formic acid (v/v) acetonitrile (ACN) (B). The sequential elution from the SCX column was by pH continuous gradient buffer, which was from pH 2.0 to pH 8.5 (from pH 8.5 to pH 2.0 in SAX) instead of previously reported pH step gradient buffer 27, as described in our recent work 47. Each of the 10 eluted fractions was on-line concentrated and desalted on the C18 trap column at a flow rate of 3 μl/min after the split, and then subjected to the analytical C18 column. The reverse-phase gradient was from 2% to 40% of the mobile phase B in 165 min at a flow rate of 100 μl/min before the split and 250 nl/min after the split. A LTQ-Orbitrap mass spectrometer (Thermo Fisher Scientific) equipped with a nanospray source was used in the MS/MS experiment with ion transfer capillary at 160 °C and NSI voltage of 1.8 kV. Normalized collision energy was 35.0%. Dynamic exclusion settings included: repeat count 2, repeat duration 30 s and exclusion duration 90 s. Full scan was performed in the Orbitrap analyzer (R = 100 000 at m/z 400) followed by MS/MS performed by CID (collision-induced dissociation) detected in the linear ion trap.
Phosphopeptide enrichment by titania
The phosphopeptide enrichment by titania was according to Wu et al. 48. Approximately 4 mg of the tryptic peptide mixture were dissolved in 500 μl of the sample loading buffer (2% TFA/65% ACN solution saturated with glutamic acid) and were incubated with 8 mg of titania beads (5 μm, GL Sciences, Japan). The titania beads were sequentially washed with 800 μl 0.5% TFA/65% ACN and 0.1% TFA/65% ACN. The bound peptides were sequentially eluted with 200 μl of 300 mM NH4OH/50% ACN and 500 mM NH4OH/60% ACN. The eluted solutions were combined and were lyophilized for 1D-RP-LC-MS/MS analysis.
Alkaline phosphatase treatment, Pro-Q staining and in-gel digestion
The precipitated proteins (∼100 μg) were dissolved in the lysis buffer consisting of 6 M urea and 100 mM Tris-HCl (pH 8.8), and were split into two aliquots, diluted to a concentration of 1 M urea and incubated with 50 U alkaline phosphatase (P0114, Sigma) and nothing for 2 h at 37 °C, respectively. Subsequently 50 μg of these two aliquots of samples were separated by a 7 cm SDS-PAGE (12.5%) minigel. The gel was fixed by methanol (50%)-acetic acid (10%) for 30 min twice, washed by water for 10 min thrice, then stained by Pro-Q diamond dye (Invitrogen) for 60 min, destained by the buffer consisting 20% acetonitrile and 50 mM sodium acetate (pH 4.0) for 30 min thrice and washed by water for 10 min twice. After visualizing on LAS-4000 (Fujifilm), the gel was stained by Coomassie brilliant blue. The gel bands containing LipL32 were excised and were in-gel digested by trypsin, as described previously 49.
1D-RP-LC-MS/MS analysis
The reverse-phase gradient was from 2% to 40% of the mobile phase B in 75 min at a flow rate of 100 μl/min before the split and 250 nl/min after the split. The MS/MS parameters were set as described in the section of the Yin-yang MDLC-MS/MS analysis. In particular, multistage activation was enabled in the MS/MS events from the sample of the titania enrichment to improve fragmentation spectra of phosphopeptides 50.
Data analysis and validation
The acquired MS/MS spectra were searched against the target-decoy databases consisting of forward and reversed sequences of CDSs in the six-frame translation database and the eventually completed CDS database using the TurboSEQUEST program in the BioWorks 3.2 software package. In particular, for searching PTMs the MS/MS spectra were searched against the target-decoy database of the eventually completed CDS database individually four times with different dynamic modifications and trypsin missed sites of: (1) phosphorylation of Ser/Thr/Tyr/His/Asp, 2 missed; (2) acetylation of Lys, 5 missed; (3) deamidation of Gln, deamidation and methylation of Gln, and methylation of Glu, 2 missed; (4) monomethylation of Lys/Arg, dimethylation of Lys/Arg, trimethylation of Lys, 5 missed. Other identical search criteria were as follows: fully tryptic specificity; carbamidomethylation of cysteine was set as a fixed modification; oxidation of methionine was set as a dynamic modification; the precursor and fragment ion mass tolerance was 500 p.p.m. and 1.0 Da (default), respectively. As a supplement, we also searched MS/MS spectra by using the above parameters including all the modifications except for those occurring on Gln. The precursor ion mass accuracy of 10 p.p.m. and the 1.0% FPR were selected to filter the identified peptides. The FPR was calculated based on the following formula: % fal = 2(nrev/(nrev+nreal)), where the % fal is the estimated false-positive rate, the nrev is the number of peptide hits from the decoy database, and nreal is the number of peptide hits from target database 28. The TurboSEQUEST results from four PTM searching were combined together to remove the different peptide identifications from the same scan using our in-house software, BuildSummary 51. All the MS/MS spectra of modified peptides and unique peptides singly assigned to the corresponding proteins were manually checked. The phosphoserine- and phosphothreonine-containing peptides were expected to show a pronounced neutral loss of phosphoric acid from the precursor ion or fragment ions. The proline-containing peptides were expected to show a pronounced cleavage N-terminal of the proline residue. For modified peptides with multiple potential modified sites, the probability score, Ascore, was calculated as described previously and those sites with an Ascore ≥ 19 were annotated as modified sites, otherwise as ambiguous sites 52. The quantitative analysis software tool, Census, was used to extract and compare the peptide intensities of proteins before and after alkaline phosphatase treating 53.
Bioinformatics analysis
BLASTP of proteins against KEGG (Kyoto Encyclopedia of Genes and Genomes) database (http://www.genome.jp/kegg/) was used to obtain the pathway information. COG (Clusters of Orthologous Genes) descriptions in NCBI were used to acquire the function classification of proteins 54. Orthologs of L. interrogans proteins across 132 species from firmicutes to human were determined via two-directional BLASTP. First, homology search was performed in the protein databases of 113 bacteria, 10 archaea and 9 eukaryotes from GenBank. To eliminate possible influences of the genome size on protein conservation analysis, 49 bacterial species with more than 3 500 predicted CDSs were eventually selected from 113 bacteria to analyze protein conservation. The measurement of protein evolutionary conservation was performed as described previously 25. The enrichment analysis for sequence motifs and protein function classes was performed using hypergeometric test with correction for multiple hypothesis testing. Because more than 64.0% of the predicted proteins were identified in our study and the corrected probability P-values were calculated to be a little larger in all enrichment analysis if the data set of predicted proteins in L. interrogans database was used as the reference data set, we presumed that it was more reasonable to select the data set of identified total proteins as the reference data set. Therefore, we eventually used the data set of identified total proteins instead of the data set of predicted proteins as the reference data set in all enrichment analysis described in this work. The sequence motifs and protein function classes that were significant with hypergeometric P < 0.05 were selected as overrepresented.
( Supplementary information is linked to the online version of the paper on the Cell Research website.)
References
Levett PN . Leptospirosis. Clin Microbiol Rev 2001; 14:296–326.
WHO. Leptospirosis worldwide, 1999. Wkly Epidemiol Rec 1999; 74:237–242.
Ren SX, Fu G, Jiang XG, et al. Unique physiological and pathogenic features of Leptospira interrogans revealed by whole-genome sequencing. Nature 2003; 422:888–893.
Nascimento AL, Ko AI, Martins EA, et al. Comparative genomics of two Leptospira interrogans serovars reveals novel insights into physiology and pathogenesis. J Bacteriol 2004; 186:9.
Bulach DM, Zuerner RL, Wilson P, et al. Genome reduction in Leptospira borgpetersenii reflects limited transmission potential. Proc Natl Acad Sci USA 2006; 103:14560–14565.
Picardeau M, Bulach DM, Bouchier C, et al. Genome sequence of the saprophyte Leptospira biflexa provides insights into the evolution of Leptospira and the pathogenesis of Leptospirosis. PLoS ONE 2008; 3:e1607.
Qin JH, Sheng YY, Zhang ZM, et al. Genome-wide transcriptional analysis of temperature shift in L. interrogans serovar Lai strain 56601. BMC Microbiol 2006; 6:51.
Lo M, Bulach DM, Powell DR, et al. Effects of temperature on gene expression patterns in Leptospira interrogans serovar Lai as assessed by whole-genome microarrays. Infect Immun 2006; 74:5848–5859.
Cullen PA, Cordwell SJ, Bulach DM, Haake DA, Adler B . Global analysis of outer membrane proteins from Leptospira interrogans serovar Lai. Infect Immun 2002; 70:2311–2318.
Nally JE, Whitelegge JP, Aguilera R, et al. Purification and proteomic analysis of outer membrane vesicles from a clinical isolate of Leptospira interrogans serovar Copenhageni. Proteomics 2005; 5:144–152.
Nally JE, Whitelegge JP, Bassilian S, Blanco DR, Lovett MA . Characterization of the outer membrane proteome of Leptospira interrogans expressed during acute lethal infection. Infect Immun 2007; 75:766–773.
Guerreiro H, Croda J, Flannery B, et al. Leptospiral proteins recognized during the humoral immune response to leptospirosis in humans. Infect Immun 2001; 69:4958–4968.
Sakolvaree Y, Maneewatch S, Jiemsup S, et al. Proteome and immunome of pathogenic Leptospira spp. revealed by 2DE and 2DE-immunoblotting with immune serum. Asian Pac J Allergy Immunol 2007; 25:53–73.
Malmstrom J, Beck M, Schmidt A, et al. Proteome-wide cellular protein concentrations of the human pathogen Leptospira interrogans. Nature 2009; 460:762–765.
Ishino Y, Okada H, Ikeuchi M, Taniguchi H . Mass spectrometry-based prokaryote gene annotation. Proteomics 2007; 7:4053–4065.
de Souza GA, Malen H, Softeland T, et al. High accuracy mass spectrometry analysis as a tool to verify and improve gene annotation using Mycobacterium tuberculosis as an example. BMC Genomics 2008; 9:316.
Tanner S, Shen Z, Ng J, et al. Improving gene annotation using peptide mass spectrometry. Genome Res 2007; 17:231–239.
Polevoda B, Sherman F . Methylation of proteins involved in translation. Mol Microbiol 2007; 65:590–606.
Wehenkel A, Bellinzoni M, Grana M, et al. Mycobacterial Ser/Thr protein kinases and phosphatases: physiological roles and therapeutic potential. Biochim Biophys Acta 2008; 1784:193–202.
Bendt AK, Burkovski A, Schaffer S, et al. Towards a phosphoproteome map of Corynebacterium glutamicum. Proteomics 2003; 3:1637–1646.
Pawson T, Scott JD . Protein phosphorylation in signaling--50 years and counting. Trends Biochem Sci 2005; 30:286–290.
Paik WK, Paik DC, Kim S . Historical review: the field of protein methylation. Trends Biochem Sci 2007; 32:146–152.
Bayle JH, Crabtree GR . Protein acetylation: more than chromatin modification to regulate transcription. Chem Biol 1997; 4:885–888.
Macek B, Mijakovic I, Olsen JV, et al. The serine/threonine/tyrosine phosphoproteome of the model bacterium Bacillus subtilis. Mol Cell Proteomics 2007; 6:697–707.
Macek B, Gnad F, Soufi B, et al. Phosphoproteome analysis of E. coli reveals evolutionary conservation of bacterial Ser/Thr/Tyr phosphorylation. Mol Cell Proteomics 2008; 7:299–307.
Zhang J, Sprung R, Pei J, et al. Lysine acetylation is a highly abundant and evolutionarily conserved modification in Escherichia coli. Mol Cell Proteomics 2009; 8:215–225.
Dai J, Jin WH, Sheng QH, et al. Protein phosphorylation and expression profiling by Yin-yang multidimensional liquid chromatography (Yin-yang MDLC) mass spectrometry. J Proteome Res 2007; 6:250–262.
Peng J, Elias JE, Thoreen CC, Licklider LJ, Gygi SP . Evaluation of multidimensional chromatography coupled with tandem mass spectrometry (LC/LC-MS/MS) for large-scale protein analysis: the yeast proteome. J Proteome Res 2003; 2:43–50.
MacCoss MJ, McDonald WH, Saraf A, et al. Shotgun identification of protein modifications from protein complexes and lens tissue. Proc Natl Acad Sci USA 2002; 99:7900–7905.
Jaffe JD, Stange-Thomann N, Smith C, et al. The complete genome and proteome of Mycoplasma mobile. Genome Res 2004; 14:1447–1461.
McBride AE, Silver PA . State of the arg: protein methylation at arginine comes of age. Cell 2001; 106:5–8.
Tatusov RL, Galperin MY, Natale DA, Koonin EV . The COG database: a tool for genome-scale analysis of protein functions and evolution. Nucleic Acids Res 2000; 28:33–36.
Delcher AL, Harmon D, Kasif S, White O, Salzberg SL . Improved microbial gene identification with GLIMMER. Nucleic Acids Res 1999; 27:4636–4641.
Frishman D, Mironov A, Mewes HW, Gelfand M . Combining diverse evidence for gene recognition in completely sequenced bacterial genomes. Nucleic Acids Res 1998; 26:2941–2947.
Haldenwang WG . The sigma factors of Bacillus subtilis. Microbiol Rev 1995; 59:1–30.
Min KT, Hilditch CM, Diederich B, Errington J, Yudkin MD . Sigma F, the first compartment-specific transcription factor of B. subtilis, is regulated by an anti-sigma factor that is also a protein kinase. Cell 1993; 74:735–742.
Ramakrishnan R, Schuster M, Bourret RB . Acetylation at Lys-92 enhances signaling by the chemotaxis response regulator protein CheY. Proc Natl Acad Sci USA 1998; 95:4918–4923.
Cullen PA, Haake DA, Adler B . Outer membrane proteins of pathogenic spirochetes. FEMS Microbiol Rev 2004; 28:291–318.
Vivian JP, Beddoe T, McAlister AD, et al. Crystal structure of LipL32, the most abundant surface protein of pathogenic Leptospira spp. J Mol Biol 2009; 387:1229–1238.
Soufi B, Gnad F, Jensen PR, et al. The Ser/Thr/Tyr phosphoproteome of Lactococcus lactis IL1403 reveals multiply phosphorylated proteins. Proteomics 2008; 8:3486–3493.
Yang XJ, Seto E . Lysine acetylation: codified crosstalk with other posttranslational modifications. Mol Cell 2008; 31:449–461.
Greenstein AE, MacGurn JA, Baer CE, et al. M. tuberculosis Ser/Thr protein kinase D phosphorylates an anti-anti-sigma factor homolog. PLoS Pathog 2007; 3:e49.
Kehry MR, Bond MW, Hunkapiller MW, Dahlquist FW . Enzymatic deamidation of methyl-accepting chemotaxis proteins in Escherichia coli catalyzed by the cheB gene product. Proc Natl Acad Sci USA 1983; 80:3599–3603.
Perez E, Zheng H, Stock AM . Identification of methylation sites in Thermotoga maritima chemotaxis receptors. J Bacteriol 2006; 188:4093–4100.
Hunter S, Apweiler R, Attwood TK, et al. InterPro: the integrative protein signature database. Nucleic Acids Res 2009; 37 (Database issue):D211–D215.
Yang HL, Zhu YZ, Qin JH, et al. In silico and microarray-based genomic approaches to identifying potential vaccine candidates against Leptospira interrogans. BMC Genomics 2006; 7:293.
Dai J, Wang LS, Wu YB, et al. Fully automatic separation and identification of phosphopeptides by continuous pH-gradient anion exchange online coupled with reversed-phase liquid chromatography mass spectrometry. J Proteome Res 2009; 8:133–141.
Wu J, Shakey Q, Liu W, Schuller A, Follettie MT . Global profiling of phosphopeptides by titania affinity enrichment. J Proteome Res 2007; 6:4684–4689.
Jiang XS, Tang LY, Cao XJ, et al. Two-dimensional gel electrophoresis maps of the proteome and phosphoproteome of primitively cultured rat mesangial cells. Electrophoresis 2005; 26:4540–4562.
Schroeder MJ, Shabanowitz J, Schwartz JC, Hunt DF, Coon JJ . A neutral loss activation method for improved phosphopeptide sequence analysis by quadrupole ion trap mass spectrometry. Anal Chem 2004; 76:3590–3598.
Dai J, Shieh CH, Sheng QH, Zhou H, Zeng R . Proteomic analysis with integrated multiple dimensional liquid chromatography/mass spectrometry based on elution of ion exchange column using pH steps. Anal Chem 2005; 77:5793–5799.
Beausoleil SA, Villen J, Gerber SA, Rush J, Gygi SP . A probability-based approach for high-throughput protein phosphorylation analysis and site localization. Nat Biotechnol 2006; 24:1285–1292.
Park SK, Venable JD, Xu T, Yates JR 3rd . A quantitative analysis software tool for mass spectrometry-based proteomics. Nat Methods 2008; 5:319–322.
Tatusov RL, Fedorova ND, Jackson JD, et al. The COG database: an updated version includes eukaryotes. BMC Bioinformatics 2003; 4:41.
Acknowledgements
This work was supported by the National Natural Science Foundation of China (30425021, 30521005, 30670102, 30770111, 30770820), the Basic Research Foundation (2006CB910700), the CAS Project (KSCX2-YW-R-106, KSCX1-YW-02), the High-technology Project (2007AA02Z334) and the National High Technology Research and Development Program of China (2006AA02Z176).
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary information, Table S1
Ten MS-detected proteins that were missed in computational prediction. The chromosomal location includes the stop codon. (XLS 21 kb)
Supplementary information, Table S2
MS-detected proteins of L. interrogans strain Lai in our study. (XLS 564 kb)
Supplementary information, Table S4
Evolutionary conservation of 3 953 predicted proteins of L. interrogans strain Lai. The 49 bacterial, ten archaeal and ninw eukaryotic species listed in the row 1 are used to perform the evolutionary analysis. “2”, “1” and “0” denote two-dimensional homolog, one-dimensional homolog and no homolog respectively. (XLS 6556 kb)
Supplementary information, Figure S1A, Figure S1B, Figure S1C, Figure S1D
Phosphopeptide spectra (Page 2–35), Acetylpeptide spectra(Page 36–82), Glx-methylated peptide spectra(Page 83–203), K/R-methylated peptide spectra (Page 204–273) (PDF 53508 kb)
Supplementary information, Figure S2
Pro-Q staining and Coomassie Brilliant Blue (CBB) staining of the protein lysates of L. interrogans serovar Lai before (AP−) and after (AP+) alkaline phosphatase treating. (PDF 203 kb)
Rights and permissions
About this article
Cite this article
Cao, XJ., Dai, J., Xu, H. et al. High-coverage proteome analysis reveals the first insight of protein modification systems in the pathogenic spirochete Leptospira interrogans. Cell Res 20, 197–210 (2010). https://doi.org/10.1038/cr.2009.127
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cr.2009.127
Keywords
This article is cited by
-
Deciphering the lysine acetylation pattern of leptospiral strains by in silico approach
Network Modeling Analysis in Health Informatics and Bioinformatics (2023)
-
Disentangling the Impact of Sulfur Limitation on Exopolysaccharide and Functionality of Alr2882 by In Silico Approaches in Anabaena sp. PCC 7120
Applied Biochemistry and Biotechnology (2021)
-
Hypothesis: protein and RNA attributes are continuously optimized over time
BMC Genomics (2019)
-
Virulence of the zoonotic agent of leptospirosis: still terra incognita?
Nature Reviews Microbiology (2017)