Abstract
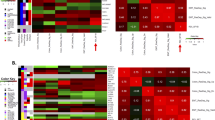
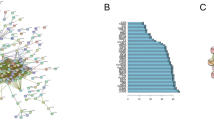
Identification of the genes that are differentially expressed between radiosensitive and radioresistant cancers by global gene analysis may help to elucidate the mechanisms underlying tumor radioresistance and improve the efficacy of radiotherapy. An integrated analysis was conducted using publicly available GEO datasets to detect differentially expressed genes (DEGs) between cancer cells exhibiting radioresistance and cancer cells exhibiting radiosensitivity. Gene Ontology (GO) enrichment analyses, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis and protein–protein interaction (PPI) networks analysis were also performed. Five GEO datasets including 16 samples of radiosensitive cancers and radioresistant cancers were obtained. A total of 688 DEGs across these studies were identified, of which 374 were upregulated and 314 were downregulated in radioresistant cancer cell. The most significantly enriched GO terms were regulation of transcription, DNA-dependent (GO: 0006355, P=7.00E-09) for biological processes, while those for molecular functions was protein binding (GO: 0005515, P=1.01E-28), and those for cellular component was cytoplasm (GO: 0005737, P=2.81E-26). The most significantly enriched pathway in our KEGG analysis was Pathways in cancer (P=4.20E-07). PPI network analysis showed that IFIH1 (Degree=33) was selected as the most significant hub protein. This integrated analysis may help to predict responses to radiotherapy and may also provide insights into the development of individualized therapies and novel therapeutic targets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Delaney G, Jacob S, Featherstone C, Barton M . The role of radiotherapy in cancer treatment: estimating optimal utilization from a review of evidence-based clinical guidelines. Cancer 2005; 104: 1129–1137.
Kristensen CA, Kjaer-Kristoffersen F, Sapru W, Berthelsen AK, Loft A, Specht L . Nasopharyngeal carcinoma. Treatment planning with IMRT and 3D conformal radiotherapy. Acta Oncol 2007; 46: 214–220.
Jeggo P, Lobrich M . Radiation-induced DNA damage responses. Radiat Prot Dosimetry 2006; 122: 124–127.
Vaupel P . Tumor microenvironmental physiology and its implications for radiation oncology. Semin Radiat Oncol 2004; 14: 198–206.
Ogawa K, Murayama S, Mori M . Predicting the tumor response to radiotherapy using microarray analysis (Review). Oncol Rep 2007; 18: 1243–1248.
Hanahan D, Weinberg RA . The hallmarks of cancer. Cell 2000; 100: 57–70.
Rodemann HP, Dittmann K, Toulany M . Radiation-induced EGFR-signaling and control of DNA-damage repair. Int J Radiat Biol 2007; 83: 781–791.
Hehlgans S, Cordes N . Caveolin-1: an essential modulator of cancer cell radio-and chemoresistance. Am J Cancer Res 2011; 1: 521–530.
Lee NH, Saeed AI . Microarrays: an overview. Methods Mol Biol 2007; 353: 265–300.
Wong YF, Sahota DS, Cheung TH, Lo KW, Yim SF, Chung TK et al. Gene expression pattern associated with radiotherapy sensitivity in cervical cancer. Cancer J 2006; 12: 189–193.
Ogawa K, Utsunomiya T, Mimori K, Tanaka F, Haraguchi N, Inoue H et al. Differential gene expression profiles of radioresistant pancreatic cancer cell lines established by fractionated irradiation. Int J Oncol 2006; 28: 705–713.
Higo M, Uzawa K, Kouzu Y, Bukawa H, Nimura Y, Seki N et al. Identification of candidate radioresistant genes in human squamous cell carcinoma cells through gene expression analysis using DNA microarrays. Oncol Rep 2005; 14: 1293–1298.
Guo WF, Lin RX, Huang J, Zhou Z, Yang J, Guo GZ et al. Identification of differentially expressed genes contributing to radioresistance in lung cancer cells using microarray analysis. Radiat Res 2005; 164: 27–35.
Kim HS, Kim SC, Kim SJ, Park CH, Jeung HC, Kim YB et al. Identification of a radiosensitivity signature using integrative metaanalysis of published microarray data for NCI-60 cancer cells. BMC Genomics 2012; 13: 348.
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M et al. NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res 2013; 41 (D1): D991–D995.
Tusher VG, Tibshirani R, Chu G . Significance analysis of microarrays applied to the ionizing radiation response. Proc Natl Acad Sci USA 2001; 98: 5116–5121.
Tabas-Madrid D, Nogales-Cadenas R, Pascual-Montano A . GeneCodis3: a non-redundant and modular enrichment analysis tool for functional genomics. Nucleic Acids Res 2012; 40 (Web Server issue): W478–W483.
Altermann E, Klaenhammer TR . PathwayVoyager: pathway mapping using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database. BMC Genomics 2005; 6: 60.
Giot L, Bader JS, Brouwer C, Chaudhuri A, Kuang B, Li Y et al. A protein interaction map of Drosophila melanogaster. Science 2003; 302: 1727–1736.
Li S, Armstrong CM, Bertin N, Ge H, Milstein S, Boxem M et al. A map of the interactome network of the metazoan C. elegans. Science 2004; 303: 540–543.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res 2003; 13: 2498–2504.
Lee JC, Lee WH, Min YJ, Cha HJ, Han MW, Chang HW et al. Development of TRAIL resistance by radiation-induced hypermethylation of DR4 CpG island in recurrent laryngeal squamous cell carcinoma. Int J Radiat Oncol Biol Phys 2014; 88: 1203–1211.
Bergman PJ, Harris D . Radioresistance, chemoresistance, and apoptosis resistance. The past, present, and future. Vet Clin North Am Small Anim Pract 1997; 27: 47–57.
Marin-Aguilera M, Codony-Servat J, Kalko SG, Fernandez PL, Bermudo R, Buxo E et al. Identification of docetaxel resistance genes in castration-resistant prostate cancer. Mol Cancer Ther 2012; 11: 329–339.
Yasui K, Mihara S, Zhao C, Okamoto H, Saito-Ohara F, Tomida A et al. Alteration in copy numbers of genes as a mechanism for acquired drug resistance. Cancer Res 2004; 64: 1403–1410.
Huang Z, Li H, Huang Q, Chen D, Han J, Wang L et al. SERPINB2 down-regulation contributes to chemoresistance in head and neck cancer. Mol Carcinog 2013; 53: 777–786.
Zhang YW, Zheng Y, Wang JZ, Lu XX, Wang Z, Chen LB et al. Integrated analysis of DNA methylation and mRNA expression profiling reveals candidate genes associated with cisplatin resistance in non-small cell lung cancer. Epigenetics 2014; 9: 896–909.
Sandfort V, Koch U, Cordes N . Cell adhesion-mediated radioresistance revisited. Int J Radiat Biol 2007; 83: 727–732.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Cancer Gene Therapy website
Supplementary information
Rights and permissions
About this article
Cite this article
Hou, DL., Chen, L., Liu, B. et al. Identification of common gene networks responsive to radiotherapy in human cancer cells. Cancer Gene Ther 21, 542–548 (2014). https://doi.org/10.1038/cgt.2014.62
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cgt.2014.62
This article is cited by
-
Thymidine phosphorylase induction by ionizing radiation antagonizes 5-fluorouracil resistance in human ductal pancreatic adenocarcinoma
Radiation and Environmental Biophysics (2022)
-
Elevated HDAC activity and altered histone phospho-acetylation confer acquired radio-resistant phenotype to breast cancer cells
Clinical Epigenetics (2020)
-
Identification of drought stress-responsive genes in rice (Oryza sativa) by meta-analysis of microarray data
Journal of Genetics (2020)
-
Effect of irradiation on cytokine secretion and nitric oxide production by inflammatory macrophages
Genes & Genomics (2016)