Abstract
Cytokine signaling is critical for proliferation, survival and differentiation of hematopoietic cell, and interleukin-3 (IL-3) is required for maintenance of many hematopoietic cell lines, such as BaF3. We have isolated apoptosis-resistant clones of BaF3 using retroviral insertional mutagenesis and the Xbp1 locus was identified as a retroviral integration site. Expression and splicing of the Xbp1 transcript was conserved in the resistant clone but was promptly disappeared on IL-3 withdrawal in parental BaF3. IL-3 stimulation of BaF3 cells enhanced Xbp1 promoter activity and induced phosphorylation of the endoplasmic reticulum stress sensor protein IRE1, resulting in the increase in Xbp1S that activates unfolded protein response. When downstream signaling from IL-3 was blocked by LY294002 and/or dn-Stat5, Xbp1 expression was downregulated and IRE1 phosphorylation was suppressed. Inhibition of IL-3 signaling as well as knockdown of Xbp1-induced apoptosis in BaF3 cells. In contrast, constitutive expression of Xbp1S protected BaF3 from apoptosis during IL-3 depletion. However, cell cycle arrest at the G1 stage was observed in BaF3 and myeloid differentiation was induced in IL-3-dependent 32Dcl3 cells. Expression of apoptosis-, cell cycle- and differentiation-related genes was modulated by Xbp1S expression. These results indicate that the proper transcriptional and splicing regulation of Xbp1 by IL-3 signaling is important in homeostasis of hematopoietic cells.
Similar content being viewed by others
Main
Cytokine signaling is important for regulating cell processes in hematopoiesis. Cytokines bind and activate distinct receptors, and downstream molecules sequentially mediate extracellular stimuli that are required for cellular survival, proliferation and/or differentiation.1 Interleukin (IL)-3 induces development of multi-lineage cell types including granulocytes, macrophages, erythroid cells and megakaryocytes, and it possesses abilities to support the cellular growth and survival of more immature cell types in vitro. IL-3 is recognized by its receptor consisting of α- and β-subunits, and the latter is common to granulocyte-macrophage colony-stimulating factor (GM-CSF) and IL-5 receptors.2, 3 IL-3 binding leads to hetero-dimerization and phosphorylation of the receptor, after which downstream molecules within the JAK/STAT, Ras/MAPK and PI3K pathways are activated.2, 3
IL-3 is secreted predominantly by T cells in vivo. Multiple lineages of hematopoietic cells as well as many leukemic cells undergo apoptosis and growth arrest in the absence of IL-3.4 The members of the Bcl2 gene family mediate the cell survival or anti-apoptosis function under IL-3.2, 5 In fact, Bcl2 and BclXL are rapidly induced by IL-3, which is mediated by Jak2 activation.6 The PI3K pathway also has a role in the anti-apoptotic function of IL-3 via Bcl2 and Bcl2 family member BAD.2, 5 IL-3 and GM-CSF signaling regulates BclXL through the activation of Stat5.7 Acquired genetic mutations related to the IL-3 signaling pathway have also been identified in human hematopoietic disorders. A gain-of-function mutation V617F of JAK2 in human myeloproliferative neoplasm induces constitutive phosphorylation of Stat5, resulting in enhanced proliferation and survival of hematopoietic precursors.8 Genetic alterations in IL-3 signaling are thus important for survival of malignant hematopoietic cells, and important molecules remain to be identified in order to understand the signaling network in the downstream of IL-3.
Retrovirus insertional mutagenesis (RIM) is a powerful tool to identify important genes responsible for growth advantage in clonal expansion.9 The technique was efficiently utilized to identify IL-3-independent phenotypes that could dissect the function of particular molecules.10 By using RIM, we have identified Xbp1, which encodes a transcription factor important for the unfolded protein response (UPR) and endoplasmic reticulum (ER) function11, 12 downstream from IL-3.
UPR is a unique signaling system associated with the ER membrane. Any conditions inducing protein misfolding in the ER induces UPR that attenuates translation of proteins entering the ER.13 UPR also activates transcription of genes involved in the degradation of nascent misfolded proteins via activation of IRE1, ATF4/ATF6 and Xbp1.13 UPR induces transcription of Xbp1 and phosphorylation of the ER transmembrane kinase IRE1 that initiates UPR-mediated mRNA splicing of Xbp1, resulting in the production of Xbp1S, an active form of a basic leucine zipper transcription factor.11, 12 In this study, we demonstrate that IL-3 stimulation regulates both transcription and splicing of Xbp1, and that Xbp1 unexpectedly affects hematopoietic cell apoptosis, growth and differentiation. These observations reveal a novel functional role of Xbp1 in hematopoiesis and leukemogenesis and possible relationship between the IL-3 signaling and UPR.
Results
Retroviral insertional mutagenesis identified Xbp1 as a survival factor in the IL-3-dependent BaF3 cell
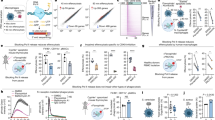
BaF3 cells (3.0 × 106) were infected with the MSCV-GFP-Neo retrovirus. The cells were incubated in the medium containing G-418 in the absence of IL-3, 48 h after infection, and were cultured for 4 weeks to select survival clone without IL-3. Three survival clones, B6F, B3G and B7B, were isolated. Southern blot analysis showed clonal integrations of the retrovirus (Figure 1a), and the retroviral integration sites (RIS) in each clone were identified using inverse polymerase chain reaction (IPCR) (Figure 1b). A RIS was identified in the B6F clone at 20 kb upstream of the Xbp1 gene on mouse chromosome 11 (Figure 1c), and the clone was subjected to further investigation. Tceb1 that encodes a 15 kDa subunit of transcription elongation factor B, Larp1, a ribonucleoprotein member gene and Ptprz1, a protein tyrosine phosphatase gene have been identified in the other two clones.
Retrovirus integration at the Xbp1 locus. (a) Southern blot analysis of survival BaF3 clones in IL-3 depletion. DNA was digested with BglII or EcoRV that cut the retrovirus once or at both long terminal repeats, respectively. The green fluorescent protein sequence was used as a probe. B6F and B3G contain a single integration and two in B7B. (b) Retrovirus integration sites of the survival clones. (c) A physical map of RIS at the Xbp1 locus
Modulation of Xbp1 expression and splicing by IL-3 signaling
To investigate relationship between IL-3 signaling and Xbp1 expression, Xbp1 alternative splicing that converts unstable Xbp1U to the stable form Xbp1S was analyzed by reverse transcription (RT)-PCR (Figure 2a). IL-3 depletion suppressed the expression of total Xbp1 and complete disappearance of Xbp1S was observed more rapidly than that of Xbp1U. On the other hand, B6F cells that have MSCV integration at the Xbp1 locus showed constitutive expression of the both Xbp1S and Xbp1U in the absence of IL-3 (Figure 2b).
IL-3 modulates Xbp1 expression and splicing. (a) An RT-PCR analysis of Xbp1. Both un-spliced and spliced forms decreased after withdrawal of IL-3. β-actin was used to confirm the quality and quantity of RNA. (b) Xbp1 expression is comparable between BaF3 with IL-3 and B6F without IL-3. (c and d) Upregulation of Xbp1S expression by IL-3 stimulation. BaF3 cells were cultured under the IL-3-free condition for 16 h. IL-3 was then added to the medium at 10 ng/ml and cells were harvested after incubation for the indicated periods. Whole cell extracts of BaF3 were subjected to RT-PCR (c) to detect Xbp1S and Xbp1U, and western blotting (d) to detect Xbp1S, phospho-IRE1, IRE1, cleaved ATF6, phospho-Stat5, GRP78 and GAPDH. Both Xbp1S upregulation and IRE1 phosphorylation was observed at 48 h after IL-3 stimulation. (e) Genomic DNA fragments encompassing the Xbp1 upstream region indicated were subcloned into the pGL3 basic vector. The constructs were nucleofected together with pRL-SV40 into 32Dcl3 and the cells were incubated for 24 h in growth medium containing IL-3. The cells were starved for IL-3 for 24 h and IL-3 was then added for additional 12 h. The cells were harvested at each time point and subjected to the luciferase assay. The bars represent the means of the relative luciferase activities, which were calculated by dividing the luciferase activity by the Renilla activity used as a transfection control. The means±S.D. from three independent experiments are shown (**P<0.05). (f) The 45 nt DNA fragment (28-bp upstream of the transcription start site of Xbp1) that contains ERSE (underlined) was subcloned into the pGL3 promoter vector. The A:G mutation at nt –59 was introduced and luciferase assays were carried out in 32Dcl3 cells in the presence of IL-3. (**P<0.05)
IL-3 stimulation induces Xbp1S expression and IRE1 phosphorylation
As expression and splicing of Xbp1 is regulated by the UPR stress response system or by the ER stress response,11, 12, 14 the effect of UPR signaling on IL-3 stimulation was investigated. Removal of a 26-nucleotide intron from Xbp1 mRNA induced by IRE1 phosphorylation was observed 48 h after addition of IL-3 to BaF3 cells (Figure 2c), which preceded accumulation of Xbp1S protein (Figure 2d). Modulation of IRE1 and Xbp1S production contrasted well with rapid phosphorylation of Stat5, a direct downstream target of IL-3 stimulation. Interestingly, expression of cleaved ATF6 was coincided with Xbp1S expression whereas GRP-78, another UPR marker,15 was not altered by IL-3 stimulation.
The ER stress response element (ERSR) within the Xbp1 promoter is responsible for the IL-3 response
As Xbp1 transcription was downregulated by IL-3 depletion, a reporter assay was carried out using IL-3-dependent 32Dcl3 murine myeloid cells to clarify whether the promoter activity of Xbp1 is modulated by IL-3 stimulation. The genomic region spanning 5 kb upstream of Xbp1 exon 1 was cloned into the pGL3 basic vector. The clone's promoter activity was 50% repressed by IL-3 deprivation or twofold activated by IL-3 (Figure 2e). An analysis of deletion mutants revealed that the minimal promoter activity is present within a 318-bp region upstream from exon 1. A previous study identified an ERSE consensus motif CCAATN9CCACG within the region between nt −61 and −42.11 A point mutation introduced in this element, −59A; G, abolished the promoter activity in the presence of IL-3 (Figure 2f), indicating that IL-3 signaling upregulates Xbp1 transcription through ERSE.
Both PI3K and Stat5 signaling regulate the expression of Xbp1 and PI3K controls splicing of Xbp1 under IL-3 signaling
To understand the upstream signaling network that modulates Xbp1 transcription, PI3K and MEK pathways were inhibited in BaF3 cells. Xbp1 expression was significantly downregulated by a PI3K inhibitor LY294002 but not by PD98059, a MEK inhibitor (Figure 3a). LY294002 treatment also suppressed proliferation of BaF3 cells (data not shown). A reporter assay showed that LY294002 treatment downregulated the promoter activity of Xbp1 both in the 5 kb fragment and in the minimal ERSE element (Figure 3b). On the other hand, addition of PD98059 did not show the suppressive effect and showed enhancement of the luciferase activity in the ERSE element at least partly due to nonspecific effect to the pGL3 promoter construct (Figure 3b). By the western blot analysis, Xbp1S expression was downregulated by the LY294002 treatment and IRE1 phosphorylation was inhibited, whereas Xbp1S expression was not reduced by MEK inhibition (Figure 3c). On the other hand, introduction of dn-Stat5 into BaF3 cells downregulated the Xbp1S expression but not IRE1 phosphorylation. These results indicate that Xbp1 expression is regulated by both PI3K and Stat5 signaling, that PI3K regulates Xbp1 activation by the IRE1 phosphorylation, and that MEK signaling is dispensable for Xbp1 expression.
Xbp1 expression and activation is regulated by PI3K and Stat5 signaling. (a) Real-time PCR of Xbp1. Xbp1 expression was reduced by LY294002 treatment (30 μM) but not PD98059 (50 μM) in BaF3 cells (**P<0.05). (b) Luciferase assay. Significant repression of Xbp1 promoter activity by LY294002 treatment in 32Dcl3 cells after 30 h exposure (**P<0.05). (c) Western blot analysis of BaF3 cells treated with PD98059 or LY294002. LY294002 treatment and dnStat5 expression downregulated Xbp1S expression and LY294002 inhibited IRE1 phosphorylation
Downregulation of Xbp1 induces apoptosis
The efficient knockdown of Xbp1 was achieved by introducing the specific short interference RNA (siRNA) into BaF3 cells (Figure 4a). Apoptosis induction and reduction of viable cells was observed by knockdown of Xbp1 (Figure 4b). Apoptotic changes of BaF3 cells were associated with caspase 3 activation (Figures 4c and d). Treatment of LY294002 or dn-Stat5 suppressed Xbp1S expression, and co-treatment with both reagents completely abolished Xbp1S expression with reduction of Xbp1U expression, which was exhibited by addition of a proteasome inhibitor MG132 (Figure 4e) Similarly, downregulation of Xbp1S induced by co-introduction of LY294002 and dn-Stat5 was accompanied with induction of apoptosis (Figure 4f). These results indicate that Xbp1S has anti-apoptotic function and acts as downstream from PI3K and Stat5 of IL-3 signaling.
Xbp1 downregulation induces cell death. (a) Xbp1 knockdown with Xbp1-specific (#1 and #2) or control siRNAs in BaF3 was confirmed by western blot. (b) The growth of cells treated with Xbp1-specific (#1: closed square, #2: closed circle) or control (closed triangle) siRNAs (**P<0.05). (c) The frequencies of annexin V-positive cells with siRNA (**P<0.05). (d) Cleaved caspase 3, an active form, was detected by Xbp1 knockdown in BaF3 cells. (e) Downregulation of Xbp1S expression by double treatment of dn-Stat5 and LY294002 (30 μM). Dn-Stat5 transfected GFP-positive BaF3 cells were collected by a cell sorter. At 24 h after sorting, the cells were treated with LY294002. (f) Annexin V-positive cells were detected. Significant apoptosis induction was observed by treatment with both dn-Stat5 and LY294002 (**P<0.05)
Xbp1S partially rescues hematopoietic cells from apoptosis by modulating expression of apoptosis-related genes
IL-3 deprivation induces rapid growth arrest and apoptosis of BaF3 cells. In the IL-3-free condition, Xbp1S transfected BaF3 cells survived longer and the number of annexin V-positive cells was reduced compared with control vector, Xbp1U and the splice site mutant Xbp1U532G, that is unable to synthesize the Xbp1S form14 (Figures 5a and b). Moreover, apoptotic induction on LY294002 and dn-Stat5 double treatment was also inhibited by exogenous expression of Xbp1S but not by Xbp1U and Xbp1U532G (Figure 5c). However, the protective effects of Xbp1S expression were incomplete and BaF3 cells failed to survive indefinitely without IL-3. The significant phenotypic changes of hematopoietic cells induced by Xbp1S expression suggest that there are global alterations in gene expression. RNA samples extracted from 32Dcl3 cells following retroviral vector infection with pMIG, pMY-Xbp1U and pMY-Xbp1S were sorted by FACS and then subjected to expression profiling using the Affymetrix GeneChip microarray system (Affymetrix, Santa Clara, CA, USA). Statistical comparisons identified common 1206 genes whose expression were >2.0-fold increased, and 1663 genes <0.5-fold in 32Dcl3-expressing Xbp1S compared with mock-infected cells (Supplementary Tables S1, S2). Sub-categorization of the significantly upregulated genes according to Gene Ontology annotations revealed that 12 upregulated genes on Xbp1S expression were highlighted for close association with apoptosis (Figure 5d). Modulation of Bcl2, Bcl-g, Casp2 and Bim expression was validated by quantitative PCR (Supplementary Figure 1). Expression of Bim, an apoptosis facilitator under IL-3 signaling,7 was significantly reduced in Xbp1S transfected cells (Figure 5e). Upregulation of each isoforms of Bim was observed by knockdown of Xbp1 in BaF3 cells using the Xbp1-specific siRNA (Figure 5f).
Xbp1 supports cell survival. (a) BaF3 cells were infected with pMIG (closed triangle), pMYs-Xbp1U532G (closed square), pMYs-Xbp1U (closed circle) or pMYs-Xbp1S (open square) retroviruses and the number of cells were measured at indicated periods after incubation in the IL-3-free growth medium (**P<0.05). (b and c) Positivities of annexin V for Xbp1-transduced BaF3 cells in IL-3-free condition (b) or treated with both LY294002 and dn-Stat5 (c). The cells were harvested 24 h after IL-3 withdrawal and subjected to the FACS analysis (**P<0.05). (d) Downregulated genes related to apoptosis in Xbp1S transduced 32Dcl3 cells. Expression was compared with pMIG or Xbp1U transduced cells by the Affimetrix GeneChip analysis. (e) Validation of Bim expression by the real-time PCR. (f) Knockdown of Xbp1 induces Bim upregulation. Each Bim isoform was detected by western blotting
Xbp1S modifies cell cycle and differentiation
Flowcytometric analyses revealed that Xbp1S-expressing BaF3 cells showed 1.7-fold increase in the G1 phase population and 40% reduction of the G2 population (Figure 6a), indicating that constitutive expression of Xbp1S induces cell cycle arrest at the G1 phase. Downregulation of cell cycle-related genes was evident in Xbp1S-expressing 32Dcl3 cells compared with those transduced with an empty vector or Xbp1U retrovirus. Significant downregulation of seven cyclin genes as well as six members of the Mcm gene family and four Cdc-related genes was observed (Figure 6b).
Xbp1S modulates cell cycle and differentiation. (a) Cell cycle arrest was induced in BaF3 cells by Xbp1S expression. Proportions of cells at G0/G1 and G2/M phases are indicated. (b) A list of downregulated genes related to cell cycle progression in Xbp1S transduced 32Dcl3 cells. (c) Wright–Giemsa-staining of cytospin samples. 32Dcl3 cells show granulocytic differentiation by Xbp1S expression, while 32Dcl3 cells transduced with pMIG or pMYs-Xbp1U remain immature. (d) Increased intensity of Mac-1 expression in 32Dcl3 cells by GFP-positive population with using pMY-Xbp1S-GFP vector (**P<0.05). (e) A list of upregulated genes associated with myeloid cell differentiation
Wright–Giemsa staining of cytospin samples showed cytoplasmic azure granules and nuclear segmentation in Xbp1S-expressing 32Dcl3 cells, typical for differentiated granulocytes (Figure 6c). Granulocytic differentiation was also confirmed by increased MAC-1 expression in Xbp1S transfected 32Dcl3 cells by using the flowcytometric analysis (Figure 6d). Granulocytic differentiation of 32Dcl3 is induced by G-CSF and is suppressed by IL-3,16 suggesting that Xbp1S may modulate the network of cytokine signaling by switching downstream transcriptional regulation in differentiation processes. Granulocytic differentiation of 32Dcl3 by Xbp1S is closely correlated with upregulation of differentiation related genes including Mef2c, Mmd2, Itgb2 (a component of MAC-1), Ccl3, Ctsd, Ctsg and Cd68 (Figure 6c). Modulation of Mef2c expression affects the number of myeloid progenitors as well as granulocyte maturation,17 suggesting that Xbp1S regulates the Mef2c-mediated differentiation process. A recent study showed downregulation of Mef2c by Xbp1S on myogenic differentiation,18 suggesting that transcriptional regulation by Xbp1S might be cell type-dependent and altered by combination with different co-factors. Carm1 and Ncoa2 could be such co-factors that cooperate with Mef2c in myogenic differentiation,19 however, these two genes were downregulated by Xbp1S in 32Dcl3 (Supplementary Table S2). To the contrary, the genes, such as Bmi1, Gata1, Gata2 and Meis1 that keep the hematopoietic cells at immature stages were downregulated by Xbp1S expression. Modulation of the expression profile by Xbp1S was validated by analyzing mRNA quantities of representative cell cycle- and differentiation-related genes using quantitative PCR (Supplementary Figure 2).
Finally, the expression profiling data shown here were compared with the data from the chromatin immunoprecipitation (ChIP)-on-chip results that exhibited promoter binding sites of Xbp1S on particular genes (Table 1).18 As expected, 109 genes that were identified in both studies were transactivated by Xbp1S, however, 19 genes were repressed, again suggesting there may be the cell type-specific regulatory mechanism. Overall, these data reveal the complexity of Xbp1 function in hematopoiesis and IL-3 signaling.
Discussion
Here, we report that the IL-3 stimulates the Xbp1 by two independent mechanisms. First, Xbp1 expression is upregulated by IL-3 at the transcriptional level via PI3K and STAT5 signaling. Second, Xbp1 mRNA is spliced to convert unstable Xbp1U to the stable form, Xbp1S, encoding a basic leucine zipper protein that is translocated into the nucleus where it exhibits transactivation activity. The PI3K pathway is also important in the second mechanism. Splicing of Xbp1 is mediated by phosphorylation of IRE1, a protein complex of ER transmembrane kinases.11, 14 The spliced form of Xbp1 mRNA is rapidly disappeared on IL-3 withdrawal, and is also decreased by the treatment with a PI3K inhibitor, LY294002. Inhibition of both PI3K and STAT5 pathways suppressed Xbp1S expression and apoptosis induction, suggesting that Xbp1 has the important role of anti-apoptotic factor in the IL-3 signaling. Upregulation of Xbp1 transcription by IL-3 signaling is mediated through ERSE in the promoter region. Previous studies reported that Xbp1 expression is regulated by Xbp1 itself, and by Atf6, both of which bind to the ERSE.12, 14 However, upregulation of Xbp1 promoter activity is more rapid than production of Xbp1S (Figures 2b-d). Moreover, Atf6 is upregulated by Xbp1S as shown in the current gene expression profile. These results suggest that there may be additional network of transcription factors governed by IL-3 and PI3K signaling, which regulates the Xbp1 promoter.
The knockdown of Xbp1-induced growth suppression and caspase 3-dependent apoptosis in BaF3, indicating its important role in cell survival. Introduction of Xbp1S cDNA indeed rescued BaF3 cells from apoptosis and growth suppression mediated IL-3 withdrawal. In addition, the anti-apoptotic effect of Xbp1 was abrogated by the splice site mutation, suggesting that Xbp1 might support cell survival using its transactivation activity and suppressing apoptosis activators, such as Bim.8 It should be noted that the enhanced UPR such as increased IRE1 activity protects cell from apoptosis.20 However, the effect was incomplete and BaF3, which overexpressed Xbp1S failed to grow without IL-3.
Constitutive expression of Xbp1S unexpectedly induced cell cycle arrest in BaF3 cells and cellular differentiation in 32Dcl3 myeloid precursor cells. Xbp1S expression is strongly stimulated after re-activation by IL-3 and it declines to the steady-state level when IL-3 is abundant (Figure 2c). Although the IL-3 stimulus activates Xbp1 promoter activity, only Xbp1S possesses a transactivation domain and has a longer half-life than Xbp1U.21 Since neither growth suppression nor differentiation induction was observed by Xbp1U introduction, constitutive expression of Xbp1S could affect homeostasis of hematopoietic cells. When Xbp1S was expressed under the control of the immunoglobulin VH promoter/Eμ enhancer in a transgenic mouse model, the mice developed multiple myeloma.22 Upregulated expression of XBP1S was also noted in human cases of multiple myeloma,22 suggesting different actions of Xbp1S in hematopoietic precursors and plasma cells. Indeed, Xbp1S is induced by plasma cell differentiation and IgM heavy chain production in B cells.23 Xbp1S also induces IL-6 secretion in B cells, however, IL-6 expression was decreased in 32Dcl3 cells, again suggesting different behavior of Xbp1S between hematopoietic precursors and differentiated B-lineage cells.
Cytokine-induced expression of Xbp1 has also been reported in case of IL-4 treatment of B cells.23 Although both IL-3 and IL-4 enhance transcription and splicing of Xbp1, induction by IL-4 is more rapid than by IL-3 and upregulation of GRP78 is observed in the presence of IL-4 but not in IL-3. Perhaps IRE1 phosphorylation is directly downstream from IL-4 signaling whereas PI3K and STAT5 signaling is mediated after cellular stimulation by IL-3. Additional studies will be needed to address the question.
Global and extensive modification of gene expression by Xbp1S in 32Dcl3 was observed as expected from gross phenotypic changes. Combining data from our expression profiling and genome-wide ChIP is useful to understand the gene regulatory network modulated by Xbp1S as summarized in Table 1. It is likely that most of these genes are direct targets of Xbp1S, and many UPR-inducible genes are included in the list. In contrast, the genes related to differentiation, cell cycle progression and apoptotic function are not included, suggesting that these biological processes may be induced as an indirect consequences of UPR. In summary, these studies identify a close relationship between the cytokine signaling system and UPR, and the important role of Xbp1 in hematopoietic cell homeostasis as well as leukemogenic process.
Materials and Methods
Cell culture and treatment
BaF3 and 32Dcl3, were maintained with 10 ng/ml of IL-3 (R & D Systems, Minneapolis, MN, USA) as described previously.16 IL-3 signaling inhibitor, PD98059 (Cell Signaling Technologies, Beverly, MA, USA) and LY294002 (Sigma, St. Louis, MO, USA) were used at the 50 μM and 30 μM, respectively.
Retrovirus infection and selection of survival clones under IL-3-free condition
The MSCV-GFP-Neo retroviral vector is composed of enhanced green fluorescent protein (EGFP) and neomycin resistance genes flanked with MSCV long terminal repeats. The plasmid was transfected to Plat-E packaging cells (a gift from Dr. Toshio Kitamura) using lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) according to the manufacture's protocol. In all, 3 × 106 BaF3 cells were added with retroviral stock with 6 μg/ml of polybrene (Sigma) and spun at 1400 × g for 2 h. After 24 h, cells were seeded into 96-well plates at 1 × 104 cells per well in G418-containing growth medium without IL-3.
Isolation of RISs
RISs were identified using the IPCR approach as described previously.24 Briefly, genomic DNA was digested with EcoRI, BamHI, BglII, NcoI, HindIII or SacI, self-ligated, and subjected to nested IPCR. The PCR primers for each restriction digestion are available on request. The PCR products were analyzed using agarose gel electrophoresis, subcloned into the pGEM T-easy plasmid (Promega, Madison, WI, USA), and subjected to sequence analysis.
Southern blot analysis
Southern blot analysis was carried out to assess the clonal insertion of the retrovirus. The genomic DNA was digested with appropriate restriction enzymes, subjected to agarose gel electrophoresis and transferred to Hybond-N nylon filter (GE Healthcare, Little Chalfont, UK) the filter was hybridized with the GFP sequence as a probe.
RT-PCR
The splicing of Xbp1 was detected by RT-PCR using (Promega) and Expand High Fidelity PCR System (Roche, Basel, Switzerland) using the Xbp1 362F (5′-TAACGGGAGAAAACTCACGGC-3′) and Xbp1 1017R (5′-CAGACAATGGCTGGATGAAAGC-3′) primers. PCR products were digested by PstI to differentiate Xbp1U from Xbp1S,25 separated by electrophoresis on 2% agarose gels and visualized by ethidium bromide staining.
Western blot analysis
Western blot analysis was performed using total cell lysates as described previously.26 Primary antibodies used were anti-GAPDH, anti-Xbp1, anti-GRP78, anti-IRE1, anti-ATF6, anti-caspase 3 (Santa Cruz Biotech, Santa Cruz, CA, USA), anti-phospho-IRE1 (Novus Biologicals, Littleton, CO, USA), anti-phospho-Stat5, anti-Akt, anti-phospho-Ser473-Akt, anti-Erk and phospho-Erk (Cell Signaling Technologies), anti-Bim (Pharmingen, San Diego, CA, USA). Secondary anti-rat, rabbit, and anti-mouse IgG antibodies were purchased from GE Healthcare. For detection to Xbp1U, cells were treated with 10 μM MG132 (Sigma) for 2 h as indicated.14
Luciferase assay
Genomic DNA fragments of the Xbp1 promoter region were amplified by PCR using a mouse BAC clone as a template and subcloned into the pGL3 basic or pGL3 promoter vectors (Promega). To induce point mutations within IL-3-responsive element, the fragment was amplified by using specific primers, –59G (5′-TGCTAGCGAGCACCGGCCGATCGCGGAC-3′) within ERSE. In all, 1 μg of the reporter plasmid together with 50 ng of pRL-SV40 were introduced into a total of 1 × 106 32Dcl3 cells by using Nucleofector device and solution V (Lonza, Cologne, Germany). Luciferase activities were measured using the Dual-Luciferase Reporter Assay System (Promega) and Lumat LB9507 (Perkin Elmer, Waltham, MA, USA).
siRNA and retroviral vector infection
In all, 2 μg of siRNA for Xbp1 purchased from QIAGEN (Hilden, Germany) was introduced into BaF3. Entire coding regions of Xbp1S and Xbp1U were amplified and FLAG-tagged by RT-PCR, and subcloned into the pMYs-IRES-GFP retroviral vector (a gift from Toshio Kitamura). To induce point mutations within Xbp1U was amplified by using specific primer 532G (5′-CCAGCAGGTGCAGGCCCAGTTGTC-3′). To inhibit IL-3 signaling, pMYs-dn-Stat5-IRES-GFP was used (a gift from Toshio Kitamura). Infection efficiency was determined by measuring GFP-positive cells using FACScalibur flow cytometer (Becton Dickinson, Franklin Lakes, NJ, USA).
Cell cycle assay, flow cytometer and apoptosis assay
Single cell suspensions were permeabilized with 0.1% triton X-100 in PBS, and 50 mg/ml propidium iodide and 1 mg/ml RNAse A was added. The cell suspensions were then analyzed by using a FACScalibur flow cytometer (Beckton Dickinson) and Modifit software (Becton Dickinson). For the surface marker analysis, suspensions of 1 × 106 cells were incubated with fluorescein isothiocyanate-conjugated anti-Mac-1 (Pharmingen) for 30 m, washed three times in PBS, applied to a flow cytometer and fluorescence intensity of GFP-positive population was detected. Apoptosis assays were performed using annexin V–PE apoptosis detection kit I (Pharmingen) according to the manufacturer's protocol.
Array hybridization
The mouse Genome 430 2.0 Array was hybridized with cRNA probes generated from Xbp1-expressing 32Dcl3 cells. Arrays were scanned using an Affymetrix confocal scanner and analyzed using Affymetrix Gene Chip Command Console Software (AGCC, Affymetrix) and GeneSpring (Agilent Technologies, Santa Clara, CA, USA) as described previously.26, 27
Statistical analysis
Results were evaluated statistically by using Student's t-test. A value of P<0.05 was considered significant.
Abbreviations
- IL-3:
-
interleukin-3
- UPR:
-
unfolded protein response
- GM-CSF:
-
granulocyte-macrophage colony-stimulating factor
- RIM:
-
retrovirus insertional mutagenesis
- ER:
-
endoplasmic reticulum
- ERSR:
-
ER stress response element
- ChIP:
-
chromatin immunoprecipitation
- EGFP:
-
enhanced green fluorescent protein
- RIS:
-
retroviral integration sites
- IPCR:
-
inverse polymerase chain reaction
- RT:
-
reverse transcription
- siRNA:
-
short interference RNA
References
Sachs L . The control of hematopoiesis and leukemia: from basic biology to the clinic. Proc Natl Acad Sci USA 1996; 93: 4742–4749.
Reddy EP, Korapati A, Chaturvedi P, Rane S . IL-3 signaling and the role of Src kinases, JAKs and STATs: a covert liaison unveiled. Oncogene 2000; 19: 2532–2547.
Steelman LS, Pohnert SC, Shelton JG, Franklin RA, Bertrand FE, McCubrey JA . JAK/STAT, Raf/MEK/ERK PI3K/Akt and BCR-ABL in cell cycle progression and leukemogenesis. Leukemia 2004; 18: 189–218.
Lotem J, Cragoe EJ, Sachs L . Rescue from programmed cell death in leukemic and normal myeloid cells. Blood 1991; 78: 953–960.
Ito T, Deng X, Carr B, May WS . Bcl-2 phosphorylation required for anti-apoptosis function. J Bio Chem 1997; 272: 11671–11673.
Sakai I, Kraft AS . The kinase domain of Jak2 mediates induction of bcl-2 and delays cell death in hematopoietic cells. J Biol Chem 1997; 272: 12350–12358.
Shinjyo T, Kuribara R, Inukai T, Hosoi H, Kinoshita T, Miyajima A et al. Downregulation of Bim a proapoptotic relative of Bcl-2, is a pivotal step in cytokine-initiated survival signaling in murine hematopoietic progenitors. Mol Cell Biol 2001; 21: 854–864.
Robert K, Francesco P, Andreas SB, Soon-Siong T, Ralph T, Jakob RP et al. A gain of function of JAK2 in myeloproliferative disorders. New Engl J Med 2005; 352: 1779–1790.
Nakamura T . Retoviral insertional mutagenesis identifies oncogenic cooperation. Cancer Sci 2005; 96: 7–12.
Kiser KF, Colombi M, Moroni C . Isolation characterization of dominant and recessive IL-3-independent hematopoietic transformants. Oncogene 2006; 25: 6595–6603.
Calfon M, Zeng H, Urano F, Till JH, Hubbard SR, Harding HP et al. IRE1 couples endoplasmic reticulum load to secretory capacity by processing the Xbp1 mRNA. Nature 2002; 415: 92–96.
Yoshida H, Okada T, Haze K, Yanagi H, Yura T et al. ATF6 activated by the proteolysis binds in the presence of NF-Y (CBF) directly to the cis-acting element responsible for the mammalian unfolded protein response. Mol Cell Biol 2000; 20: 6755–6767.
Rutkowsky DT, Kaufman RJ . A trip to the ER: coping with stress. Trends Cell Biol 2004; 14: 20–28.
Yoshida H, Oku M, Suzuki M, Mori K . pXbp1 (U) encoded in Xbp1 pre-mRNA negative regulates unfolded protein response activator pXbp1 (S) in mammalian ER stress response. J Cell Biol 2006; 172: 565–575.
Yoshida H, Haze K, Yanagi H, Yura T, Mori K . Identification of the cis-acting endoplasmic reticulum stress response element responsible for transcriptional induction of mammalian glucose-regulated proteins. J Biol Chem 1998; 273: 33741–33749.
Fujino T, Yamazaki Y, Largaespada DA, Jenkins NA, Copeland NG, Hirokawa K et al. Inhibition of myeloid differentiation by Hoxa9, Hoxb8, Meis homeobox genes. Exp Hematol 2001; 29: 856–863.
Johnnidis JB, Harris MH, Wheeler RT, Stehling-Sun S, Lam MH, Kirak O et al. Regulation of progenitor cell proliferation and granulocyte function by microRNA-223. Nature 2008; 451: 1125–1129.
Acosta-Alvear D, Zhou Y, Blais A, Tshikitis M, Lents NH, Arias C et al. Xbp1 control diverse cell type- and condition-specific transcriptional regulatory networks. Mol Cell 2007; 27: 53–66.
Chen SL, Loffler KA, Chen D, Stallcup MR, Muscat GE . The coactivator-associated arginine methyltransferase is necessary for muscle differentiation. J Biol Chem 2002; 277: 4324–4333.
Lin JH, Li H, Yasumura D, Cohen HR, Zhang C, Panning B et al. IRE1 signaling affects cell fate during the unfolded protein response. Science 2007; 318: 944–949.
Ye Y, Shibata Y, Yun C, Ron D, Rapoport TA . A member protein complex mediates retro-translocation from the ER lumen into the cytosol. Nature 2004; 429: 841–847.
Carrasco DR, Sukhdeo K, Protopopova M, Sinha R, Enos M, Carrasco DE et al. The differentiation and stress response factor Xbp1 drives multiple myeloma pathogenesis. Cancer Cell 2008; 4: 349–360.
Iwakoshi NN, Lee AH, Vallabhajosyula P, Otipoby KL, Rajewsky K, Glimcher LH . Plasma cell differentiation and the unfolded protein response intersect at the trasnscriptional factor Xbp1. Nat Immunol 2003; 4: 321–329.
Tanaka M, Jin G, Yamazaki Y, Takahara T, Takuwa M, Nakamura T . Identification of candidate cooperative genes of the Apc mutation in the trasnsformation of the colon epithelial cell by the retroviral insertional mutagenesis. Cancer Sci 2008; 99: 979–985.
Yoshida H, Matsui T, Yamamoto A, Okada T, Mori K . Xbp1 mRNA is induced by ATF6 and spliced by IRE1 in response to ER stress to produce a highly active transcriptional factor. Cell 2001; 107: 881–891.
Kawamura-Saito M, Yamazaki Y, Kaneko K, Kawaguchi N, Kanda H, Mukai H et al. Fusion between CIC and DUX4 up-regulates PEA3 family genes in Ewing-like sarcomas with t(4;19)(q35;q13) traslocation. Hum Mol Genet 2006; 15: 2125–2137.
Fujino T, Nomura K, Ishikawa Y, Makino H, Umezawa A, Aburatani H et al. Function of EWS-POU5F1 in sarcomagenesis and tumor cell maintenance. Am J Pathol 2010; 176: 1973–1982.
Acknowledgements
We thank Dr. Toshio Kitamura for Plat-E cells. This work was supported by Grant-in-Aid for Scientific Research on Priority Areas ‘Integrative Research Toward the Conquest of Cancer’ from the Ministry of Education, Culture, Sports, Science and Technology (TN), and by Grant-in-Aid for Young Scientist (Start-up) from Japan Society for the Promotion of Science (M Kurata).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by P Salomoni
Supplementary Information accompanies the paper on Cell Death and Disease website
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Kurata, M., Yamazaki, Y., Kanno, Y. et al. Anti-apoptotic function of Xbp1 as an IL-3 signaling molecule in hematopoietic cells. Cell Death Dis 2, e118 (2011). https://doi.org/10.1038/cddis.2011.1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cddis.2011.1
Keywords
This article is cited by
-
Dynamics of precision-guided projectile launch: fluid–structure interaction
Acta Mechanica (2021)
-
A negative feedback loop between XBP1 and Fbw7 regulates cancer development
Oncogenesis (2019)
-
Leptin Promotes Allergic Airway Inflammation through Targeting the Unfolded Protein Response Pathway
Scientific Reports (2018)
-
Apoptotic action of botulinum toxin on masseter muscle in rats: early and late changes in the expression of molecular markers
SpringerPlus (2016)
-
New directions in ER stress-induced cell death
Apoptosis (2013)