Abstract
Prostate cancer (CaP) is mostly composed of luminal-like differentiated cells, but contains a small subpopulation of basal cells (including stem-like cells), which can proliferate and differentiate into luminal-like cells. In cancers, CpG island hypermethylation has been associated with gene downregulation, but the causal relationship between the two phenomena is still debated. Here we clarify the origin and function of CpG island hypermethylation in CaP, in the context of a cancer cell hierarchy and epithelial differentiation, by analysis of separated basal and luminal cells from cancers. For a set of genes (including GSTP1) that are hypermethylated in CaP, gene downregulation is the result of cell differentiation and is not cancer specific. Hypermethylation is however seen in more differentiated cancer cells and is promoted by hyperproliferation. These genes are maintained as actively expressed and methylation-free in undifferentiated CaP cells, and their hypermethylation is not essential for either tumour development or expansion. We present evidence for the causes and the dynamics of CpG island hypermethylation in CaP, showing that, for a specific set of genes, promoter methylation is downstream of gene downregulation and is not a driver of gene repression, while gene repression is a result of tissue-specific differentiation.
Similar content being viewed by others
Main
It is likely that cancer originates from environmental, genetic and epigenetic perturbation of tissue stem or progenitor cells.1, 2 The end result of these aberrations is a cancer mass possessing a cellular hierarchy which mirrors that in normal tissue.3 Accumulating evidence now shows just such a hierarchical structure in many cancers4 including prostate.5 In particular, prostate cancer (CaP) is characterized by an imbalance of the differentiation process, resulting in the accumulation of proliferative differentiated luminal cancer cells (composing >99% of the tumour).6, 7, 8 A small population of basal cells (<1% of cells), which shares chromosomal changes such as a TMPRSS2-ERG fusion with the luminal cancer cells,9 still persists in CaP. Although the field is still debating on the origin and phenotype of prostate cancer stem cells,10, 11, 12, 13 several lines of evidence show that they might indeed reside within this small population of basal cells.5, 12, 13, 14
Aberrant DNA methylation patterns are found in all cancers, and hypermethylation of CpG islands in promoter regions has been associated with cancer-specific gene downregulation, promoting both cancer development and progression.15 However, it is unclear whether hypermethylation causes gene downregulation16 or simply maintains it, by adding a final ‘lock’ on the promoter, whereas the initial gene downregulation is carried out by DNA methylation-independent mechanisms.17, 18 CaP is characterized by hypermethylation of numerous promoters,19 but also an unexpectedly high intra-patient variability, where each individual develops a unique DNA methylation signature.20 Of the genes hypermethylated in CaP, GSTP1 is the best studied.21 It encodes a detoxifying enzyme, which is able to protect cells from DNA adduct formation by electrophilic compounds.22 GSTP1 hypermethylation is already present in preneoplastic lesions such as prostatic intraepithelial neoplasia (PIN),23 promoting the hypothesis that epigenetic downregulation of GSTP1 sensitizes cancer precursor cells (cell of origin) to carcinogenic insults and promotes tumour progression.24 However, the causes of GSTP1 downregulation and hypermethylation in CaP are still unknown.
This study aimed to understand the function and origin of CpG island hypermethylation in CaP, in the context of cancer hierarchy and differentiation. We report that a set of genes commonly hypermethylated in CaP (including GSTP1) is (i) downregulated as a result of prostate-specific epithelial differentiation in both CaP and benign prostatic hyperplasia (BPH); (ii) selectively hypermethylated only in differentiated (luminal) cells, most likely promoted by their hyperproliferative phenotype; (iii) actively expressed and methylation free in undifferentiated (basal) CaP cells. Downregulation and hypermethylation of these genes is however not essential for tumour development or tumour expansion. Moreover, for all these genes, the downregulation induced by prostate-specific differentiation is independent of DNA hypermethylation and is associated with detachment of RNAPolII from their promoters and a reduction in histone marks associated with active transcription.
Results
GSTP1 is downregulated through epithelial differentiation and hypermethylated only in CaP luminal cells
GSTP1 promoter methylation was quantified by pyrosequencing in a panel of CaP cell lines (Figure 1a) spanning the entire spectrum of epithelial differentiation from basal to luminal. In agreement with previous reports,25 high levels of DNA methylation (>90%) were found in luminal CaP cell lines (LnCaP, PC346C and VCaP), whereas no methylation (RC-165N/hTERT, PNT1A, PNT2-C2) or little methylation (BPH-1) was found in benign cell lines. Cancer cell lines with an intermediate differentiation phenotype (PC3, DU145)26 showed partial methylation (50–60%). Strikingly, no methylation was found in cancer-derived cell lines with a basal phenotype (P4E6, Bob and SerBob).27, 28
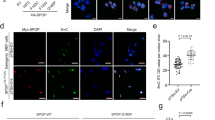
GSTP1 is hypomethylated and highly expressed in undifferentiated basal prostate cancer cells. (a) Pyrosequencing methylation analysis of the GSTP1 promoter performed in prostate cell lines (bars=single CpG sites; n=3 technical replicates; mean±S.D.; line=average of 14 CpG sites, Positive Control=RC-165N/hTERT DNA methylated with SssI methyltransferase). (b) Pyrosequencing methylation analysis of GSTP1 performed in basal and luminal cells derived from BPH and CaP (each dot represents the average of 14 CpG sites analysed in a single sample; boxplots show minimum, 25%, median, 75% and maximum, hypermethylation threshold (dot-dashed line)=average methylation of BPH basal+2 S.D., P-values from Mann–Whitney test). (c) qRT-PCR analysis of GSTP1 expression relative to GAPDH in basal and luminal cells derived from BPH and CaP (boxplots show minimum, 25%, median, 75% and maximum; each dot represents a single sample, P-values from Mann–Whitney test). (d) qRT-PCR analysis of GSTP1 expression relative to GAPDH in primary prostate cancer xenografts generated in RAG2−/− γC−/− mice. (e) Pyrosequencing methylation analysis of GSTP1 performed in primary prostate cancer xenografts (left panel), MACS selected cells from disaggregated xenografts tumours (central panel), and matched xenografts and original tumour tissue (right panel) (bars=single CpG sites; n=3 technical replicates; mean±S.D.; line=average of 14 CpG sites)
We then analysed GSTP1 methylation and expression levels in separated basal and luminal cells isolated from BPH and CaP tissues. Lin−/CD31−/CD24+ luminal cells were isolated from disaggregated prostate primary tissues (Supplementary Figure 1). As a source of basal cells, primary prostate epithelial cultures were generated as previously described5, 29, 30 from fresh BPH and CaP tissues. Cultures from CaP tissues still retain cancer features: (i) increased invasion capacity and (ii) proliferative potential,5 genomic rearrangements such as (iii) TMPRSS2-ERG fusion5, 9, 29 and (iv) microsatellite instability,5 (v) high telomerase expression and activity (Rane et al, in preparation), (vi) overexpression of cancer-associated genes (eg, AMACR and MMP9) and downregulation of PTEN.29 We have confirmed the cells’ phenotype by immunofluorescence and qRT-PCR: Lin−/CD31−/CD24+ cells were PanCytokeratin+/GSTP1−/Cytokeratin5−/AR+/Cytokeratin8+/PSA+, while prostate epithelial cultures were PanCytokeratin+/GSTP1+/Cytokeratin5+/AR−/Cytokeratin8−/PSA− (Supplementary Figures 2 and 3A and B). We defined Lin−/CD31−/CD24+ cells as ‘luminal’ cells and primary epithelial cultures as ‘basal’ cells.
CaP luminal cells clearly showed hypermethylation of GSTP1 promoter (11/17 samples), only rarely seen in BPH luminal cells (2/16 samples with <10% methylation) (Figure 1b). Strikingly, CaP basal cells showed no GSTP1 hypermethylation.
GSTP1 mRNA levels (Figure 1c) were high in both BPH and CaP basal cells, with a significant downregulation in luminal cells and no significant difference between BPH and CaP. High expression in CaP basal cells was confirmed by the reanalysis of our previous microarray data29 (Supplementary Figure 4).
As an alternative source for undifferentiated CaP cells, we utilised ‘near-patient’ xenografts generated in BALB/c/RAG2−/−γC−/− mice.31 Xenografts mainly show an intermediate phenotype co-expressing basal and luminal markers, with less than 5% of partially differentiated cells (CD24+/ARlow). Compared with P4E6 (basal) and LNCaP (luminal), all the xenografts analysed actively expressed GSTP1, with small variations between samples (Figure 1d). Moreover, none of the xenografts showed hypermethylation of GSTP1 either as an unfractionated tissue (Figure 1e, left) or as fractionated cell populations representing heterogeneous (Lin−), partially differentiated (Lin−/CD24+) or undifferentiated cells (Lin−/CD44+ and Lin−/CD133+) (Figure 1e, central). For one xenograft (H027/10), we confirmed that hypermethylation of GSTP1 was present in the original tumour tissue but lost upon grafting (Figure 1e, right). Altogether these results show that GSTP1 is not hypermethylated and is highly expressed in undifferentiated basal-like cancer cells.
GSTP1 methylation correlates with the differentiation status of hyperproliferating prostate epithelial cells
The previous results strongly indicated that GSTP1 is hypermethylated only in luminal CaP cells. As these cells are highly proliferative compared with their normal counterparts,32 significantly upregulate DNMT3A compared with basal cells (Supplementary Figures 3C–F) and DNA methyltransferases are mostly active in the S phase of the cell cycle,33 we hypothesized that a combination of cell differentiation, high DNMTs and hyperproliferation could be the primary cause of GSTP1 hypermethylation in CaP. To test this, we dissected GSTP1 hypermethylation heterogeneity in BPH-1 cells: an established cell model for hyperproliferating prostate cells with an intermediate phenotype.
BPH-1 cells34 have a short doubling time of 20 h, indicating hyperproliferation, express intermediate levels of GSTP1, and have a partially methylated GSTP1 promoter (Supplementary Figures 5A and B and Figure 1). Immunofluorescence analysis revealed heterogeneous expression of GSTP1, Cytokeratin5 and Cytokeratin8 in individual cells (Supplementary Figure 5A), while qRT-PCR showed low but detectable levels of AR, PAP and PSA (Supplementary Figure 5B), indicative of a heterogeneous intermediate phenotype. An almost perfect correlation was found between GSTP1 expression and basal cell markers Cytokeratin5 and p63 (Supplementary Figure 6), indicating GSTP1 upregulation in more undifferentiated cells. To dissect BPH-1 heterogeneity, eighteen clones were derived from single BPH-1 cells. These clones show extreme variability in GSTP1 mRNA levels, protein expression (Figures 2a and d) and promoter methylation (Figure 2b), with a strong inverse correlation between expression and methylation (Figure 2c). Taken altogether, these results suggest that GSTP1 expression and methylation are strictly correlated and linked to the cells’ differentiation state.
GSTP1 expression and promoter methylation correlates with differentiation of hyperproliferating prostate epithelial cells. (a) RT-PCR analysis of GSTP1 and ACTB (β-actin) expression in 18 randomly selected clones of BPH-1 cells. Five independent preparations of the parental cell line were used as a control for the stability of GSTP1 expression and reliability of the technique (WP-1–WP-5). (b) Pyrosequencing methylation analysis of the GSTP1 promoter performed on the same clones normalized versus the parental cell line (WP) (bars=single CpG sites; n=3 technical replicates; mean±SD; line=average of 14 CpG sites). (c) GSTP1 expression plotted against promoter methylation in BPH-1 clones. GSTP1 expression was normalized to β-actin and calibrated against the average of the five WP samples (open circle) (each dot represent a single clone, dashed line=linear regression). (d) Western blot analysis for GSTP1 on three hypermethylating clones (B8, C1, C3), two hypomethylating clones (C6, C11) and two clones (C4, C9) with average methylation levels comparable to the parental cell line (WP). (e) Immunofluorescence analysis of Cytokeratin5 (KRT5) and GSTP1 levels in BPH-1 colonies. (f) Pyrosequencing methylation analysis of the GSTP1 performed on the DNA extracted from the colonies shown in e after the immunofluorescence pattern was recorded
To directly confirm this link, more BPH-1 clones were derived, colonies were fixed after 7 days’ growth, and co-stained for GSTP1 and Cytokeratin5 (Figure 2e). Colonies with an overall overexpression or downregulation of both markers were identified (exemplified in Figure 2e). Methylation analysis performed on the same cells after immunofluorescence revealed that colonies with low levels of GSTP1 and Cytokeratin5 also hypermethylated GSTP1, compared with colonies with high levels of both proteins (Figure 2f). This confirmed that, in a hyperproliferating system, GSTP1 hypermethylation is present preferentially in differentiated cells.
Identification of DAH genes: a set of genes downregulated through prostate-specific differentiation and selectively hypermethylated in CaP
We then investigated to what extent the conclusions made for GSTP1 (downregulated in CaP, downregulated in luminal cells and highly expressed in basal CaP cells) could be extrapolated to all the genes hypermethylated in CaP. We generated a comprehensive list of promoters hypermethylated in CaP from publicly available resources and assessed gene expression in three data sets: CaP versus normal prostate tissues,35 luminal versus basal primary prostate cells,36 CaP versus BPH-derived basal cells.29 As expected, ‘genes hypermethylated in CaP’ were significantly enriched for genes downregulated in CaP (Supplementary Figure 7A), confirming the general correlation between hypermethylation and downregulation in cancer. In luminal versus basal prostate cells, although an almost equal proportion of genes was up or downregulated, there was a clear enrichment for genes strongly downregulated in luminal cells (Supplementary Figure 7B), suggesting that a significant subset of these genes is already repressed in normal luminal cells. Surprisingly, most ‘genes hypermethylated in CaP’ remained unchanged between CaP and BPH basal cells (Supplementary Figure 7C), indicating that undifferentiated cancer cells could have a profoundly different expression and methylation profiles compared with differentiated cells, and that many other genes could indeed behave similarly to GSTP1.
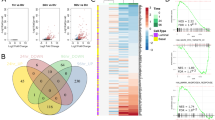
We then generated a list of genes highly expressed in prostate basal cells from BPH and CaP29 and intersected it with the list of genes hypermethylated in CaP, finding 206 genes that are potentially regulated similarly to GSTP1 (Figure 3a and Supplementary Table 1). We named these genes ‘Differentiation-Associated Hypermethylated’ (DAH). Expression analysis of DAH genes in the same data sets35, 36 revealed that they are strongly enriched for genes downregulated in CaP tissues, and also in normal luminal cells (Figures 3b and c), supporting the hypothesis that these genes are, like GSTP1, mainly downregulated during differentiation and subsequently hypermethylated in CaP.
Identification of DAH genes: a set of genes behaving similarly to GSTP1. (a) Schematic representation of the workflow undertaken to select genes hypermethylated in prostate cancer luminal-like cells, but actively expressed in prostate cancer basal cells (described in details in materials and methods section). (b, c) Expression analysis of DAH genes in CaP versus Normal (b), and Luminal versus Basal normal prostate cells (c) (differentially expressed= all probes with P<0.05 in a t-test comparing the two sample groups; P-values from Mann-Whitney test; Red-blue=difference of the mean log2 fold change for each set of genes). (d) Chromatin status of DAH promoters in H1-hESCs (data retrieved from ENCODE database). (e) Selection of representative DAH genes for further analysis
As hypermethylation in cancer has been associated with areas marked by bivalent chromatin in embryonic stem cells (ESCs),37 we interrogated the ENCODE database for the presence of H3K4me3 and H3K27me3 on the promoters of DAH genes in H1-hESCs38 (Figure 3d). More than 40% of DAH genes were marked by both H3K4me3 and H3K27me3, indicative of bivalent chromatin. However, almost half of DAH genes were marked only by H3K4me3, whereas a very small proportion of genes was marked by H3K27me3. This suggested that DAH genes are either in an active or in poised state in ESCs, are then maintained as active or activated in prostate stem cells (high expression) and repressed during prostate specific differentiation.
We also checked the frequency of hypermethylation of the DAH promoters in CaP (Kobayashi et al.39 data set). The results indicated a wide variation ranging from high (74%) to very low (1%) (Supplementary Figure 7D). Thus, downregulation in hyperproliferating luminal cells may not always be sufficient to cause consistent gene hypermethylation.
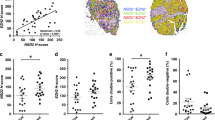
We then selected seven representative DAH genes for further analysis, through two independent processes (Figure 3e): CCND2, CSTA and S100A14 were selected by the bioinformatic analysis described in Supplementary Figure 8; while SFN, THBS1, DKK3 and LDHB displayed the most consistent previously published evidence for promoter hypermethylation in CaP and downregulation during prostate epithelial differentiation (Figure 3e and Supplementary Table 2). All seven genes were consistently downregulated in luminal cells compared with basal cells in both BPH and CaP (Figure 4a), while the DNA methylation analysis showed that they were almost never hypermethylated in basal cells from both BPH and CaP (Figure 4b). In luminal cells, hypermethylation was either frequent in both BPH and CaP (CSTA, S100A14 and SFN), frequent only in CaP (CCND2) or rare but CaP specific in (DKK3, LDHB, THBS1). Taken together, these results show that several genes do behave similarly to GSTP1: they are consistently downregulated through differentiation in both BPH and CaP, while DNA methylation arises preferentially in luminal cells, with some genes being hypermethylated only in CaP, while others in both BPH and CaP.
CCND2, CSTA, DKK3, LDHB, S100A14, SFN and THBS1 are hypomethylated and highly expressed in basal fraction of primary CaP. (a) qRT-PCR analysis of CCND2 (i), CSTA (ii), DKK3 (iii), LDHB (iv), S100A14 (v), SFN (vi) and THBS1 (vii) expression relative to GAPDH in basal and luminal cells derived from BPH and CaP (boxplots show minimum, 25%, median, 75% and maximum; each dot represents a single sample, P-values from Mann–Whitney test). (b) Pyrosequencing methylation analysis of CCND2 (i), CSTA (ii), DKK3 (iii), LDHB (iv), S100A14 (v), SFN (vi) and THBS1 (vii) in basal and luminal cells derived from BPH and CaP (each dot represent the average of all the CpG sites analysed in a single sample; boxplots show minimum, 25%, median, 75% and maximum, hypermethylation threshold (dot-dashed line)=average methylation of BPH basal+2 standard deviations, P-values from Mann–Whitney test, the two BPH basal samples that clearly hypermethylated SFN were excluded from the threshold calculation)
Differentiation of prostate epithelial cells induces downregulation of DAH genes, promoter transcriptional inactivation, but not DNA hypermethylation
To better understand the regulation of DAH genes, we studied their behaviour in a prostate epithelial differentiation model in vitro, by culturing the cells as acinus-like spheroids in Matrigel40 (3D conditions). To monitor differentiation, BPH-1 cells were transduced with a lentiviral vector where the expression of mOrange is under the control of PSAPb promoter, specifically activated in the later stages of prostatic differentiation30 (Figure 5a). After 7 days of 3D culture, mOrange was upregulated in ∼50% of the spheroids (Figure 5a). mOrange+ cells were localized in the centre of the spheroid, indicative of prototypic gland formation.27 AR and PSA transcripts were upregulated in 3D cultures compared with standard 2D cultures (Supplementary Figures 9A and B), whereas both Cytokeratin5 and GSTP1 were downregulated in the inner part of the spheroid (Supplementary Figure 9C), confirming prostate-specific differentiation and indicating polarization of the spheroid. AR protein was found only in rare cells (Supplementary Figure 9C) suggesting that complete luminal differentiation was reached only in a subfraction of cells. Around 80% of the spheres also contained cells expressing activated-caspase3 in the inner part of the spheroid. These cells were Cytokeratin5LOW and Cytokeratin8+, indicating that a proportion of the differentiated cells was unstable and underwent apoptosis (Supplementary Figure 10). As previously described,40 cell death is not essential for complete spheroid formation and is probably a limitation of the in vitro conditions of this model.
Differentiation of BPH-1 cells induces downregulation and transcriptional inactivation of differentiation-associated hypermethylated genes, but not promoter hypermethylation. (a, top) Representative image of BPH1-PPO cells grown in 2D (standard culture conditions) and 3D (differentiating conditions) showing induction of mOrange expression in differentiating conditions, indicative of successful differentiation. (a, bottom) Schematic representation of the viral vector used to generate BPH-1 luminal reporter cells. (b) qRT-PCR analysis of DAH candidate genes expression relative to GAPDH in BPH-1 cells grown in 2D and 3D. (c–f): ChIP-qPCR analysis carried out in BPH-1 cells grown in 2D and 3D with (c) anti-RNAPolII, (d) anti-H3K4Me3, (e) anti-acetylated H3 (f) anti-H3K27Me3 (data presented as % of immunoprecipitated DNA). (g) Pyrosequencing methylation analysis of DAH candidate genes performed on BPH-1 PPO cells grown in 2D and 3D (bars=single CpG sites; n=3 technical replicates; mean±S.D.; line=average of 14 CpG sites)
Expression of all the DAH genes tested, except CCND2, was significantly reduced in 3D conditions compared with 2D (Figure 5b), confirming that these genes are downregulated during prostate differentiation. This was accompanied by a repression of the transcriptional activity of their promoters, measured by a decrease in bound RNAPolII and a reduction of the active chromatin marks H3K4Me3 and/or H3 acetylation; while the repressive mark H3K27Me3 remained almost constant in all genes (Figures 5c–f). However, only a very small and nonsignificant increase in DNA hypermethylation (Figure 5g) was found, suggesting that transcriptional inactivation and chromatin rearrangements precede DNA methylation during DAH genes’ downregulation.
To clarify the role of established DNA methylation patterns on DAH genes’ expression, we pharmacologically inhibited DNA methylation (5-aza-2′deoxycytidine, 1 μM for 96 h) in P4E6 (basal) and LNCaP (luminal) cells. As expected, all DAH genes (except LDHB) were hypermethylated in LNCaP (Supplementary Figure 10A). After treatment, all the genes showed upregulation in LNCaP (Supplementary Figure 10B), suggesting that, once established, DNA methylation could either promote further downregulation or act as a primary repression signal.
Downregulation and hypermethylation of DAH genes are not associated with tumourigenic induction of prostate epithelial cells
To understand whether the downregulation and hypermethylation of DAH genes are strictly associated with prostate tumour formation, BPH-1 cells were recombined with either normal prostate fibroblasts (NPF – inducing normal growth) or cancer-associated fibroblasts (CAF – inducing tumour formation) and grafted in the renal capsule of NOD-SCID mice41 (Figure 6a). BPH-1+CAF tumours exhibit squamous-like differentiation and do not express AR in their epithelial component.41, 42 Both BPH-1+NPF and BPH-1+CAF grafts expressed high levels of PAP and undetectable levels of PSA (Figure 6b), confirming partial differentiation. Expression analysis of all the DAH genes showed no differences between BPH-1+NPF grafts and BPH-1+CAF grafts (Figure 6c), again confirming that the regulation of these genes is not linked to the malignant nature of the cells and that downregulation of DAH genes is not necessary for tumour formation. Compared with BPH-1, expression of GSTP1, CSTA, DKK3, S100A14 and SFN was upregulated in the grafts, whereas CCND2 and LDHB remained unchanged and THBS1 was downregulated (Figure 6c), suggesting that the correct differentiation programme (prostate adenomatous-like versus squamous-like) is necessary for the downregulation of DAH genes. As expected, DNA methylation of these genes did not change between BPH-1+NPF and BPH-1+CAF grafts (Figure 6d), while GSTP1, CSTA and S100A14 methylation was considerably lower in grafts compared with BPH-1 cells, again showing a direct correlation between methylation and expression, but no relationship to tumorigenesis.
Expression and methylation of DAH genes is not related to induction of prostate tumorigenesis. (a) Diagram explaining the in vivo experimental design. (b) qRT-PCR for PSA and PAP in BPH-1 cells and renal grafts generated by recombining BPH-1 cells with patient matched normal prostate fibroblasts (BPH-1+NPF, n=3) or cancer-associated fibroblasts (BPH-1+CAF, n=3). qRT-PCR (c) and pyrosequencing methylation (d) analysis of DAH candidate genes in the same samples
Discussion
CpG island hypermethylation has been demonstrated in all cancer types at multiple genomic loci. Because of its early appearance and frequency, it is thought to be one of the cancer’s founding alterations and thus occurs in the cancer cell of origin, potentially even before DNA mutation. In all cancers, including CaP, hypermethylation is responsible for the downregulation of tumour suppressor genes,16 promoting both cancer development and progression. However, the mechanisms by which CpG island hypermethylation originates in cancer are still poorly understood. Here we show a direct link between tissue-specific differentiation, gene downregulation and hypermethylation in CaP. In order to dissect intra-tumour cellular heterogeneity, we have analysed primary prostate basal and luminal cells derived from BPH and CaP separately. In this way, we identified a set of genes (DAH) frequently hypermethylated in CaP, which is primarily downregulated through tissue-specific differentiation, both in normal tissues and cancer. For these genes, we hypothesize that DNA methylation can arise only after gene downregulation and is aided by cell hyperproliferation. Moreover, downregulation and hypermethylation of these genes are not essential for either tumour development or expansion.
This set of genes includes GSTP1, the most studied hypermethylated gene in CaP. According to the literature, promoter hypermethylation directly represses GSTP1 in CaP, rendering CaP cells (or precancerous cells) more susceptible to carcinogenic insults by electrophilic compounds.22, 43, 44 However, this theory is based on findings indicating a strong correlation, but not a causal relationship, between gene downregulation and hypermethylation in CaP. In contrast, a detailed molecular analysis of GSTP1 regulation showed that gene silencing precedes de novo methylation, and that a random ‘seed’ of methylation is necessary for the initiation of promoter hypermethylation.45 However, the mechanisms of the initial GSTP1 downregulation and methylation ‘seeding’ were not defined as yet.
Our data now clearly show that GSTP1 is primarily downregulated through epithelial differentiation in both BPH and CaP, while promoter hypermethylation arises only in differentiated (luminal) cancer cells, most likely promoted by the hyperproliferation of these cells.
We found DAH genes to be equally downregulated in luminal cells from CaP and BPH, while basal cells constitutively expressed these genes, irrespective of their origin. Moreover, DAH genes were also significantly downregulated in CaP compared with normal prostate. If we consider that hormone naive cancers are mostly composed of luminal-like cells (<1:100 basal/luminal cell ratio in CaP versus ∼1:1 to ∼1:2.7 in the benign epithelium),6, 7, 8, 46 we can infer that the cancer-specific downregulation seen is due to differences in ratios of cell types, rather than to a cancer-cell-specific gene repression. This conclusion is also in line with recent reports showing that hypermethylation in cancer occurs more frequently in genes already repressed in normal and precancerous tissues.47
Our data also clearly show that the small fraction of basal cancer cells continues to express high levels of DAH genes. Basal cancer cells are considered to contain the prostate cancer stem cells that can self-renew, proliferate and differentiate in luminal cancer cells.5, 12, 13, 14, 31 In the context of a cancer cell hierarchy, we conclude that DAH genes are primarily downregulated during the differentiation of cancer stem cells into cancer luminal cells, in a DNA methylation-independent manner.
As both BPH and CaP-derived luminal cells express low levels of DAH genes, it is of critical importance to understand the mechanisms inducing hypermethylation specifically in cancer. For GSTP1, we have shown that methylation arises preferentially in differentiated cells that are aberrantly proliferating, where the gene has been already downregulated, as previously hypothesized;48 this process could also be aided by the high levels of DNMT3A found in luminal cells. In line with this, GSTP1 hypermethylation is present in a proportion of precancerous lesions (PIN),23 where luminal cells are first observed starting to hyperproliferate.32 However, as both the frequency and levels of hypermethylation in DAH genes are extremely variable, and as some of the DAH genes analysed have a very low frequency of hypermethylation in CaP, we can hypothesise that other (unknown) factors are required for the successful methylation of DAH genes in luminal cells.
Because of its high frequency and early onset, promoter hypermethylation is thought to be a key step in cancer development. Our results challenge this hypothesis, showing that DAH genes’ methylation is lost in primary CaP xenografts, mainly composed of undifferentiated cells, and it is not induced in the BPH-1+CAF tumour model. This indicates that hypermethylation of DAH genes is (i) strictly linked to complete prostate luminal differentiation; (ii) not essential for the in vivo expansion of CaP cells; and (iii) not essential for induction of prostate tumour formation.
In summary, we show that DAH genes are both expressed and unmethylated in BPH and CaP basal cells, downregulated as a result of prostate epithelial differentiation, in a process involving changes in histone modifications, and hypermethylated solely in CaP luminal cells, through their hyperproliferative phenotype (Figure 7). Overall, this is the first report to explain the causes and the dynamics of CpG island hypermethylation in CaP, where, for this specific set of genes, which represents 5% of all genes reported to be hypermethylated in CaP, promoter methylation is downstream of gene downregulation and is not a driver of gene repression.
Therefore our results clearly show that basal CaP cells do not hypermethylate DAH genes, which are detected as methylated in unfractionated CaP tissues. Although DNA hypermethylation remains an extremely important cancer-specific biomarker, currently being fully assessed in both tissues and body fluids, our results pose important questions about the development of epigenetic therapies for cancer, which do not consider the basal subpopulation of cells. Future investigations should be focused on understanding which genes are affected by aberrant epigenetic modifications in cancer stem cells as well as differentiated cells, as these could represent valuable alternative therapeutic targets for effective cancer treatment of tumours with an acknowledged poor chemotherapeutic response rate.
Material and Methods
BPH and CaP tissues collection
Prostate tissues were obtained from patients undergoing TURP or radical prostatectomy at York Hospital (York, UK) and Castle Hill Hospital (Cottingham, UK) with informed patient consent and approval from the NRES Committee Yorkshire & The Humber. Tissues were sampled immediately after surgery. For TURPs, a proportion of the prostate chips was collected for analysis. For radical prostatectomies, three core needle biopsies were taken from four different sites (left base, left apex, right base, right apex) and were directed by previous pathology, imaging and palpation. Tissues were transported in RPMI-1640 with 5% FCS and 100 U/ml antibiotic/antimycotic solution at 4 °C, and processed within 6 h. BPH or CaP diagnosis was confirmed by histological examination of representative adjacent fragments. A list of the samples utilized in this study is presented in Supplementary Table 3.
Selection of Lin−/CD31−/CD24+ cells from BPH and CaP tissues and establishment of primary cultures
Tissues were disaggregated as previously described,5, 49 all the digestions and incubations were performed in the presence of 10 nM 5α-dihydrotestosterone (5α-DHT) in order to preserve viability of prostate luminal cells. Single-cell suspensions were cleaned from cell debris and dead cells by centrifugation on a density gradient of Lymphocyte Separation Media (#0850494X, MP Biomedicals, Cambridge, UK), and then labelled with Lineage Cell Depletion Kit (human) and CD31 MicroBead Kit (#130-092-211 and #130-091-935 Miltenyi Biotec, Surrey, UK) following the manufacturer’s instruction. Lin+/CD31+ cells were depleted twice using MACS LS Columns (#130-042-401 Miltenyi Biotec). Lin−/CD31− cells were then labelled with CD24 MicroBead Kit (#130-095-951 Miltenyi Biotec) and Lin−/CD31−/CD24+ cells were selected twice using MACS MS Columns (#130-042-201 Miltenyi Biotec) (Supplementary Figure 1A). To determine the purity of the populations, cells were labelled with CD24-PE (human) (#130-095-953 Miltenyi Biotec) following the manufacturer’s instructions and analysed on a CyAn ADP flow cytometer (Dako Cytomation, Stockport, UK) (Supplementary Figure 1B). CD24− cells were plated on to type I Collagen-coated 100 mm plates (BD Biosciences, Oxford, UK) in the presence of irradiated STO feeder cells as previously described.5, 30, 50 Primary cultures were subsequently analysed at passage 0 or 1.
Generation and maintenance of CaP xenografts
Xenografts were generated from tissue biopsies from four patients undergoing either radical prostatectomy (Y042/07) or palliative channel TURP (H016, Y019 and H027/10). Patient details are shown in Supplementary Table 3. Tissue biopsies were engrafted subcutaneously into intact male BALB/c/RAG2−/−γC−/− mice. Mice were supplemented with a slow release 5α-DHT tablet (Innovative Research of America) if the patient was hormone naive at the time of biopsy. Once tumours reached 1.5 cm3, which was considered a humane end point, the mice were killed and the tumours were either reimplanted into further mice or the tissue was digested for further experiments. To maintain the tumour xenograft as ‘near-patient,’ tumours were re-established from frozen cells after five passages in mice. To generate single cells from xenografts, the same method used for human CaP tissues was utilized, and endothelial and haematopoietic cells were depleted using Lineage Cell Depletion Kit, mouse (#130-090-858 Miltenyi Biotec). Then, cell populations were separated using magnetic sorting and these kits (following the manufacturer’s instructions): CD24 MicroBead Kit (#130-095-951), CD44 MicroBeads (#130-095-194), CD133 MicroBead Kit (#130-050-801, all from Miltenyi Biotec).
Cell lines
A list of the cell lines used, origin and culture conditions is provided in Supplementary Table 4. Cells were cultured at 37 °C in a humidified atmosphere with 5% CO2, handled under good laboratory practice conditions in defined passage windows, monthly certified free of Mycoplasma and genotyped to ensure authenticity.
Generation of BPH-1 clones
BPH-1 cells were seeded at 312, 156 or 78 cells/dish in 10 cm Petri dishes (Corning, Amsterdam, The Netherlands). After 7 days, clones were isolated using sterile cloning rings and cells were expanded until 80% confluent in a T75 flask (Corning).
Generation of BPH-1 PSAPb-mOrange cells
Lentiviral vectors for the expression of mOrange under the control of the PSAPb promoter were produced, and BPH-1 cells were infected and selected with Blasticidin 4 μg/ml as previously described.30, 51
In vitro differentiation of BPH-1 cells
BPH-1 cells were grown in 3D cultures under differentiating conditions as previously described40, 52, 53 with some modifications: BPH-1 cells were seeded on a 50%(v/v) Matrigel plug and cultured in KSFM with 2% FCS, 2 mM L-glutamine, 10 nM R1881 and 10 nM β-estradiol, 5 ng/ml EGF, 1 μg/ml FGF, 4% (v/v) Matrigel, in co-culture with prostate stroma derived from a patient with high Gleason grade cancer.
Generation of BPH-1 grafts
Primary cultures of matched NPFs and CAFs were established from radical prostatectomy specimens with patient consent and human ethics approval (Human Ethics Research Approvals 34306 at Epworth Hospital, 03-14-04-08 at Cabrini Hospital and RMO 2006/6108-2004000145 at Monash University). BPH-1 cells were recombined with fibroblasts from three independent patients and grafted into immune-deficient NOD-SCID mice as previously described41 in accordance with Monash University ethics approval (Approval Numbers: MMCA/2007/04 and MMCA/2008/33).
RNA extraction and qRT-PCR analysis
RNA was extracted using the RNeasy Mini Kit (Qiagen, Manchester, UK) or RNeasy Micro Kit (Qiagen), and reverse transcribed using random hexamers and reverse transcriptase (Superscript III, Invitrogen, Life Technologies Ltd, Paisley, UK). Real-time PCR was carried out using the enzyme mix SsoFast EvaGreen Supermix (Bio-Rad, Hemel Hempstead, UK) and specific primers (Supplementary Table 5) in a CFX96 real-time PCR detection system. Data were analysed using the Bio-Rad CFX Manager 2.0 (Bio-Rad).
DNA extraction, bisulphite conversion and pyrosequencing methylation analysis
DNA was extracted using the DNeasy Blood & Tissue Kit (Qiagen) or classic phenol/chloroform extraction for small samples. A 100% methylated control was generated by treating RC-165N/hTERT DNA with SssI methyltransferase (New England Biolabs, Hitchin, UK) for 4 h at 37 °C. Fifty nanograms to 1 μg of DNA was bisulphite converted using the EpiTect Bisulphite Kit (Qiagen). Pyrosequencing was carried out as previously described50 using gene-specific primers (Supplementary Table 5 and Supplementary Figure 11 for assay localization). Hypermethylation was defined as a signal higher than the average methylation +2 S.D. of the BPH basal population, as described in Håvik et a.l54
Chromatin Immunoprecipitation
Chromatin immunoprecipitation was performed as described.55 In brief, the chromatin of cells grown in 2D or 3D was cross-linked using formaldehyde and sonicated with a Bioruptor UCD-200 (Diagenode, Seraing, Belgium) (five cycles of 5 min each, 30 s ON 30s OFF, at full power and 4 °C) to obtain a chromatin average size of 300–400 bp. Immunoprecipitation was carried out overnight with 2.4 μg of antibody O/N at 4 °C. Immunoprecipitated DNA was extracted by phenol/chloroform extraction using linear acrylamide (Applied Biosystems, Life Technologies Ltd) as a carrier. DNA was then analysed by qPCR as described above, using specific primers (Supplementary Table 5 and Supplementary Figure 11 for assay localization).
Immunofluorescence analysis and image analysis
Immunofluorescence was carried out on cultured cells as previously described.29 Lin−/CD31−/CD24+ cells were spotted on a 3-aminopropyltriethoxysilane-treated glass slide before proceeding with the same protocol. Immunofluorescence on BPH-1 cells cultured in 3D was carried out as previously described.40 A list of antibodies is presented in Supplementary Table 6.
Western Blot analysis
Western blot analysis was performed as described.56 Antibodies used were anti GSTP1 (HPA019779) and anti B-actin (A5316) from Sigma-Aldrich (Dorset, UK), anti DNMT3A (ab13888), DNMT3B (ab13604), DNMT1 (ab13537) and GAPDH (ab9485) from Abcam (Cambridge, UK).
Identification of ‘Differentiation-Associated Hypermethylated’ genes
First, a comprehensive list of 3891 unique genes reported to be hypermethylated in CaP was generated by combining publicly available data in multiple publications39, 48, 57, 58 and the CaP section of the following databases: pubmeth.org59 and MethCancerDB.60 Second, a list of 1407 genes highly expressed in prostate basal cells (from both BPH and CaP) was generated by a re-analysis of the microarray data set previously published by our lab29 (raw data available in the ArrayExpress Database, accession E-MEXP-993). In brief, raw data were reanalyzed by robust multi-array averaging (RMA) and calculated as log2 expression. Genes were selected on the basis of these criteria: (i) no significant difference (P>0.05 in a student’s t-test) between BPH and CaP in any of the subpopulations (CD133+/α2β1hi or CD133−/α2β1low) for all of the probes matching each gene; (ii) average log2 expression across all samples greater than 10. The intersect of these two lists generated 206 ‘Differentiation-Associated Hypermethylated’ genes (DAH genes) potentially behaving similarly to GSTP1.
Accession codes
Abbreviations
- 5α-DHT:
-
5α-dihydrotestosterone
- BPH:
-
benign prostatic hyperplasia
- CAF:
-
cancer-associated fibroblasts
- CaP:
-
prostate cancer
- DAH:
-
differentiation-associated hypermethylated genes
- ESCs:
-
embryonic stem cells
- NPF:
-
normal prostate fibroblasts
- PIN:
-
prostatic intraepithelial neoplasia
- RMA:
-
robust multi-array averaging
References
Feinberg AP, Ohlsson R, Henikoff S . The epigenetic progenitor origin of human cancer. Nat Rev Genet 2006; 7: 21–33.
Sell S . On the stem cell origin of cancer. Am J Pathol 2010; 176: 2584–2494.
Miller SJ, Lavker RM, Sun T-T . Interpreting epithelial cancer biology in the context of stem cells: tumor properties and therapeutic implications. Biochim Biophys Acta 2005; 1756: 25–52.
Clevers H . The cancer stem cell: premises, promises and challenges. Nat Med 2011; 17: 313–319.
Collins AT, Berry PA, Hyde C, Stower MJ, Maitland NJ . Prospective identification of tumorigenic prostate cancer stem cells. Cancer Res 2005; 65: 10946–10951.
El-Alfy M, Pelletier G, Hermo LS, Labrie F . Unique features of the basal cells of human prostate epithelium. Microsc Res Tech 2000; 51: 436–446.
Humphrey PA . Diagnosis of adenocarcinoma in prostate needle biopsy tissue. J Clin Pathol 2007; 60: 35–42.
Nagle RB, Ahmann FR, McDaniel KM, Paquin ML, Clark VA, Celniker A . Cytokeratin characterization of human prostatic carcinoma and its derived cell lines. Cancer Res 1987; 47: 281–286.
Polson ES, Lewis JL, Celik H, Mann VM, Stower MJ, Simms MS et al. Monoallelic expression of TMPRSS2/ERG in prostate cancer stem cells. Nat Commun 2013; 4: 1623.
Shen MM, Wang X, Economides KD, Walker D, Abate-Shen C . Progenitor cells for the prostate epithelium: roles in development, regeneration, and cancer. Cold Spring Harb Symp Quant Biol 2008; 73: 529–538.
Wang ZA, Shen MM . Revisiting the concept of cancer stem cells in prostate cancer. Oncogene 2011; 30: 1261–1271.
Oldridge EE, Pellacani D, Collins AT, Maitland NJ . Prostate cancer stem cells: are they androgen-responsive? Mol Cell Endocrinol 2012; 360: 14–24.
Goldstein AS, Stoyanova T, Witte ON . Primitive origins of prostate cancer: in vivo evidence for prostate-regenerating cells and prostate cancer-initiating cells. Mol Oncol 2010; 4: 385–396.
Rane JK, Pellacani D, Maitland NJ . Advanced prostate cancer-a case for adjuvant differentiation therapy. Nat Rev Urol 2012; 9: 595–602.
Portela A, Esteller M . Epigenetic modifications and human disease. Nat Biotechnol 2010; 28: 1057–1068.
Cedar H, Bergman Y . Programming of DNA methylation patterns. Annu Rev Biochem 2012; 81: 97–117.
Jones PA . Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet 2012; 13: 484–492.
Clark SJ, Melki J . DNA methylation and gene silencing in cancer: which is the guilty party? Oncogene 2002; 21: 5380–5387.
Goering W, Kloth M, Schulz WA . DNA methylation changes in prostate cancer. Methods Mol Biol 2012; 863: 47–66.
Aryee MJ, Liu W, Engelmann JC, Nuhn P, Gurel M, Haffner MC et al. DNA methylation alterations exhibit intraindividual stability and interindividual heterogeneity in prostate cancer metastases. Sci Transl Med 2013; 5: 169ra10.
Meiers I, Shanks JH, Bostwick DG . Glutathione S-transferase pi (GSTP1) hypermethylation in prostate cancer: review 2007. Pathology 39: 299–304.
Nelson CP, Kidd LC, Sauvageot J, Isaacs WB, De Marzo AM, Groopman JD et al. Protection against 2-hydroxyamino-1-methyl-6-phenylimidazo[4,5-b]pyridine cytotoxicity and DNA adduct formation in human prostate by glutathione S-transferase P1. Cancer Res 2001; 61: 103–109.
Nakayama M, Bennett CJ, Hicks JL, Epstein JI, Platz EA, Nelson WG et al. Hypermethylation of the human glutathione S-transferase-pi gene (GSTP1) CpG island is present in a subset of proliferative inflammatory atrophy lesions but not in normal or hyperplastic epithelium of the prostate: a detailed study using laser-capture microdissection. Am J Pathol 2003; 163: 923–933.
Nelson WG, De Marzo AM, Deweese TL, Lin X, Brooks JD, Putzi MJ et al. Preneoplastic prostate lesions: an opportunity for prostate cancer prevention. Ann N Y Acad Sci 2001; 952: 135–144.
Yegnasubramanian S, Kowalski J, Gonzalgo ML, Zahurak M, Piantadosi S, Walsh PC et al. Hypermethylation of CpG islands in primary and metastatic human prostate cancer. Cancer Res 2004; 64: 1975–1986.
van Leenders GJ, Aalders TW, Hulsbergen-van de Kaa CA, Ruiter DJ, Schalken JA . Expression of basal cell keratins in human prostate cancer metastases and cell lines. J Pathol 2001; 195: 563–570.
Lang SH, Smith J, Hyde C, Macintosh C, Stower M, Maitland NJ . Differentiation of prostate epithelial cell cultures by matrigel/ stromal cell glandular reconstruction. In Vitro Cell Dev Biol Anim 2006; 42: 273–280.
Attard G, Rizzo S, Ledaki I, Clark JP, Reid AHM, Thompson A et al. A novel, spontaneously immortalized, human prostate cancer cell line, Bob, offers a unique model for pre-clinical prostate cancer studies. Prostate 2009; 69: 1507–1520.
Birnie R, Bryce SD, Roome C, Dussupt V, Droop A, Lang SH et al. Gene expression profiling of human prostate cancer stem cells reveals a pro-inflammatory phenotype and the importance of extracellular matrix interactions. Genome Biol 2008; 9: R83.
Frame FM, Hager S, Pellacani D, Stower MJ, Walker HF, Burns JE et al. Development and limitations of lentivirus vectors as tools for tracking differentiation in prostate epithelial cells. Exp Cell Res 2010; 316: 3161–3171.
Maitland NJ, Frame FM, Polson ES, Lewis JL, Collins AT . Prostate cancer stem cells: do they have a basal or luminal phenotype? Horm Cancer 2011; 2: 47–61.
Ananthanarayanan V, Deaton RJ, Yang XJ, Pins MR, Gann PH . Alteration of proliferation and apoptotic markers in normal and premalignant tissue associated with prostate cancer. BMC Cancer 2006; 6: 73.
Robertson KD, Keyomarsi K, Gonzales FA, Velicescu M, Jones PA . Differential mRNA expression of the human DNA methyltransferases (DNMTs) 1, 3a and 3b during the G(0)/G(1) to S phase transition in normal and tumor cells. Nucleic Acids Res 2000; 28: 2108–2113.
Hayward SW, Dahiya R, Cunha GR, Bartek J, Deshpande N, Narayan P . Establishment and characterization of an immortalized but non-transformed human prostate epithelial cell line: BPH-1. In Vitro Cell Dev Biol Anim 1995; 31: 14–24.
Liu P, Ramachandran S, Ali Seyed M, Scharer CD, Laycock N, Dalton WB et al. Sex-determining region Y box 4 is a transforming oncogene in human prostate cancer cells. Cancer Res 2006; 66: 4011–4019.
Oudes AJ, Campbell DS, Sorensen CM, Walashek LS, True LD, Liu AY . Transcriptomes of human prostate cells. BMC Genomics 2006; 7: 92.
Easwaran H, Johnstone SE, Van Neste L, Ohm J, Mosbruger T, Wang Q et al. A DNA hypermethylation module for the stem/progenitor cell signature of cancer. Genome Res 2012; 22: 837–849.
Raney BJ, Cline MS, Rosenbloom KR, Dreszer TR, Learned K, Barber GP et al. ENCODE whole-genome data in the UCSC genome browser (2011 update). Nucleic Acids Res 2011; 39: D871–D875.
Kobayashi Y, Absher DM, Gulzar ZG, Young SR, McKenney JK, Peehl DM et al. DNA methylation profiling reveals novel biomarkers and important roles for DNA methyltransferases in prostate cancer. Genome Res 2011; 21: 1017–1027.
Pearson JF, Hughes S, Chambers KF, Lang SH . Polarized fluid movement and not cell death, creates luminal spaces in adult prostate epithelium. Cell Death Differ 2009; 16: 475–482.
Taylor RA, Toivanen R, Frydenberg M, Pedersen J, Harewood L et alAustralian Prostate Cancer Bioresource. Human epithelial basal cells are cells of origin of prostate cancer, independent of CD133 status. Stem Cells 2012; 30: 1087–1096.
Hayward SW, Wang Y, Cao M, Hom YK, Zhang B, Grossfeld GD et al. Malignant transformation in a nontumorigenic human prostatic epithelial cell line. Cancer Res 2001; 61: 8135–8142.
Lin X, Tascilar M, Lee WH, Vles WJ, Lee BH, Veeraswamy R et al. GSTP1 CpG island hypermethylation is responsible for the absence of GSTP1 expression in human prostate cancer cells. Am J Pathol 2001; 159: 1815–1826.
Nakayama M, Gonzalgo ML, Yegnasubramanian S, Lin X, De Marzo AM, Nelson WG . GSTP1 CpG island hypermethylation as a molecular biomarker for prostate cancer. J Cell Biochem 2004; 91: 540–552.
Song JZ, Stirzaker C, Harrison J, Melki JR, Clark SJ . Hypermethylation trigger of the glutathione-S-transferase gene (GSTP1) in prostate cancer cells. Oncogene 2002; 21: 1048–1061.
Hudson DL, Guy AT, Fry P, O’Hare MJ, Watt FM, Masters JR . Epithelial cell differentiation pathways in the human prostate: identification of intermediate phenotypes by keratin expression. J Histochem Cytochem 2001; 49: 271–278.
Sproul D, Kitchen RR, Nestor CE, Dixon JM, Sims AH, Harrison DJ et al. Tissue of origin determines cancer-associated CpG island promoter hypermethylation patterns. Genome Biol 2012; 13: R84.
Schulz WA, Hatina J . Epigenetics of prostate cancer: beyond DNA methylation. J Cell Mol Med 2006; 10: 100–125.
Richardson GD, Robson CN, Lang SH, Neal DE, Maitland NJ, Collins AT . CD133, a novel marker for human prostatic epithelial stem cells. J Cell Sci 2004; 117: 3539–3545.
Pellacani D, Packer RJ, Frame FM, Oldridge EE, Berry PA, Labarthe M-C et al. Regulation of the stem cell marker CD133 is independent of promoter hypermethylation in human epithelial differentiation and cancer. Mol Cancer 2011; 10: 94.
Hager S, Frame FM, Collins AT, Burns JE, Maitland NJ . An internal polyadenylation signal substantially increases expression levels of lentivirus-delivered transgenes but has the potential to reduce viral titer in a promoter-dependent manner. Hum Gene Ther 2008; 19: 840–850.
Chambers KF, Pearson JF, Aziz N, O’Toole P, Garrod D, Lang SH . Stroma regulates increased epithelial lateral cell adhesion in 3D culture: a role for actin/cadherin dynamics. PLoS One 2011; 6: e18796.
Chambers KF, Pearson JF, Pellacani D, Aziz N, Guzvic M, Klein CA et al. Stromal upregulation of lateral epithelial adhesions: Gene expression analysis of signalling pathways in prostate epithelium. J Biomed Sci 2011; 18: 45.
Håvik AB, Brandal P, Honne H, Dahlback H-SS, Scheie D, Hektoen M et al. MGMT promoter methylation in gliomas-assessment by pyrosequencing and quantitative methylation-specific PCR. J Transl Med 2012; 10: 36.
Dahl JA, Collas P . A rapid micro chromatin immunoprecipitation assay (microChIP). Nat Protoc 2008; 3: 1032–1045.
Rivera-Gonzalez GC, Droop AP, Rippon HJ, Tiemann K, Pellacani D, Georgopoulos LJ et al. Retinoic acid and androgen receptors combine to achieve tissue specific control of human prostatic transglutaminase expression: a novel regulatory network with broader significance. Nucleic Acids Res 2012; 40: 4825–4840.
Li L-C, Okino ST, Dahiya R . DNA methylation in prostate cancer. Biochim Biophys Acta 2004; 1704: 87–102.
Jerónimo C, Bastian PJ, Bjartell A, Carbone GM, Catto JWF, Clark SJ et al. Epigenetics in prostate cancer: biologic and clinical relevance. Eur Urol 2011; 60: 753–766.
Ongenaert M, Van Neste L, De Meyer T, Menschaert G, Bekaert S, Van Criekinge W . PubMeth: a cancer methylation database combining text-mining and expert annotation. Nucleic Acids Res 2008; 36: D842–D846.
Lauss M, Visne I, Weinhaeusel A, Vierlinger K, Noehammer C, Kriegner A . MethCancerDB - aberrant DNA methylation in human cancer. Br J Cancer 2008; 98: 816–817.
Acknowledgements
We thank Dr Simon Hayward for provision of BPH-1 cells, Dr Johng Rhim for provision of RC-165N/hTERT cells and Dr David Hudson for provision of Bob and SerBob cells. We also thank Dr Stefanie Hager for the generation of the BPH-1 PSAPb-mOrange cells, Katherine M Wilson and Dr Euan S Polson for their help with the BPH-1 clones, Dr Jayant K Rane for his help with the DNMTs experiments, and Dr Guillermo Rivera-Gonzalez and Dr David Hudson for critical evaluation of the manuscript. This work was supported by Yorkshire Cancer Research (Project grant Y256 and Program grant) and donations from The Freemasons’ Grand Charity and York Vikings Rotary Club.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by JP Medema
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Supplementary information
Rights and permissions
About this article
Cite this article
Pellacani, D., Kestoras, D., Droop, A. et al. DNA hypermethylation in prostate cancer is a consequence of aberrant epithelial differentiation and hyperproliferation. Cell Death Differ 21, 761–773 (2014). https://doi.org/10.1038/cdd.2013.202
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2013.202
Keywords
This article is cited by
-
DNA hypermethylation driven by DNMT1 and DNMT3A favors tumor immune escape contributing to the aggressiveness of adrenocortical carcinoma
Clinical Epigenetics (2023)
-
DNA methylation profile is associated with the osteogenic potential of three distinct human odontogenic stem cells
Signal Transduction and Targeted Therapy (2018)
-
Phenotype-independent DNA methylation changes in prostate cancer
British Journal of Cancer (2018)
-
MiR-455-3p inhibits the degenerate process of chondrogenic differentiation through modification of DNA methylation
Cell Death & Disease (2018)
-
Methylation of PITX2, HOXD3, RASSF1 and TDRD1 predicts biochemical recurrence in high-risk prostate cancer
Journal of Cancer Research and Clinical Oncology (2014)