Abstract
Cytotoxic lymphocyte protease granzyme M (GrM) is a potent inducer of tumor cell death. The apoptotic phenotype and mechanism by which it induces cell death, however, remain poorly understood and controversial. Here, we show that GrM-induced cell death was largely caspase-dependent with various hallmarks of classical apoptosis, coinciding with caspase-independent G2/M cell cycle arrest. Using positional proteomics in human tumor cells, we identified the nuclear enzyme topoisomerase II alpha (topoIIα) as a physiological substrate of GrM. Cleavage of topoIIα by GrM at Leu1280 separated topoIIα functional domains from the nuclear localization signals, leading to nuclear exit of topoIIα catalytic activity, thereby rendering it nonfunctional. Similar to the apoptotic phenotype of GrM, topoIIα depletion in tumor cells led to cell cycle arrest in G2/M, mitochondrial perturbations, caspase activation, and apoptosis. We conclude that cytotoxic lymphocyte protease GrM targets topoIIα to trigger cell cycle arrest and caspase-dependent apoptosis.
Similar content being viewed by others
Main
Cytotoxic T lymphocytes and natural killer (NK) cells form the first line of defense against virus-infected and tumor cells.1 Upon target cell recognition, these cytotoxic lymphocytes can initiate target cell death via the death receptor or the granule exocytosis pathway.2 In the latter, cytotoxic lymphocytes release the pore-forming protein perforin and a set of structurally homologous serine proteases known as granzymes. Perforin facilitates entry of granzymes into the target cell, where they can cleave intracellular substrates to induce cell death. In humans, five granzymes have been identified.1, 3, 4, 5
Human granzyme M (GrM)6 is expressed in NK, γδ-T, NKT, and CD8+ T cells.7, 8, 9 GrM is unique in that it preferably cleaves after a Leu or Met residue.10, 11 GrM efficiently kills tumor cell lines such as HeLa, Jurkat, HL60, and K562 cells.12, 13, 14, 15, 16 In addition, a GrM-anti CD64 fusion protein has recently been shown to induce cell death in human primary leukemic cells in vitro.16 The in vivo importance of GrM is still unclear. In one study, GrM knockout mice clear tumors just as efficiently as wild-type (wt) mice.17 However, in another study, GrM is important in the anti-tumor effect mediated by adoptively transferred NK cells.18 The lack of a clear-cut phenotype in GrM knockout mice may be due to the redundancy of the murine granzymes or to species-specific differences between the human and mouse GrM orthologues.10, 19
The apoptotic phenotype and molecular mechanism of GrM-mediated cell death in human tumor cells are still unclear and remain controversial in the literature. Several studies have shown that GrM triggers cell death in a caspase-independent fashion, without fragmentation of DNA or perturbation of the mitochondria.13, 14 In contrast, other studies reported that GrM-mediated cell death occurs in the presence of caspase-3 activation, DNA fragmentation, reactive oxygen species (ROS) generation, and cytochrome c release from the mitochondria.15, 20, 21, 22 Over the years, several GrM substrates have been identified.10, 12, 13, 15, 20, 21, 22, 23 Of these, only Fas-associated protein with death domain (FADD) was univocally proven to have an important role in GrM-mediated apoptosis.15 Cleavage of human FADD by GrM promotes pro-caspase-8 recruitment and activation and subsequent initiation of the caspase cascade.15, 19 However, FADD-deficient cancer cells are only partially resistant to GrM,15 indicating that there is at least one other important mediator via which GrM induces apoptosis.
In the present study, we comprehensively characterized the phenotype of GrM-induced cell death. GrM treatment resulted in largely caspase-dependent cell death exhibiting classical hallmarks of apoptosis. Furthermore, we showed for the first time that GrM triggered G2/M cell cycle arrest. In the absence of caspase-8 – and thus the GrM-FADD-caspase-8 pathway15– both cell cycle arrest and caspase activation still occurred. To understand these caspase-8/FADD-independent GrM functions, we used positional proteomics in HeLa tumor cells to identify DNA topoisomerase II alpha (topoIIα) as a novel GrM substrate. We conclude that, next to FADD-cleavage mediated caspase activation, cytotoxic lymphocyte protease GrM targets topoIIα to trigger cell cycle arrest and caspase-dependent apoptosis.
Results
GrM triggers classical hallmarks of apoptosis
The phenotype of GrM-mediated cell death remains controversial in the literature. Therefore, we comprehensively characterized apoptotic hallmarks in GrM-treated human tumor cells. Recombinant human GrM or catalytically inactive GrM-SA (inactive GrM mutant in which the catalytic site Ser residue has been mutated to an Ala residue) were delivered into cells with the perforin-analogue streptolysin O (SLO). GrM triggered cell death in HeLa cells as measured by a WST-1 cell viability assay, reflecting the number of metabolically active, adherent cells (Figure 1a). Similarly, when Jurkat cells were treated with GrM, an increase in cells with fragmented DNA (subG1) was observed (Figure 1b). To further characterize the type of cell death induced by GrM, HeLa cells were stained with AnnexinV-fluos (AnnV) and propidium iodide (PI) and analyzed by flow cytometry (Figures 1c and d) or fluorescence microscopy (Supplementary Figure S1a). GrM-treated cells first became AnnV positive, and later AnnV/PI double-positive, suggesting death via classical apoptosis. Similar results were obtained for Jurkat cells treated with GrM delivered by SLO (data not shown) and perforin (Supplementary Figure S1b). Typically, upon induction of classical apoptosis, DNases are activated, leading to DNA fragmentation. Indeed, in GrM-treated cells, an increase in TdT dUTP nick-end labeling (TUNEL)-positive cells was observed (Figure 1e), indicative of DNA fragmentation. In addition, loss of mitochondrial membrane potential – as measured with the fluorescent dye DiOC6 – accompanied by an increase in ROS and the release of cytochrome c were observed (Figures 1f–h). Interestingly, treatment with GrM resulted in rapid changes in cellular morphology (Supplementary Figure S1c). At 2 h after treatment – before AnnV/PI positivity – HeLa cells displayed long, thick protrusions, and irregularly shaped nuclei. These changes in morphology were also verified in FSC/SSC plots: GrM-treated cells were reduced in size (FSC) but displayed increased granularity (SCC) (Supplementary Figure S1d). Using time-lapse microscopy, we tracked these morphological alterations over an 8-h time course (Supplementary Figure S1e; Supplementary Movie S1). The rapid changes in morphology preceded apoptotic blebbing. Finally, no upregulation of the stress-inducible protein CHOP24 and no accumulation of ubiquitylated proteins were detected (Supplementary Figure S1f), indicating that GrM-mediated apoptosis is not accompanied by endoplasmic reticulum stress.
Cell death induced by GrM is characterized by apoptotic hallmarks. (a) HeLa cells were treated with 1 μM GrM (M), 1 μM GrM-SA (MSA), or 100 nM GrB, and 0.5 μg/ml SLO or no SLO for 20 h at 37 °C. Metabolically active cells were quantified using the WST-1 assay, and the relative loss of viability compared with untreated (SLO-negative) cells was calculated. (b) Jurkat cells were treated with 1 μM GrM (M), 1 μM GrM-SA (MSA), or 100 nM GrB in the absence or presence of 0.1 μg/ml SLO for 16 h. DNA fragmentation was determined by PI staining and flow cytometry to assess the percentage of subG1 cells. (c) HeLa cells were treated as in panel (a) and tested for induction of apoptosis with AnnV/PI flow cytometry. Results are representative of at least three experiments. (d) HeLa cells were treated as in panel (a) for 4, 8, or 20 h and tested for induction of apoptosis with AnnV/PI flow cytometry. AnnV and/or PI-positive cells are plotted on the y axis. (e–h) HeLa cells were treated as in panel (a) and tested for (e) DNA fragmentation with TUNEL (results are representative of at least three experiments), (f) loss of mitochondrial membrane potential (ΔΨm) with DiOC6, (g) generation of ROS with CM-H2DCFDA, and (h) release of cytochrome c using flow cytometry. Bar graphs depict the mean±S.D., with *P values<0.05
GrM indirectly activates caspases
It remains unclear whether or not caspase activation is involved in GrM-mediated cell death.13, 14, 15, 20, 22 Therefore, cell-free protein extracts from HeLa cells were incubated for 4 h with increasing concentrations of GrM, in the presence or absence of the pan-caspase inhibitor zVAD-fmk. Caspase activation of a subset of the caspase family was monitored using immunoblot (Figure 2a). Whereas granzyme B (GrB) cleaved caspase-3, -8, and -9 in lysate (and indirectly activated caspase-6),25 GrM treatment did not lead to any detectable caspase activation in lysate. However, as a control, known GrM substrates nucleophosmin (NPM),13 α-tubulin,12 and FADD15 were cleaved by GrM in a caspase-independent manner (Figure 2a). Remarkably, GrM-mediated proteolysis of the previously reported GrM substrate TRAP/Hsp7521 could not be verified. Similar results were obtained in Jurkat lysate (data not shown). These data indicate that GrM cannot directly activate caspases.
GrM indirectly activates caspases-3, -6, -8, and -9 and induces caspase-8-dependent and -independent cell death. (a) 5 μg of HeLa cell lysate was incubated with the indicated concentrations of GrM (0–500 nM) or 500 nM GrB for 2 h at 37 °C in the absence (dimethyl sulfoxide (DMSO)) or presence of 100 μM z-VAD-fmk, after which caspase activation was assessed using immunoblot. The effects of GrM on known substrates TRAP,21 NPM,13 α-tubulin,12 and FADD15 were also determined. Nm23H1 served as loading control. (b) HeLa cells were treated with 1 μM GrM (M), 1 μM GrM-SA (MSA), or 100 nM GrB (B) and 0.5 μg/ml SLO in the absence (DMSO) or presence of 100 μM z-VAD-fmk for 8 h at 37 °C. Caspase activation and substrate cleavage were visualized using immunoblot with nm23H1 serving as the loading control. Known GrM substrates (NPM, α-tubulin, and FADD) were used as positive controls, of which α-tubulin cleavage was partially mediated by caspases. (c) Jurkat cells were treated for 6 h at 37 °C with 1 μM GrM, 1 μM GrM-SA, or 50 nM GrB and 0.1 μg/ml SLO. Caspase-3/-7 activation was detected using the CaspaseGlo assay. Relative light units (RLU) are depicted. (d–f) HeLa cells were treated with 1 μM GrM or 1 μM GrM-SA and 0.5 μg/ml SLO. Caspase activation (d) was detected in time using immunoblot analysis, and the kinetics of ΔΨm (DiOC6) loss (e) and cytochrome c release (f) were also determined at various time intervals (n=3, means±S.E.M.). (g) HeLa cells were treated as in panel (b), and the effects of 100 μM zVAD on GrM-induced cell death were determined using AnnV/PI flow cytometry. (h) Wild-type (wt) and caspase-8-deficient (casp-8−/−) Jurkat cells were treated with 1 μM GrM and 0.1 μg/ml SLO in the absence (DMSO) or presence of 100 μM z-VAD-fmk. Cell death induction was assessed using AnnV/PI flow cytometry
Next, HeLa cells were treated with GrM in the presence or absence of zVAD-fmk, and again, caspase activation was monitored (Figure 2b). GrB served as a control for monitoring caspase activation and directly activated caspase-3 and -8, whereas caspase-6 and -9 activation was at least partially dependent on the activity of other caspases (Figure 2b). In contrast to cell-free tumor lysates, GrM was capable of activating caspase-3, -6, -8 and -9 in living cells. This activation was inhibited by zVAD-fmk, confirming that GrM did not directly proteolyze caspases. Similar results were obtained with Jurkat cells in which GrM was delivered with SLO (data not shown) and with perforin (Supplementary Figure S1g). Caspase-3/-7 activation in GrM-treated Jurkat cells was further validated using the Caspase-Glo activity assay (Figure 2c). To study the kinetics of caspase activation in HeLa cells, caspase activation was determined at various time points following GrM treatment (Figure 2d). At 4 h after treatment, activation of caspase-3, -6 and -9 could be detected; after 6 h, caspase-8 activation was also detected (Figure 2d). Loss of mitochondrial membrane potential and the release of cytochrome c were already observed after 2 h of GrM treatment (and thus before measurable caspase activation), suggesting that targeting of mitochondria occurs relatively early in GrM-mediated cell death (Figures 2d–f).
Consistent with the zVAD-sensitivity of GrM-mediated caspase activation (Figure 2b), zVAD-fmk treatment of HeLa cells almost completely inhibited GrM-induced apoptosis (Figure 2g). Recently, Wang et al.15 demonstrated that GrM activates caspase-8 via cleavage of FADD, resulting in mitochondrial damage and initiation of the caspase cascade. In caspase-8-deficient (casp-8−/−) Jurkat cells (Supplementary Figure S2a), GrM-induced cell death was indeed reduced (∼50%) as compared with wt Jurkat cells, but it was not completely inhibited (Figure 2h). This suggests that GrM activates an alternative cell death pathway that does not rely on FADD-cleavage/caspase-8 activation. zVAD-fmk even further inhibited GrM-induced apoptosis in casp-8−/− cells (Figure 2h), whereas Bcl-2 overexpression did not (Supplementary Figure S2b). This suggests that in casp-8−/− Jurkat cells, GrM-induced cell death is independent of caspase-8 but (at least partially) relies on the indirect activation of other caspases. zVAD-fmk did not affect GrM-mediated morphological changes (Supplementary Figure S2c) and mitochondrial perturbations (Supplementary Figure S2d). Taken together, these data indicate that GrM triggers cellular morphological changes, mitochondrial perturbations, and activation of the caspase cascade that is only partially dependent on caspase-8, leading to DNA fragmentation and apoptosis.
Proteomic identification of GrM substrates in living cancer cells
To identify candidates via which GrM activates caspase-8-independent apoptosis, we compared the N-terminomes of control HeLa cells (SLO treated) with those of HeLa cells treated for 15 or 60 min with SLO and GrM (Figure 3a) to increase the overall possibility of identifying both early and late proteolytic events. In this way, 1361 N-terminal peptides were identified, 597 potentially hinting to proteolytic neo-N-termini, that is, peptides that were N-terminally trideutero-acetylated and did not start at position 1 or 2. Of these, 38 neo-N-termini were uniquely identified in (one of) the GrM-treated setups and could thus be assigned to 34 (direct or indirect) GrM substrates (Supplementary Table S1). Amino acids surrounding the scissile bond are denoted as Pn-P2-P1↓P1’-P2’-Pn’.26 Out of 38 GrM-generated neo-N-termini, 25 (65.8%) were cleaved after a P1 Leu or Met. A multiple sequence alignment of the 38 cleavage sites by iceLogo27 (Figure 3b) clearly showed the P4-P1 sequence specificity profile [K/R][V/E][P/A]L↓, which closely matches previously characterized GrM specificity profiles.10, 11, 28 TopoIIα was among the 25 GrM substrates identified upon cleavage at a P1 Leu or Met (Figure 3c). TopoIIα is a nuclear enzyme that can alter DNA topology to resolve DNA overwinding and is often highly expressed in cancers where it correlates with poor patient survival. The identified GrM cleavage site in topoIIα here, Leu1280, resides inside the first nuclear localization signal (NLS) of the enzyme (Figure 3d).
Positional proteomics-based identification of GrM-mediated topoIIα cleavage at Leu1280 in HeLa cells. (a) Schematic overview of the N-terminal positional proteomics strategy. L-Arg SILAC-labeled HeLa cells were incubated with 1 μM GrM and 0.5 μg/ml SLO for either 15 (12C6) or 60 (13C6) min or with 0.5 μg/ml SLO only for 60 min (13C615N4) at 37 °C. N-terminal COFRADIC was used to identify in vivo GrM substrates. (b) IceLogo visualization of the 38 non-redundant P10-P10’ GrM cleavage site motifs identified. Multiple sequence alignments27 of peptide substrate motifs from P10 to P10’ are given with cleavage of the substrate occurring between P1 and P1’. Statistically significant residues with a P-value threshold of≤0.05 are plotted. The amino-acid heights are indicative of their degree of conservation at the indicated position. The frequency of the amino-acid occurrence at each position in the sequence set was compared with the human Swissprot 2011_11 database.27 (c) MS/MS spectrum of the N-terminally trideutero-acetylated neo-N-terminus of topoIIα (1280AFKPIKKGKKR, Swiss-Prot accession: P11388) with the observed b- and y-type fragment ions indicated. (d) Schematic domain representation of human topoIIα. Proteomics identified GrM cleavage of topoIIα at Leu1280 in the regulatory C-terminal domain, within the first NLS. NES, nuclear export signal; TOPRIM, topoisomerase-primase domain; WHD, winged-helix domain
GrM targets topoIIα
To validate topoIIα as a GrM substrate, HeLa cells were treated with GrM in the presence or absence of zVAD-fmk. TopoIIα cleavage by GrM was confirmed by immunoblotting and was independent of caspase activation (Figure 4a). Full-length topoIIα (∼170 kDa) disappeared and the expected N-terminal fragment (∼140 kDa) was detected. In Jurkat cells treated with GrM and perforin, topoIIα was also efficiently cleaved (Supplementary Figure S1h). Expression levels of topoIIα, however, did not change in GrM-treated Jurkat cells (Supplementary Figure S2f). Similarly, in cell-free lysates, topoIIα was cleaved with high efficiency: already within 30 min, 10 nM of GrM had completely proteolyzed topoIIα (Figure 4b, upper panel). GrM cleaved purified recombinant topoIIα, validating topoIIα as a direct GrM substrate (Figure 4c). Finally, Cos7 cells were transfected with eGFP-linked topoIIα and co-cultured with increasing effector:target (E:T) ratios of KHYG1 NK cells,29 which are known to express GrM.30 Using immunoblotting with an anti-eGFP antibody, cleavage of eGFP-topoIIα inside target cells was observed (Figure 4d), indicating that topoIIα is a physiological substrate during NK-cell-mediated tumor cell killing. No cleavage fragment was detected, which may be due to further processing and degradation of cleaved topoIIα during NK-cell-mediated cytotoxicity.
GrM efficiently and directly cleaves topoIIα. (a) HeLa cells were incubated with 1 μM GrM or 1 μM GrM-SA, and 0.5 μg/ml SLO in the absence (dimethyl sulfoxide (DMSO)) or presence of 100 μM z-VAD-fmk for 6 h at 37 °C. TopoIIα cleavage was determined using immunoblot analysis. Hsp90β was used as loading control. (b) 10 μg HeLa cell lysate was incubated at 37 °C with indicated concentrations of GrM or 50 nM GrM-SA for 30 min or with 5 nM GrM for the indicated lengths of time or 60 min with 5 nM GrM-SA. TopoIIα cleavage was determined using immunoblot analysis. β-Tubulin was used as a loading control. (c) Twenty units of purified recombinant human topoIIα were incubated with 100 nM GrM or GrM-SA for 30 min at 37 °C. Cleavage was determined using immunoblot. (d) Cos7 cells were transiently transfected with enhanced green fluorescent protein (eGFP)-topoIIα and co-cultured with increasing effector:target (E:T) ratios of NK cells (KHYG1) for 4 h at 37 °C. Cleavage of eGFP-topoIIα was determined using immunoblot for eGFP. An arrow is used to indicate the topoIIα cleavage fragment
GrM disrupts topoIIα nuclear localization
GrM cleaved topoIIα at Leu1280, which is located in its C-terminal regulatory domain within the first NLS (Figure 3d). Previous studies have shown that the N-terminal domain of topoIIα can function in vitro without its C-terminal regulatory domain,31, 32 suggesting that topoIIα functionality should not necessarily be affected by GrM cleavage. Human DNA topoIIα regulates DNA topology and can relax supercoiled DNA. An in vitro DNA relaxation assay was performed to study the effect of GrM cleavage on topoIIα functioning. Recombinant topoIIα was first cleaved by GrM (Figure 4c), after which supercoiled pUC18 DNA was added. As expected, topoIIα fragments rendered by GrM cleavage were still capable of relaxing supercoiled DNA (Figure 5a), suggesting that the catalytic topoIIα domains remain active following GrM cleavage.
GrM disrupts topoIIα nuclear localization. (a) Twenty units of recombinant topoIIα were treated with 100 nM GrM or GrM-SA for 30 min at 37 °C (see also Figure 4c). These samples were incubated with 500 ng of supercoiled pUC18 plasmid for 30 min at 37 °C. pUC18 DNA relaxation was assessed on a 0.7% agarose gel. Supercoiled and relaxed pUC18 are indicated. (b) HeLa cells were transfected with full-length eGFP-topoIIα or topoIIα fragments that mimic GrM cleavage, that is, the N-terminal eGFP-topoIIα 1–1280 and C-terminal eGFP-topoIIα 1280–1531 fragments. Expression of the fragments was verified by immunoblotting for GFP. β-Tubulin was used as a loading control. (c) Cells were transfected as in panel (b), and the cell viability was determined using PI staining. (d) Cells were transfected as in panel (b), and H2B-mCherry was co-transfected to stain nuclei. Localization was determined by fluorescent microscopy. (e) HeLa cells were incubated with 1 μM GrM and 0.5 μg/ml SLO for 30 min or mock treated, after which they were fractionated. Whole cell (WC) lysate, cytosolic (C), and nuclear (N) fractions were immunoblotted for topoIIα. An arrow is used to indicate the topoIIα cleavage fragment. (f) Semi-quantitative analysis was performed on the immunoblots of cells prepared in panel (e), and ratios of fragment/full-length topoIIα are plotted for WC, C, and N fractions. (g and h) HeLa cells were pretreated with 12 nM LMB for 2 h, after which they were treated with 0.5 μg/ml SLO in the absence or presence of 1 μM GrM. After a 30-min incubation, samples were immunoblotted for topoIIα (g). In addition, GrM-treated samples were fractionated and immunoblotted for topoIIα and for TAF5 and Hsp90β (as controls for the nuclear and cytosolic fractions, respectively) (h). (i) HeLa cells were treated with 1 μM GrM in the presence or absence of 0.5 μg/ml SLO, followed by a 30-min incubation. Cells were then washed, fixed, stained, and GrM localization was visualized using confocal microscopy. DAPI (4,6-diamidino-2-phenylindole) was used to stain the nuclei
Although GrM cleavage did not affect topoIIα functionality in vitro, the generated fragments could be proapoptotic in living cells. Full-length eGFP-linked topoIIα was expressed in HeLa cells, as well as the N- and C-terminal topoIIα fragments that mimic GrM cleavage (eGFP-topoIIα 1–1280 and eGFP-topoIIα 1280–1531, respectively). Expression of the fragments (Figure 5b) did not affect cell viability (Figure 5c). However, whereas full-length topoIIα and the C-terminal fragment containing the NLS clearly localized to the nuclei of transfected cells, the N-terminal fragment bearing the topoIIα catalytic domains and nuclear export signals33 localized to the cytoplasm (Figure 5d). Expression of protein lacking a NLS is not identical to the removal of a localization signal of a nuclear protein. Therefore, the effects of GrM-mediated cleavage of endogenous nuclear topoIIα were investigated. HeLa cells were treated with GrM, fractionated, and topoIIα localization was monitored (Figures 5e and f). As expected, full-length topoIIα in untreated control cells was only present in the nuclear fraction. In GrM-treated cells, however, the N-terminal topoIIα fragment relocalized to the cytoplasmic fraction, indicative of a cleavage-induced nuclear exit of this fragment. Pretreatment of cells with the nuclear export inhibitor leptomycin B (LMB) (Supplementary Figure S2e), which prevents topoIIα nuclear/cytoplasmic shuttling,33 did not prevent GrM-mediated cleavage of topoIIα nor did it prevent the nuclear exit of the cleavage fragment (Figures 5g and h). This suggests that the cleavage fragment may exit the nucleus via alternative mechanisms, for example, as a result of nuclear disintegration. For GrA and GrB, nuclear localization has been described previously.34, 35, 36, 37, 38 For GrM, however, nuclear localization remains unknown. To test this, GrM localization was determined after intracellular delivery using immunofluorescence and confocal microscopy (Figure 5i). GrM entered the nucleus, already within 30 min after GrM delivery. Combined, these data indicate that GrM enters the nucleus where it cleaves topoIIα, resulting in nuclear exit of the N-terminal fragment, impairing topoIIα function.
TopoIIα depletion phenotypically resembles GrM-induced apoptosis
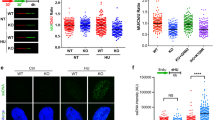
As GrM abolishes nuclear topoIIα function, the effects of genetic topoIIα depletion were compared with GrM-mediated cell death. TopoIIα was conditionally depleted from HTETOP cells by adding doxycycline (dox).39 At 48 h after administration of dox, topoIIα levels were reduced to undetectable levels (Figure 6a, inset; Supplementary Figure S2g). To examine general cell behavior, topoIIα-depleted cells were monitored real-time by xCELLigence over a period of 96 h (Figures 6a and b). Whereas the cell index (CI) in control (–dox) cells increased over time due to cell proliferation, topoIIα-depleted cells (+dox) were markedly impaired. Decreased CI values correlated with lower numbers of adherent topoIIα-depleted HTETOP cells as visualized by light microscopy (Supplementary Figure S2h). This decrease in adherent cells could be due to decreased proliferation, increased apoptosis, or a combination. TopoIIα depletion is known to result in G2/M arrest and increased polyploidy,39 which was confirmed using cell cycle analysis (Figure 6c). G2/M arrest was also observed within 3 days after topoIIα depletion (data not shown). If topoIIα function is impaired by GrM, one would expect that GrM also triggers a G2/M arrest in tumor cells. Indeed, the topoIIα-depleted cell cycle profile was remarkably similar to that of GrM-treated Jurkat cells, which also arrested in G2/M (Figure 6d). Likewise, G2/M arrest was also seen in GrM-treated casp-8−/− Jurkat cells, indicating that this arrest is independent of caspase-8 activation upon FADD cleavage (Figure 6e). In addition, topoIIα-depleted cells also displayed increased rates of caspase-dependent apoptosis (Figure 6f). Caspase-3/-7 was activated in topoIIα-depleted cells (Figure 6g), and activation of caspase-3 was further confirmed by immunoblotting (Figure 6h). Finally, caspase-independent perturbation of the mitochondrial membrane potential was observed in topoIIα-depleted cells (Figure 6i), suggesting that this contributes to the induction of apoptosis. Thus, topoIIα depletion in tumor cells phenotypically resembles GrM-induced apoptosis, including G2/M cell cycle arrest, mitochondrial perturbations, and caspase activation.
TopoIIα depletion phenotypically resembles GrM-induced apoptosis. (a) Treatment with 1 μg/ml dox for 48 h depleted topoIIα in HTETOP cells as determined using immunoblot (inset). Hsp90β was used as loading control. HTETOP cells depleted of topoIIα for 48 h were seeded in the xCELLigence system, and CI was monitored in time (every 30 min for 96 h). Medium was refreshed after 48 h. Graph depicts n=4 with mean±S.D. of a single experiment that is representative of three independent experiments. (b) The increase in CI (slope) between 12 and 96 h after seeding was determined for three independent xCELLigence experiments. (c) The cell cycle profile of HTETOP cells that had been depleted of topoIIα was determined 5 days after the seeding of the cells using PI-cell cycle flow cytometry. (d and e) Both wild-type (wt) and casp-8−/− Jurkat cells were treated in the presence of z-VAD-fmk with 1 μM GrM or GrM-SA and 0.1 μg/ml SLO for 24 h, after which cell cycle distributions were determined using PI-cell cycle flow cytometry. (f) Cell death of topoIIα-depleted HTETOPs grown in the absence (dimethyl sulfoxide (DMSO)) or presence of 100 μM z-VAD-fmk was determined at different time points after seeding using AnnV/PI flow cytometry. (g) Caspase-3/-7 activity was assessed in topoIIα-depleted HTETOPs at 48 h after seeding using the CaspaseGlo assay. Nm23H1 served as a loading control. (h) Cells were seeded as in panel (g), and caspase-3 activation was detected using immunoblot. GAPDH (glyceraldehyde 3-phosphate dehydrogenase) was used as a loading control. (i) Loss of mitochondrial membrane potential was measured at different time points after seeding of topoIIα-depleted HTETOPs grown in the absence (DMSO) or presence of 100 μM z-VAD-fmk. In all experiments, medium (supplemented with dox) was refreshed every 48 h
Discussion
Granzymes are important effector molecules in the cellular immune response against tumor and virus-infected cells. Although the literature clearly describes GrM as a significant inducer of cell death, the general characteristics of cell death induction remain controversial and unclear. We have therefore comprehensively characterized the GrM-mediated apoptotic phenotype (Figures 1 and 2). GrM triggered classical hallmarks of apoptosis, including the initial appearance of AnnV single-positive (early apoptotic) cells, followed by the appearance of AnnV/PI double-positive (late apoptotic/necrotic) cells. This is in agreement with some studies20, 22 but in contradiction with other studies that describe only the formation of AnnV and PI double-positive cells.13, 14 Kelly et al.14 find no evidence of DNA fragmentation after GrM treatment, whereas Lu et al.22 do. In our study, GrM also induced DNA fragmentation. Similarly, Kelly et al.14 do not detect any changes in mitochondrial membrane potential or release of cytochrome c, whereas GrM is found to disturb the mitochondria in studies by Hua et al.,21 Wang et al.,15 and in the present study. Furthermore, we confirmed that GrM treatment results in ROS generation, as first described by Hua et al.21 Discrepancies have also been found concerning the activation of caspases upon GrM treatment. Lu et al.22 describe that GrM can directly cleave pro-caspase-3 in lysates; this has, however, been refuted by studies done by Kelly et al.,14 Cullen et al.,13 Bovenschen et al.,12 and the present study. Although we did not find caspase activation by GrM in lysates, we did find caspase activation in living cells. This observation is in agreement with studies by Hu et al.20 and Wang et al.,15 but in disagreement with studies by Kelly et al.14 and Cullen et al.13 However, in the latter study, caspase activation has been monitored in the presence of zVAD-fmk, which means that indirect activation of the caspases by GrM would not have been detected, as shown here to be the case in zVAD-fmk pretreated cells (Figure 2b).13 The presence or absence of (indirect) caspase activation is likely an important determinant of the GrM cell death phenotype: caspases are known to activate DNases, which leads to positive TUNEL-staining, and are also known to lead to cytochrome c release and AnnV-positivity. Therefore, the inconsistency in caspase activation between studies may lie at the base of all other discrepancies. Participation of caspases in GrM-mediated tumor cell death is further supported by the findings that GrM targets hnRNP K23 and survivin,20 which stabilize the caspase-inhibitor XIAP.20, 40 This suggests that GrM coordinates induction of the caspase cascade by both inducing caspase activation and reducing caspase inhibition. Next to the general apoptotic hallmarks described above, we showed for the first time that GrM rapidly triggered G2/M cell cycle arrest in tumor cells (Figure 6). Thus, we propose that GrM triggers classical hallmarks of apoptosis with mitochondrial damage, caspase activation, and cell cycle arrest.
To date, FADD is the only univocally proven GrM substrate that has a direct role in GrM-mediated cell death.15 Cleavage of human FADD by GrM promotes pro-caspase-8 recruitment and activation and subsequent initiation of the caspase cascade.15 FADD-deficient tumor cells, however, are still partially sensitive to GrM.15 Consistently, we showed that GrM still induced caspase-dependent apoptosis in casp-8−/− cells (Figure 2h). Interestingly, neither caspase-8-deficiency nor chemical caspase inhibition blocked GrM-induced G2/M cell cycle arrest, which may precede apoptosis.41 Thus, these data indicate that GrM triggers caspase-dependent cell death via at least one other mechanism that is independent of FADD/caspase-8, likely involving G2/M cell cycle arrest. To elucidate this pathway, we used positional proteomics in HeLa cells and identified DNA topoIIα as a novel GrM substrate, a nuclear enzyme essential in proliferating cells where it assists in chromosome condensation, segregation, and replication.42 Overexpression of topoIIα correlates with poor prognosis in breast, ovarian, and peritoneum cancer and oligodendrogliomas.43, 44, 45 TopoIIα is a current anti-cancer target for chemotherapeutics such as etoposide and doxorubicin46 and depletion of topoIIα leads to G2/M cell cycle arrest and apoptosis39 (Figure 6). GrM cleaved topoIIα after Leu1280 (Supplementary Table S1), thereby separating the topoIIα functional domains from its NLSs. Indeed, GrM cleavage led to nuclear exit of the N-terminal fragment of topoIIα that harbors the catalytic domains (Figure 5), thereby rendering the protein nonfunctional. Interestingly, the GrM apoptotic phenotype, including G2/M arrest, mitochondrial perturbation, and caspase activation (Figures 1 and 2), resembled the phenotype of topoIIα depletion (Figure 6). Although the relative importance of this pathway remains to be investigated, these data suggest that inactivation of topoIIα by GrM contributes to G2/M arrest and caspase-dependent apoptosis (Figure 7).
Schematic model of GrM-induced cell death. Perforin facilitates entry of GrM into the target cell via pore formation. Upon entry, GrM can cleave FADD,15 resulting in the recruitment and auto-processing of caspase-8, and subsequently activation of caspase-3 – either directly or via Bid-mediated targeting of the mitochondria and apoptosome formation. In addition, GrM can cleave nuclear topoIIα, resulting in the formation of a C-terminal fragment that bears the NLS and remains in the nucleus, and an N-terminal fragment, which bears the catalytic domains, and translocates to the cytosol. The depletion of functional topoIIα in the nucleus results in G2/M cell cycle arrest, which also leads to mitochondrial perturbations and caspase-dependent cell death, via a yet to be identified mechanism. Furthermore, GrM can also cleave and inactivate survivin20 and hnRNP K,23 resulting in a decrease in the levels of caspase inhibitors, thereby facilitating caspase-induced apoptosis
Although previous work has hinted towards a role of GrB in targeting mitosis-regulating kinases,47, 48 we present – to our knowledge – the first evidence of cell cycle arrest induction by a granzyme. GrM-induced cell cycle arrest occurred independently of caspases (Figure 6). This raises the interesting possibility that if the immune system cannot kill a tumor cell – for example, because it expresses certain inhibitors of apoptosis,49 such as Bcl2, XIAP,50 or the GrB inhibitor SERPINB951, 52, 53, 54 – GrM can still trigger apoptosis or cell cycle arrest to reduce tumor growth. Alternatively, GrM-induced effects on DNA topology may also contribute to help dispose of the cell corpse and facilitate DNA degradation post mortem.
TopoIIα belongs to a general family of isomerases that act on the topology of DNA, for example, during DNA replication and cell division.42 This family also includes topoisomerases I and IIβ. Interestingly, proteolytic targeting of topoisomerases during apoptosis has been observed previously. GrB cleaves topoI, which is involved in breaking and rejoining of DNA single strands,55 and caspases can cleave topoisomerases I, IIα, and IIβ56, 57, 58 at multiple sites.56 Finally, we have preliminary data showing that in tumor cell lysate GrA and GrB can also cleave topoIIα in close proximity to the GrM cleavage site (data not shown). These findings suggest that targeting of topoisomerases is a common phenomenon during apoptosis, although the effects of these cleavage events on cell cycle progression remain unknown. In this context, we and others have previously established that GrM and GrB can cleave α-tubulin, resulting in the stabilization of microtubule polymers12, 59, 60 in a manner that resembles the effects of the anti-cancer drug paclitaxel (Taxol). Paclitaxel is known to induce G2/M arrest, and granzyme-mediated cleavage of α-tubulin may therefore also contribute to the induction of a G2/M arrest in GrM-treated cells. The relative contribution of these substrates and potentially other pathways to the induction of granzyme-induced cell cycle arrest and cytotoxicity remains to be elucidated. Whether granzymes other than GrM also induce cell cycle arrest remains an interesting question that deserves further study.
Materials and Methods
Cells
HeLa and Cos7 cells were grown in DMEM supplemented with 10% fetal calf serum (FCS), 100 units/ml penicillin and 100 μg/ml streptomycin (pen/strep; Life Technologies, Grand Island, NY, USA). HTETOP cells39 were grown in the same medium, with L-glutamine (2 mM) and non-essential amino acids (20 ml/500 ml; Life Technologies). To deplete topoIIα in HTETOP cells, medium was supplemented with 1 μg/ml dox and refreshed every 2–3 days. Jurkat and caspase-8-deficient Jurkat cells (clone I 9.2, ATCC) were maintained in RPMI-1640 medium with 10% FCS, pen/strep, and 2 g/l sodium bicarbonate. The NK cell line KHYG1 (Japan Health Sciences Foundation, Tokyo, Japan; JCRB0156) was maintained in the same medium with 50 ng/ml recombinant human interleukin-2 (Tebu-Bio, Heerhugowaard, the Netherlands). Cells were transfected using polyethylenimine (Polysciences, Eppelheim, Baden-Württemberg, Germany). For combined fractional diagonal chromatography (COFRADIC), HeLa cells were grown in Glutamax-containing SILAC (stable isotope labeling by amino acids in cell culture) DMEM (Life Technologies) with 10% dialyzed FCS (Life Technologies) and pen/strep, containing natural, 13C6- or 13C615N4-labeled L-arginine (80 μM; Cambridge Isotope Laboratories, Tewksbury, MA, USA). Cells were cultured for six population doublings (8 days) for complete incorporation of labeled arginine. For leptomycin B (LMB) pretreatment, cells were incubated with 12 nM LMB (Sigma-Aldrich, St. Louis, MO, USA) for 2 h. For caspase inhibition, 100 μM zVAD-fmk (Enzo Life Sciences, Farmingdale, NY, USA) was used. Cells were fractionated as described.61
Proteins
Granzyme production and granzyme treatment with SLO (Aalto Bio Reagents, Dublin, Ireland) and human perforin (Enzo Life Sciences) were performed as described previously.10, 12 Briefly, human GrB, GrM, and its catalytically inactive mutant GrM-SA, in which the Ser195 residue has been replaced with Ala, were produced in Pichia pastoris and purified using cation-exchange chromatography. Catalytic activity was verified using synthetic chromogenic substrates.
Antibodies and cloning
Antibodies were anti-acetyl-histone-H3 (06–599; Upstate, Merck Millipore, Billerica, MA, USA), anti-α-tubulin (B-5–1–2; Sigma, St. Louis, MO, USA), anti-β-tubulin (Tub2.1; Sigma), anti-caspase 3 (H-277, Santa Cruz Biotechnology, Santa Cruz, CA, USA), anti-Asp175 cleaved caspase 3 (Cell Signaling Technologies, Boston, MA, USA), anti-caspase 6 (3E8; MBL International Corporation, Woburn, MA, USA), anti-caspase 8 (5F7; MBL International Corporation), anti-Asp315 cleaved caspase 9 (Cell Signaling Technologies), anti-CHOP (L63F7; Cell Signaling Technologies), anti-Erk-2 (C14, Santa Cruz Biotechnology), anti-GAPDH (6C5; Abcam, Cambridge, UK), anti-GFP (clones 7.1 and 13.1, Roche, Basel, Switzerland), anti-GrM (4B2G4),8 anti-Hsp90β (K3701; Enzo Life Sciences), anti-nm23H1 (C-20; Santa Cruz Biotechnology), anti-NPM (FC-61991; Life Technologies), anti-topoIIα (Ki-S1, DAKO, Glostrup, Denmark), anti-TRAP (TR-1A, Abcam), and HRP-conjugated anti-ubiquitin (Enzo Life Sciences). Fragments of topoIIα cDNA were cloned in pCDNA3.1+ (Life Technologies) using pBL-eGFP-topoIIα39 as a template.
Viability assays and flow cytometry
WST-1 assays were performed as described by the manufacturer (Roche). For AnnV (Life Technologies) and PI (Sigma) analysis, cells were stained for 15 min in 140 mM NaCl, 4 mM KCl, 0.75 mM MgCl2, 1.5 mM CaCl2, and 10 mM HEPES pH 7.4. Flow cytometry was performed on a FACSCalibur with CellQuest Pro software (BD Biosciences, Franklin Lakes, NJ, USA). AnnV- and PI-negative cells were considered viable. Cells were stained with 20 nM DiOC6 (Tebu-Bio) for 15 min at 37 °C. TUNEL (Merck Millipore), CM-H2DCFDA (Life Technologies), and cytochrome c (VWR Omnilabo, Radnor, PA, USA) staining were performed according to the manufacturers’ protocols. For cell cycle analysis, cells were washed in PBS and fixed in 70% ice-cold ethanol for >15 min at 4 °C. Cells were washed with 0.1% Triton-X100 in PBS and stained in 0.1% Triton-X100, 15 μg/ml RNase A, and 100 μg/ml PI in PBS for 2 h. Cells were plotted on FL2-A and FL2-W to discriminate between individual cells and aggregates. Caspase activation was determined using the CaspaseGlo assay (Promega, Madison, WI, USA).
Fluorescence microscopy
Untransfected cells (living or stained with Giemsa) and cells transfected with fluorescently-tagged constructs (H2B-mCherry and eGFP-topoIIα variants) were directly imaged by inverted fluorescence microscopy (Leica DMI4000b, Leica DFC300 FX camera, Leica Microsystems, Wetzlar, Germany). Confocal microscopy was performed on a Leica LSM700 (Leica Microsystems).
xCELLigence
The xCELLigence system (Roche) measures electrical impedance (represented in the CI) across micro-electrodes integrated on the bottom of 16-well plates, which provides quantitative information about the overall status of cells (cell number, viability, adherence, morphology). Untreated or topoIIα-depleted HTETOP cells were seeded at 1250 cells/well. Dox was refreshed after 48 h. CI values were measured every 30 min for 96 h.
In vitro topoIIα activity assay
Recombinant topoIIα (Affymetrix/USB, Santa Clara, CA, USA; 20 units) was incubated with 100 nM GrM or GrM-SA for 30 min at 37 °C. Subsequently, 500 ng of supercoiled plasmid pUC18 was added for 30 min at 37 °C, electrophorized using a 0.7% agarose gel, and stained with Midori Green Advance DNA stain (Biolegio, Nijmegen, the Netherlands).
N-terminal COFRADIC analysis
L-arginine labeled HeLa cells were treated with SLO for 60 min (control setup; 13C615N4-Arg) or with SLO and 1 μM GrM for 15 min (12C6-Arg) or 60 min (13C6-Arg) (Figure 3a). Cells were lysed on ice for 5 min in lysis buffer containing 100 mM NaCl, 1% CHAPS, 0.5 mM EDTA, and 50 mM sodium phosphate buffer (pH 7.5) and freshly added protease inhibitor cocktail, EDTA-free. Cellular debris was spun down for 1 min at 18 000 g. Protein concentration in the supernatant was determined by the Bradford assay. Guanidinium hydrochloride was added to a final concentration of 4 M to inactivate GrM and denature all proteins. Before mixing equal amounts of samples, proteins were reduced and alkylated using TCEP.HCl (Tris carboxyethyl phosphine.HCl; 1 mM) and iodoacetamide (2 mM) respectively, for 1 h at 30 °C. N-terminal COFRADIC was performed as described previously.62 The protein mixture was digested overnight at 37 °C with sequencing-grade, modified trypsin (Promega) (enzyme/substrate of 1/100, w/w). Liquid chromatography-mass spectrometry/mass spectrometry (LC-MS/MS) analysis and data processing were performed as described previously63 (Supplementary Materials and Methods).
Statistical analysis
Unless otherwise indicated, data are depicted as mean±S.D. of at least three independent experiments. Statistical analyses were performed using independent samples’ t-test. *P<0.05 was considered statistically significant.
Abbreviations
- AnnV:
-
AnnexinV-fluos
- Casp-8−/−:
-
caspase-8 deficient
- CI:
-
cell index
- COFRADIC:
-
combined fractional diagonal chromatography
- dox:
-
doxycycline
- FADD:
-
Fas-associated protein with death domain
- FCS:
-
fetal calf serum
- GrB:
-
granzyme B
- GrM:
-
granzyme M
- GrM-SA:
-
inactive GrM mutant in which the catalytic site Ser residue has been mutated to an Ala residue
- LC-MS/MS:
-
liquid chromatography-mass spectrometry/mass spectrometry
- NK:
-
natural killer
- NLS:
-
nuclear localization signal
- NPM:
-
nucleophosmin
- PI:
-
propidium iodide
- ROS:
-
reactive oxygen species
- SILAC:
-
stable isotope labeling by amino acids in cell culture
- SLO:
-
streptolysin O
- TCEP:
-
Tris carboxyethyl phosphine
- topoIIα:
-
topoisomerase II alpha
- TUNEL:
-
TdT dUTP nick-end labeling
- wt:
-
wild type
References
Lieberman J . Anatomy of a murder: how cytotoxic T cells and NK cells are activated, develop, and eliminate their targets. Immunol Rev 2010; 235: 5–9.
Cullen SP, Brunet M, Martin SJ . Granzymes in cancer and immunity. Cell Death Differ 2010; 17: 616–623.
Grossman WJ, Revell PA, Lu ZH, Johnson H, Bredemeyer AJ, Ley TJ . The orphan granzymes of humans and mice. Curr Opin Immunol 2003; 15: 544–552.
Anthony DA, Andrews DM, Watt SV, Trapani JA, Smyth MJ . Functional dissection of the granzyme family: cell death and inflammation. Immunol Rev 2010; 235: 73–92.
Bovenschen N, Kummer JA . Orphan granzymes find a home. Immunol Rev 2010; 235: 117–127.
de Koning PJ, Kummer JA, Bovenschen N . Biology of granzyme M: a serine protease with unique features. Crit Rev Immunol 2009; 29: 307–315.
Bade B, Boettcher HE, Lohrmann J, Hink-Schauer C, Bratke K, Jenne DE et al. Differential expression of the granzymes A, K and M and perforin in human peripheral blood lymphocytes. Int Immunol 2005; 17: 1419–1428.
de Koning PJ, Tesselaar K, Bovenschen N, Colak S, Quadir R, Volman TJ et al. The cytotoxic protease granzyme M is expressed by lymphocytes of both the innate and adaptive immune system. Mol Immunol 2010; 47: 903–911.
Smyth MJ, Sayers TJ, Wiltrout T, Powers JC, Trapani JA . Met-ase: cloning and distinct chromosomal location of a serine protease preferentially expressed in human natural killer cells. J Immunol 1993; 151: 6195–6205.
de Poot SA, Westgeest M, Hostetter DR, Van Damme P, Plasman K, Demeyer K et al. Human and mouse granzyme M display divergent and species-specific substrate specificities. Biochem J 2011; 437: 431–442.
Mahrus S, Craik CS . Selective chemical functional probes of granzymes A and B reveal granzyme B is a major effector of natural killer cell-mediated lysis of target cells. Chem Biol 2005; 12: 567–577.
Bovenschen N, de Koning PJ, Quadir R, Broekhuizen R, Damen JM, Froelich CJ et al. NK cell protease granzyme M targets alpha-tubulin and disorganizes the microtubule network. J Immunol 2008; 180: 8184–8191.
Cullen SP, Afonina IS, Donadini R, Luthi AU, Medema JP, Bird PI et al. Nucleophosmin is cleaved and inactivated by the cytotoxic granule protease granzyme M during natural killer cell-mediated killing. J Biol Chem 2009; 284: 5137–5147.
Kelly JM, Waterhouse NJ, Cretney E, Browne KA, Ellis S, Trapani JA et al. Granzyme M mediates a novel form of perforin-dependent cell death. J Biol Chem 2004; 279: 22236–22242.
Wang S, Xia P, Shi L, Fan Z . FADD cleavage by NK cell granzyme M enhances its self-association to facilitate procaspase-8 recruitment for auto-processing leading to caspase cascade. Cell Death Differ 2012; 19: 605–615.
Schiffer S, Letzian S, Jost E, Mladenov R, Hristodorov D, Huhn M et al. Granzyme M as a novel effector molecule for human cytolytic fusion proteins: CD64-specific cytotoxicity of Gm-H22(scFv) against leukemic cells. Cancer Lett 2013 doi:10.1016/j.canlet.2013.08.005.
Pao LI, Sumaria N, Kelly JM, van Dommelen S, Cretney E, Wallace ME et al. Functional analysis of granzyme M and its role in immunity to infection. J Immunol 2005; 175: 3235–3243.
Pegram HJ, Haynes NM, Smyth MJ, Kershaw MH, Darcy PK . Characterizing the anti-tumor function of adoptively transferred NK cells in vivo . Cancer Immunol Immunother 2010; 59: 1235–1246.
de Poot SA, Lai KW, Hovingh ES, Bovenschen N . Granzyme M cannot induce cell death via cleavage of mouse FADD. Apoptosis 2013; 18: 533–534.
Hu D, Liu S, Shi L, Li C, Wu L, Fan Z . Cleavage of survivin by Granzyme M triggers degradation of the survivin-X-linked inhibitor of apoptosis protein (XIAP) complex to free caspase activity leading to cytolysis of target tumor cells. J Biol Chem 2010; 285: 18326–18335.
Hua G, Zhang Q, Fan Z . Heat shock protein 75 (TRAP1) antagonizes reactive oxygen species generation and protects cells from granzyme M-mediated apoptosis. J Biol Chem 2007; 282: 20553–20560.
Lu H, Hou Q, Zhao T, Zhang H, Zhang Q, Wu L et al. Granzyme M directly cleaves inhibitor of caspase-activated DNase (CAD) to unleash CAD leading to DNA fragmentation. J Immunol 2006; 177: 1171–1178.
van Domselaar R, Quadir R, van der Made AM, Broekhuizen R, Bovenschen N . All human granzymes target hnRNP K that is essential for tumor cell viability. J Biol Chem 2012; 287: 22854–22864.
Oyadomari S, Mori M . Roles of CHOP/GADD153 in endoplasmic reticulum stress. Cell Death Differ 2004; 11: 381–389.
Adrain C, Murphy BM, Martin SJ . Molecular ordering of the caspase activation cascade initiated by the cytotoxic T lymphocyte/natural killer (CTL/NK) protease granzyme B. J Biol Chem 2005; 280: 4663–4673.
Schechter I, Berger A . On the size of the active site in proteases. I. Papain. Biochem Biophys Res Commun 1967; 27: 157–162.
Colaert N, Helsens K, Martens L, Vandekerckhove J, Gevaert K . Improved visualization of protein consensus sequences by iceLogo. Nat Methods 2009; 6: 786–787.
Mahrus S, Kisiel W, Craik CS . Granzyme M is a regulatory protease that inactivates proteinase inhibitor 9, an endogenous inhibitor of granzyme B. J Biol Chem 2004; 279: 54275–54282.
Yagita M, Huang CL, Umehara H, Matsuo Y, Tabata R, Miyake M et al. A novel natural killer cell line (KHYG-1) from a patient with aggressive natural killer cell leukemia carrying a p53 point mutation. Leukemia 2000; 14: 922–930.
Suck G, Branch DR, Smyth MJ, Miller RG, Vergidis J, Fahim S et al. KHYG-1, a model for the study of enhanced natural killer cell cytotoxicity. Exp Hematol 2005; 33: 1160–1171.
Dickey JS, Osheroff N . Impact of the C-terminal domain of topoisomerase IIalpha on the DNA cleavage activity of the human enzyme. Biochemistry 2005; 44: 11546–11554.
Jensen S, Andersen AH, Kjeldsen E, Biersack H, Olsen EH, Andersen TB et al. Analysis of functional domain organization in DNA topoisomerase II from humans and Saccharomyces cerevisiae . Mol Cell Biol 1996; 16: 3866–3877.
Turner JG, Engel R, Derderian JA, Jove R, Sullivan DM . Human topoisomerase IIalpha nuclear export is mediated by two CRM-1-dependent nuclear export signals. J Cell Sci 2004; 117 (Pt 14): 3061–3071.
Shi L, Mai S, Israels S, Browne K, Trapani JA, Greenberg AH et al. Granzyme B (GraB) autonomously crosses the cell membrane and perforin initiates apoptosis and GraB nuclear localization. J Exp Med 1997; 185: 855–866.
Trapani JA, Browne KA, Smyth MJ, Jans DA . Localization of granzyme B in the nucleus. A putative role in the mechanism of cytotoxic lymphocyte-mediated apoptosis. J Biol Chem 1996; 271: 4127–4133.
Jans DA, Jans P, Briggs LJ, Sutton V, Trapani JA . Nuclear transport of granzyme B (fragmentin-2). Dependence of perforin in vivo and cytosolic factors in vitro . J Biol Chem 1996; 271: 30781–30789.
Jans DA, Briggs LJ, Jans P, Froelich CJ, Parasivam G, Kumar S et al. Nuclear targeting of the serine protease granzyme A (fragmentin-1). J Cell Sci 1998; 111 (Pt 17): 2645–2654.
Blink EJ, Jiansheng Z, Hu W, Calanni ST, Trapani JA, Bird PI et al. Interaction of the nuclear localizing cytolytic granule serine protease granzyme B with importin alpha or beta: modulation by the serpin inhibitor PI-9. J Cell Biochem 2005; 95: 598–610.
Carpenter AJ, Porter AC . Construction, characterization, and complementation of a conditional-lethal DNA topoisomerase IIalpha mutant human cell line. Mol Biol Cell 2004; 15: 5700–5711.
Xiao Z, Ko HL, Goh EH, Wang B, Ren EC . hnRNP K suppresses apoptosis independent of p53 status by maintaining high levels of endogenous caspase inhibitors. Carcinogenesis 2013; 34: 1458–1467.
Andersen JL, DeHart JL, Zimmerman ES, Ardon O, Kim B, Jacquot G et al. HIV-1 Vpr-induced apoptosis is cell cycle dependent and requires Bax but not ANT. PLoS Pathog 2006; 2: e127.
Vos SM, Tretter EM, Schmidt BH, Berger JM . All tangled up: how cells direct, manage and exploit topoisomerase function. Nat Rev Mol Cell Biol 2011; 12: 827–841.
Depowski PL, Rosenthal SI, Brien TP, Stylos S, Johnson RL, Ross JS . Topoisomerase IIalpha expression in breast cancer: correlation with outcome variables. Mod Pathol 2000; 13: 542–547.
Costa MJ, Hansen CL, Holden JA, Guinee D Jr . Topoisomerase II alpha: prognostic predictor and cell cycle marker in surface epithelial neoplasms of the ovary and peritoneum. Int J Gynecol Pathol 2000; 19: 248–257.
Miettinen HE, Jarvinen TA, Kellner U, Kauraniemi P, Parwaresch R, Rantala I et al. High topoisomerase IIalpha expression associates with high proliferation rate and and poor prognosis in oligodendrogliomas. Neuropathol Appl Neurobiol 2000; 26: 504–512.
Nitiss JL . Targeting DNA topoisomerase II in cancer chemotherapy. Nat Rev Cancer 2009; 9: 338–350.
Chen G, Shi L, Litchfield DW, Greenberg AH . Rescue from granzyme B-induced apoptosis by Wee1 kinase. J Exp Med 1995; 181: 2295–2300.
Shi L, Chen G, He D, Bosc DG, Litchfield DW, Greenberg AH . Granzyme B induces apoptosis and cyclin A-associated cyclin-dependent kinase activity in all stages of the cell cycle. J Immunol 1996; 157: 2381–2385.
Altieri DC . Survivin and IAP proteins in cell-death mechanisms. Biochem J 2011; 430: 199–205.
Goping IS, Barry M, Liston P, Sawchuk T, Constantinescu G, Michalak KM et al. Granzyme B-induced apoptosis requires both direct caspase activation and relief of caspase inhibition. Immunity 2003; 18: 355–365.
Bird CH, Sutton VR, Sun J, Hirst CE, Novak A, Kumar S et al. Selective regulation of apoptosis: the cytotoxic lymphocyte serpin proteinase inhibitor 9 protects against granzyme B-mediated apoptosis without perturbing the Fas cell death pathway. Mol Cell Biol 1998; 18: 6387–6398.
Medema JP, de Jong J, Peltenburg LT, Verdegaal EM, Gorter A, Bres SA et al. Blockade of the granzyme B/perforin pathway through overexpression of the serine protease inhibitor PI-9/SPI-6 constitutes a mechanism for immune escape by tumors. Proc Natl Acad Sci USA 2001; 98: 11515–11520.
Bladergroen BA, Meijer CJ, ten Berge RL, Hack CE, Muris JJ, Dukers DF et al. Expression of the granzyme B inhibitor, protease inhibitor 9, by tumor cells in patients with non-Hodgkin and Hodgkin lymphoma: a novel protective mechanism for tumor cells to circumvent the immune system? Blood 2002; 99: 232–237.
van Houdt IS, Oudejans JJ, van den Eertwegh AJ, Baars A, Vos W, Bladergroen BA et al. Expression of the apoptosis inhibitor protease inhibitor 9 predicts clinical outcome in vaccinated patients with stage III and IV melanoma. Clin Cancer Res 2005; 11: 6400–6407.
Casciola-Rosen L, Andrade F, Ulanet D, Wong WB, Rosen A . Cleavage by granzyme B is strongly predictive of autoantigen status: implications for initiation of autoimmunity. J Exp Med 1999; 190: 815–826.
Casiano CA, Martin SJ, Green DR, Tan EM . Selective cleavage of nuclear autoantigens during CD95 (Fas/APO-1)-mediated T cell apoptosis. J Exp Med 1996; 184: 765–770.
Casiano CA, Ochs RL, Tan EM . Distinct cleavage products of nuclear proteins in apoptosis and necrosis revealed by autoantibody probes. Cell Death Differ 1998; 5: 183–190.
Fischer U, Janicke RU, Schulze-Osthoff K . Many cuts to ruin: a comprehensive update of caspase substrates. Cell Death Differ 2003; 10: 76–100.
Goping IS, Sawchuk T, Underhill DA, Bleackley RC . Identification of {alpha}-tubulin as a granzyme B substrate during CTL-mediated apoptosis. J Cell Sci 2006; 119 (Pt 5): 858–865.
Adrain C, Duriez PJ, Brumatti G, Delivani P, Martin SJ . The cytotoxic lymphocyte protease, granzyme B, targets the cytoskeleton and perturbs microtubule polymerization dynamics. J Biol Chem 2006; 281: 8118–8125.
Suzuki K, Bose P, Leong-Quong RY, Fujita DJ, Riabowol K . REAP: a two minute cell fractionation method. BMC Res Notes 2010; 3: 294.
Staes A, Impens F, Van Damme P, Ruttens B, Goethals M, Demol H et al. Selecting protein N-terminal peptides by combined fractional diagonal chromatography. Nat Protoc 2011; 6: 1130–1141.
Van Damme P, Hole K, Pimenta-Marques A, Helsens K, Vandekerckhove J, Martinho RG et al. NatF contributes to an evolutionary shift in protein N-terminal acetylation and is important for normal chromosome segregation. PLoS Genet 2011; 7: e1002169.
Vizcaino JA, Cote RG, Csordas A, Dianes JA, Fabregat A, Foster JM et al. The PRoteomics IDEntifications (PRIDE) database and associated tools: status in 2013. Nucleic Acids Res 2013; 41 (Database issue): D1063–D1069.
Acknowledgements
This work was supported by the Dutch Cancer Society (UU-2009-4302 to NB), the Dutch Organization for Scientific Research (NWO) (grant number 916.66.044 (to NB)), and PRIME-XS (grant agreement number 262067, funded by the European Union 7th Framework Program). KP is supported by a PhD grant from the Institute for the Promotion of Innovation through Science and Technology in Flanders (IWT-Vlaanderen). PVD is a Postdoctoral Fellow of the Research Foundation-Flanders (FWO-Vlaanderen). The MS proteomics data have been deposited to the ProteomeXchange Consortium (http://proteomecentral.proteomexchange.org) via the PRIDE partner repository64 with the data set identifier PXD000252 and DOI 10.6019/PXD000252. Confocal microscopy was performed at the Cell Microscopy Center at the University Medical Center Utrecht, Utrecht, the Netherlands.
Author Contributions
SdP participated in the design of the study, performed experiments, analyzed and interpreted the data, and wrote the paper. KWL and LvdW performed experiments. KP and PVD participated in the design of the study, performed experiments, and analyzed and interpreted the data. KG participated in the design of the study and analyzed and interpreted the data. AP analyzed and interpreted the data and provided vital unique reagents. NB participated in the design of the study, analyzed and interpreted the data, and wrote the paper.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Edited by G Melino
Supplementary Information accompanies this paper on Cell Death and Differentiation website
Rights and permissions
About this article
Cite this article
de Poot, S., Lai, K., van der Wal, L. et al. Granzyme M targets topoisomerase II alpha to trigger cell cycle arrest and caspase-dependent apoptosis. Cell Death Differ 21, 416–426 (2014). https://doi.org/10.1038/cdd.2013.155
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/cdd.2013.155
Keywords
This article is cited by
-
Resistance of HIV-infected macrophages to CD8+ T lymphocyte–mediated killing drives activation of the immune system
Nature Immunology (2018)
-
Perforin and granzymes: function, dysfunction and human pathology
Nature Reviews Immunology (2015)
-
Granzyme M: behind enemy lines
Cell Death & Differentiation (2014)
-
NK cell intrinsic regulation of MIP-1α by granzyme M
Cell Death & Disease (2014)